Recognition: unknown
Vib2Conf: AI-driven discrimination of molecular conformations from vibrational spectra
Pith reviewed 2026-05-07 17:29 UTC · model grok-4.3
The pith
A deep learning model discriminates three-dimensional molecular conformations from vibrational spectra even for near-isomeric structures differing by roughly one angstrom in root-mean-square deviation.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
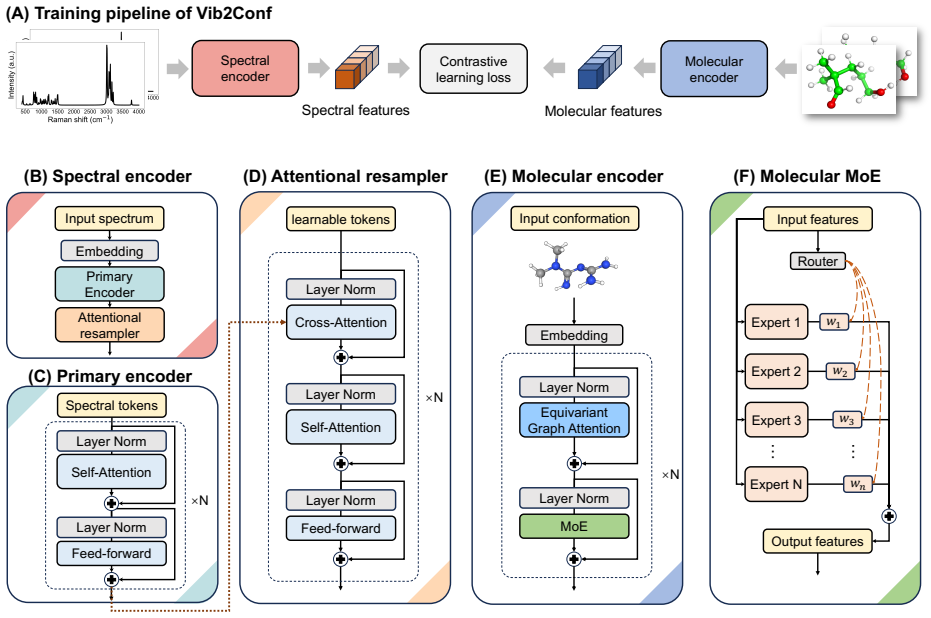
Vib2Conf directly discriminates 3D molecular conformations from vibrational spectra by distilling conformation-sensitive features from sparse signals with an attentional resampler and partitioning the conformational space with Mixture-of-Experts for precise geometric mapping, reaching state-of-the-art top-1 recall exceeding 95 percent on QM9S, VB-Mols, and QMe14S while achieving 82.06 percent top-1 recall on the VB-Confs test set for conformational isomers that differ by an RMSD of only approximately 1 Å.
What carries the argument
Attentional resampler that extracts conformation-sensitive features from sparse spectral signals together with Mixture-of-Experts that partitions conformational space for geometric mapping.
If this is right
- Spectrum-to-structure retrieval extends from two-dimensional connectivity to direct three-dimensional conformational discrimination.
- Conformers that differ by only about one angstrom RMSD become distinguishable from vibrational data alone.
- The same attentional and expert-based architecture yields state-of-the-art recall on multiple established spectrum-structure benchmarks.
- Fine-grained spectrum-to-conformation analysis becomes feasible for general use in molecular identification.
Where Pith is reading between the lines
- The approach could reduce the need for complementary experimental techniques such as crystallography when only spectroscopic data are available.
- Integration with existing computational workflows might allow real-time conformation assignment during spectroscopic experiments.
- The method opens a route to test whether similar modular networks can resolve other ambiguous inverse problems in molecular spectroscopy.
Load-bearing premise
The training distributions from QM9S, VB-Mols, QMe14S, and VB-Confs sufficiently represent the conformational heterogeneity and spectral noise found in real experimental measurements.
What would settle it
Measure vibrational spectra of a set of known near-isomeric conformers (RMSD ~1 Å) that were never seen during training and test whether Vib2Conf still returns the correct 3D structure as its top prediction.
Figures




read the original abstract
Retrieving or generating two-dimensional molecular structures on the basis of vibrational spectra has been well demonstrated via deep learning models. However, deciphering three-dimensional molecular conformations is still challenging, primarily due to spectral ambiguities caused by conformational heterogeneity, which are difficult to resolve. To address this limitation, we propose Vib2Conf, a deep learning model directly discriminating 3D molecular conformations from vibrational spectra. We implement an attentional resampler to distill conformation-sensitive features from sparse spectral signals, and integrate Mixture-of-Experts (MoE) to partition the conformational space for precise geometric mapping. These modules enable Vib2Conf to achieve state-of-the-art top-1 recall exceeding 95% on traditional spectrum-structure benchmarks, including QM9S, VB-Mols, and QMe14S. More importantly, Vib2Conf can discriminate near-isomeric conformers with a top-1 recall of 82.06% on VB-Confs test set, where conformational isomers differ by a root-mean-square deviation (RMSD) of only ~1 {\AA}. In general, Vib2Conf is a promising method for fine-grained spectrum-to-conformation analysis.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces Vib2Conf, a deep learning model that uses an attentional resampler to extract conformation-sensitive features from vibrational spectra and a Mixture-of-Experts (MoE) module to map these to 3D conformations. It reports state-of-the-art top-1 recall exceeding 95% on the QM9S, VB-Mols, and QMe14S benchmarks, and a top-1 recall of 82.06% on the VB-Confs test set for distinguishing near-isomeric conformers that differ by an RMSD of only ~1 Å.
