Recognition: unknown
Colloidal layer deposition with a controllable number of layers and compositional order
Pith reviewed 2026-05-09 17:58 UTC · model grok-4.3
The pith
DNA-decorated colloids and surfaces self-assemble into crystallites with controlled layer count and alternating composition
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
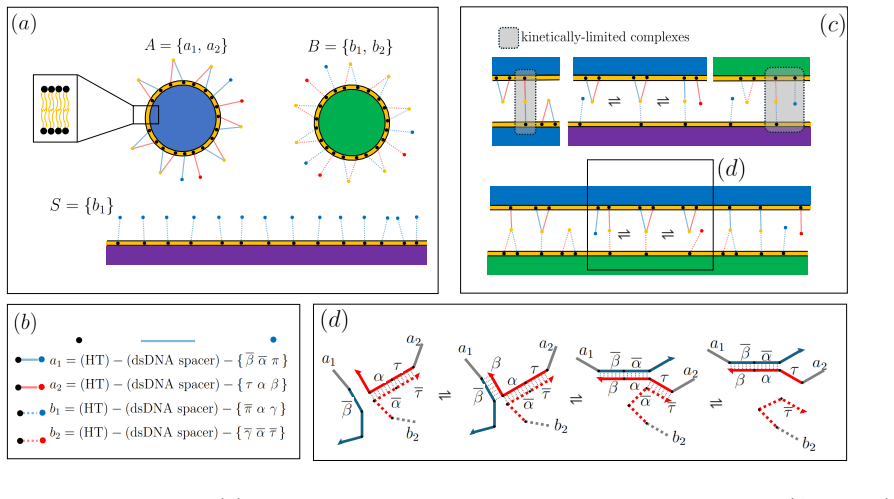
We design a system with a binary suspension of colloids and a surface that triggers the self-assembly of crystallites with a finite thickness. The proposed design allows controlling the number of layers forming the aggregate and constrains the two types of particles to lie on different planes. These functionalities are achieved by decorating the colloids and the surface with multiple DNA oligomers featuring specific interactions. The surface triggers a chain of reactions between DNA oligomers, leading to localized self-assembly. Equilibrium principles control the thickness of the aggregates. Instead, compositional order is achieved by engineering the reaction kinetics between DNA oligomers.
What carries the argument
Multiple DNA oligomers with specific interactions attached to colloids and surface, triggering a localized reaction chain whose equilibrium sets aggregate thickness and whose kinetics enforce compositional order by limiting same-type binding.
Load-bearing premise
That DNA oligomers can be designed and attached so the intended specific bindings dominate without significant cross-reactivity or off-target effects, and that the reaction-diffusion model captures the actual multibody colloidal behavior.
What would settle it
An experiment or simulation showing aggregates whose thickness varies independently of equilibrium conditions or where same-type colloids occupy the same plane would falsify the claimed control.
Figures











read the original abstract
We design a system with a binary suspension of colloids and a surface that triggers the self-assembly of crystallites with a finite thickness. The proposed design allows controlling the number of layers forming the aggregate and constrains the two types of particles to lie on different planes. These functionalities are achieved by decorating the colloids and the surface with multiple DNA oligomers featuring specific interactions. The surface triggers a chain of reactions between DNA oligomers, leading to localized self-assembly. Equilibrium principles control the thickness of the aggregates. Instead, compositional order is achieved by engineering the reaction kinetics between DNA oligomers in a way that limits interactions between colloids of the same type. We validate our design using theory and reaction-diffusion simulation algorithms, which capture the multibody nature of the interactions. This work demonstrates how engineering the kinetics provides a new avenue for controlling the morphology of aggregates assembled by DNA.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proposes a design for a binary colloidal suspension and functionalized surface decorated with multiple DNA oligomers that triggers localized self-assembly into crystallites of finite, controllable thickness with compositional order (particles of different types constrained to distinct planes). Thickness is asserted to be set by equilibrium binding principles, while compositional order is imposed by kinetic engineering that suppresses same-type colloid interactions; the design is validated through theory and reaction-diffusion simulations that incorporate multibody interactions.
Significance. If the equilibrium-kinetics separation can be realized without feedback, the approach offers a route to morphology control in DNA-mediated colloidal assembly that goes beyond purely thermodynamic designs, with potential applications in layered materials. The explicit use of reaction-diffusion simulations to capture multibody effects is a positive methodological choice, though the absence of quantitative error analysis, specific parameter values, or direct comparisons to equilibrium-only cases in the validation limits the strength of the supporting evidence.
major comments (2)
- [Design and validation sections] The central design claim requires decoupling equilibrium control of thickness from kinetic control of order, but DNA sequence design inherently links on-rates, off-rates, and equilibrium constants (K_eq = k_on/k_off). No derivation or parameter scan is provided showing that sequences can be chosen to suppress A-A/B-B kinetics while preserving the surface-particle and inter-layer affinities needed for finite stacking. This appears in the design description and validation sections.