Significance. If the performance claims hold under scrutiny, the work would represent a meaningful advance in spectrum-to-structure retrieval by addressing conformational ambiguity, which remains a core limitation in vibrational spectroscopy applications. The attentional resampler and MoE integration provide a plausible mechanism for handling sparse signals and partitioning conformational space. The reported results on multiple benchmarks, including the challenging near-isomer case, constitute a strength, though the idealized DFT-derived training data limit immediate claims of experimental utility.
major comments (2)
- [Results (VB-Confs)] Results section on VB-Confs benchmark: The central claim of 82.06% top-1 recall for ~1 Å RMSD conformers is load-bearing, yet no ablation studies are presented that isolate the contribution of the attentional resampler versus the MoE (or a baseline without either); without these, it is impossible to confirm that the architecture, rather than dataset artifacts, drives the discrimination of subtle spectral differences.
- [Methods] Methods section describing datasets and training: No details are provided on train/test splits, cross-validation strategy, random seeds, or any spectral noise augmentation for QM9S, VB-Mols, QMe14S, or VB-Confs; this omission directly affects assessment of whether the 82.06% recall reflects generalization or potential overfitting to noise-free DFT spectra.
minor comments (2)
- [Abstract] Abstract: The statement 'exceeding 95%' on the three benchmarks would be more informative if the exact per-dataset recalls were stated rather than aggregated.
- [Results] Notation: The RMSD threshold of '~1 Å' is used without a precise definition or distribution statistics for the VB-Confs pairs; adding a table or histogram of RMSD values would clarify the difficulty of the test cases.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed review of our manuscript. We address each major comment point by point below. Where the comments identify omissions that affect the strength of our claims, we commit to revisions that will incorporate the requested information and analyses.
read point-by-point responses
-
Referee: [Results (VB-Confs)] Results section on VB-Confs benchmark: The central claim of 82.06% top-1 recall for ~1 Å RMSD conformers is load-bearing, yet no ablation studies are presented that isolate the contribution of the attentional resampler versus the MoE (or a baseline without either); without these, it is impossible to confirm that the architecture, rather than dataset artifacts, drives the discrimination of subtle spectral differences.
Authors: We agree that ablation studies are necessary to rigorously attribute the 82.06% top-1 recall on VB-Confs to the attentional resampler and MoE components rather than dataset properties. In the revised manuscript we will add a dedicated ablation subsection in the Results, reporting performance of the full model against (i) a variant without the attentional resampler, (ii) a variant without the MoE module, and (iii) a simple baseline without either component. These experiments will be performed on the same VB-Confs test set and will be accompanied by statistical significance tests. revision: yes
-
Referee: [Methods] Methods section describing datasets and training: No details are provided on train/test splits, cross-validation strategy, random seeds, or any spectral noise augmentation for QM9S, VB-Mols, QMe14S, or VB-Confs; this omission directly affects assessment of whether the 82.06% recall reflects generalization or potential overfitting to noise-free DFT spectra.
Authors: We thank the referee for highlighting this gap. The revised Methods section will explicitly state the train/test split ratios and selection criteria for each benchmark, the cross-validation strategy employed, the random seeds used for all experiments (to ensure reproducibility), and whether spectral noise augmentation was applied (none was used, as all spectra were generated from noise-free DFT calculations). These additions will allow readers to evaluate generalization versus potential overfitting. revision: yes
Circularity Check
No circularity: performance claims rest on held-out test sets
full rationale
The paper reports empirical top-1 recall metrics (95%+ on QM9S/VB-Mols/QMe14S; 82.06% on VB-Confs) evaluated on explicitly held-out test sets whose construction is independent of the model's fitted parameters. The attentional resampler and MoE modules are architectural choices whose outputs are validated against external benchmarks rather than being defined in terms of those outputs. No equations, uniqueness theorems, or self-citations are invoked to force the reported numbers by construction. The central claims therefore remain falsifiable against the test distributions and do not reduce to the training inputs.
Axiom & Free-Parameter Ledger
Reference graph
Works this paper leans on
-
[1]
(2) Wang, Y.; Shen, Y.; Chen, S.; Wang, L.; Ye, F.; Zhou, H. Learning harmonic molecular representations on riemannian manifold.arXiv preprint arXiv:2303.155202023, (3) Du, Y.; Wang, L.; Feng, D.; Wang, G.; Ji, S.; Gomes, C. P.; Ma, Z.-M.; others A new perspective on building efficient and expressive 3D equivariant graph neural networks. Advances in neura...
-
[2]
The information bottleneck method
(21) Frisch, M.; Trucks, G.; Schlegel, H.; Scuseria, G.; Robb, M.; Cheeseman, J.; Scal- mani, G.; Barone, V.; Petersson, G.; Nakatsuji, H.; others Gaussian, Inc., Wallingford CT, 2016.Gaussian09, Revision D2016,1. (22) Best, R. B.; Zhu, X.; Shim, J.; Lopes, P. E.; Mittal, J.; Feig, M.; MacKerell Jr, A. D. Optimization of the additive CHARMM all-atom prote...
work page internal anchor Pith review arXiv 2016
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.