- [Validation section] The abstract and validation statement assert that theory and reaction-diffusion simulations confirm the design, yet no quantitative metrics (e.g., layer-number histograms, order parameters, or comparison of simulated vs. predicted thickness) or parameter values are referenced. Without these, it is impossible to assess whether the simulations actually demonstrate independent control or merely reproduce the input assumptions.
minor comments (2)
- Notation for the distinct DNA oligomers (e.g., labels for surface vs. particle strands) should be introduced with a table or explicit definitions early in the manuscript to improve readability.
- The reaction-diffusion model description would benefit from a brief statement of the diffusion coefficients and reaction rate constants used, even if they are illustrative.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed comments on our manuscript. We address each major comment below and will revise the manuscript to incorporate additional details and quantitative validation as suggested.
read point-by-point responses
-
Referee: [Design and validation sections] The central design claim requires decoupling equilibrium control of thickness from kinetic control of order, but DNA sequence design inherently links on-rates, off-rates, and equilibrium constants (K_eq = k_on/k_off). No derivation or parameter scan is provided showing that sequences can be chosen to suppress A-A/B-B kinetics while preserving the surface-particle and inter-layer affinities needed for finite stacking. This appears in the design description and validation sections.
Authors: We agree that for any given hybridization reaction the on-rate, off-rate and equilibrium constant are linked. However, our design uses multiple orthogonal DNA sequences assigned to distinct interaction types (surface-A, A-B, B-surface, etc.). For A-A and B-B pairs we select sequences with minimal or no complementarity, rendering their k_on and K_eq negligibly small and thereby suppressing same-type interactions. The sequences for the required surface-particle and hetero inter-layer bindings are chosen independently to maintain strong complementarity and the affinities needed for finite stacking. Equilibrium thus governs layer number via the hetero-binding energies, while the absence of same-type binding enforces compositional order. We will add to the revised manuscript a short derivation of the binding-energy conditions together with example sequence choices and a parameter scan confirming that finite thickness and order can be achieved simultaneously. revision: yes
-
Referee: [Validation section] The abstract and validation statement assert that theory and reaction-diffusion simulations confirm the design, yet no quantitative metrics (e.g., layer-number histograms, order parameters, or comparison of simulated vs. predicted thickness) or parameter values are referenced. Without these, it is impossible to assess whether the simulations actually demonstrate independent control or merely reproduce the input assumptions.
Authors: We acknowledge that the validation section presents results qualitatively and does not include explicit quantitative metrics, tabulated parameters, or comparisons to equilibrium-only cases. In the revised manuscript we will add: (i) histograms of assembled layer numbers, (ii) compositional order parameters (e.g., layer-resolved type fractions), (iii) a table of all simulation parameters, and (iv) a direct comparison of simulated average thickness against the equilibrium-theory prediction. We will also include equilibrium-only control simulations to demonstrate the necessity of the kinetic suppression. These additions will allow quantitative assessment of independent control. revision: yes
Circularity Check
No circularity: design claims rest on independent engineering choices, not self-referential fits or citations
full rationale
The paper presents a colloidal design using DNA oligomers where thickness is controlled by equilibrium binding affinities and compositional order by separate kinetic engineering to suppress same-type interactions. No equations, fitted parameters, or derivation steps are exhibited that reduce either claim to a quantity defined by the result itself. Validation via reaction-diffusion simulations is asserted to capture multibody effects without evidence that the model parameters are constructed from the target morphology. No self-citation load-bearing steps or uniqueness theorems imported from prior author work appear in the provided text. The separation of equilibrium and kinetics is framed as a design feature rather than a tautological prediction.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption DNA oligomers can be engineered to exhibit the required binding affinities and kinetic rates without significant cross-reactivity
Reference graph
Works this paper leans on
-
[1]
A.; Letsinger, R
Mirkin, C. A.; Letsinger, R. L.; Mucic, R. C.; Storhoff, J. J. A DNA -based method for rationally assembling nanoparticles into macroscopic materials. Nature 1996, 382, 607--609
1996
-
[2]
P.; Johnsson, K
Alivisatos, A. P.; Johnsson, K. P.; Peng, X.; Wilson, T. E.; Loweth, C. J.; Bruchez, M. P.; Schultz, P. G. Organization of 'nanocrystal molecules' using DNA . Nature 1996, 382, 609--611
1996
-
[3]
R.; Seeman, N
Jones, M. R.; Seeman, N. C.; Mirkin, C. A. Programmable materials and the nature of the DNA bond. Science 2015, 347, 1260901
2015
-
[4]
A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics
SantaLucia, J. A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics. Proc.\ Natl.\ Acad.\ Sci.\ U.\ S.\ A. 1998, 95, 1460--1465
1998
-
[5]
E.; Porubsky, N
Fornace, M. E.; Porubsky, N. J.; Pierce, N. A. A unified dynamic programming framework for the analysis of interacting nucleic acid strands: enhanced models, scalability, and speed. ACS Synthetic Biology 2020, 9, 2665--2678
2020
-
[6]
H.; Eiser, E.; Foffi, G
Varrato, F.; Di Michele, L.; Belushkin, M.; Dorsaz, N.; Nathan, S. H.; Eiser, E.; Foffi, G. Arrested demixing opens route to bigels. Proc.\ Natl.\ Acad.\ Sci.\ U.\ S.\ A. 2012, 109, 19155--19160
2012
-
[7]
L.; Wang, T.; Emamy, H.; Li, H.; Yager, K
Liu, W.; Tagawa, M.; Xin, H. L.; Wang, T.; Emamy, H.; Li, H.; Yager, K. G.; Starr, F. W.; Tkachenko, A. V.; Gang, O. Diamond family of nanoparticle superlattices. Science 2016, 351, 582--586
2016
-
[8]
C.; McGinley, J
Wang, Y.; Jenkins, I. C.; McGinley, J. T.; Sinno, T.; Crocker, J. C. Colloidal crystals with diamond symmetry at optical lengthscales. Nat.\ Commun. 2017, 8, 14173
2017
-
[9]
Ducrot, \'E .; He, M.; Yi, G.-R.; Pine, D. J. Colloidal alloys with preassembled clusters and spheres. Nat.\ Mater. 2017, 16, 652
2017
-
[10]
Table of Elements
Macfarlane, R. J.; O'Brien, M. N.; Petrosko, S. H.; Mirkin, C. A. Nucleic Acid‐Modified Nanostructures as Programmable Atom Equivalents: Forging a New “Table of Elements”. Angew.\ Chem., Int.\ Ed. 52, 5688--5698
-
[11]
M.; van der Lelie, D.; Gang, O
Nykypanchuk, D.; Maye, M. M.; van der Lelie, D.; Gang, O. DNA -guided crystallization of colloidal nanoparticles. Nature 2008, 451, 549--552
2008
-
[12]
Y.; Lytton-Jean, A
Park, S. Y.; Lytton-Jean, A. K. R.; Lee, B.; Weigand, S.; Schatz, G. C.; Mirkin, C. A. DNA -programmable nanoparticle crystallization. Nature 2008, 451, 553--556
2008
-
[13]
Preparation Techniques, Design Strategies of Responsive Photonic Crystals and Their Typical Applications in the Field of Sensing
Ye, R.; Yang, R.; Hu, L.; Li, Z.; Luo, Z.; Chen, X. Preparation Techniques, Design Strategies of Responsive Photonic Crystals and Their Typical Applications in the Field of Sensing. Crystals 2026, 16, 232
2026
-
[14]
D.; Tkachenko, A
Halverson, J. D.; Tkachenko, A. V. DNA-programmed mesoscopic architecture. Phys.\ Rev.\ E 2013, 87, 062310
2013
-
[15]
Patra, N.; Tkachenko, A. V. Layer-by-layer assembly of patchy particles as a route to nontrivial structures. Phys.\ Rev.\ E 2017, 96, 022601
2017
-
[16]
K.; Mognetti, B
Jana, P. K.; Mognetti, B. M. Surface-triggered cascade reactions between DNA linkers direct the self-assembly of colloidal crystals of controllable thickness. Nanoscale 2019, 11, 5450--5459
2019
-
[17]
A.; Holmes-Cerfon, M
Marbach, S.; Zheng, J. A.; Holmes-Cerfon, M. The nanocaterpillar's random walk: diffusion with ligand--receptor contacts. Soft Matter 2022, 18, 3130--3146
2022
-
[18]
Lowensohn, J.; Stevens, L.; Goldstein, D.; Mognetti, B. M. Sliding across a surface: Particles with fixed and mobile ligands. The Journal of Chemical Physics 2022, 156
2022
-
[19]
Phase-transition-like behaviors of sequence-selective dynamic bonds
Dai, X.; Wang, Y.; Wei, W.; Jiao, Z.; Chen, W.; Cheng, L.; Yan, L.-T. Phase-transition-like behaviors of sequence-selective dynamic bonds. Proceedings of the National Academy of Sciences 2026, 123, e2514117123
2026
-
[20]
S.; Weck, M.; Pine, D
Wang, Y.; Wang, Y.; Zheng, X.; Ducrot, \'E .; Yodh, J. S.; Weck, M.; Pine, D. J. Crystallization of DNA-coated colloids. Nature communications 2015, 6, 7253
2015
-
[21]
L.; Kim, A
Biancaniello, P. L.; Kim, A. J.; Crocker, J. C. Colloidal interactions and self-assembly using DNA hybridization. Physical review letters 2005, 94, 058302
2005
-
[22]
Proceedings of the National Academy of Sciences 2020, 117, 5617--5623
Zhou, X.; Yao, D.; Hua, W.; Huang, N.; Chen, X.; Li, L.; He, M.; Zhang, Y.; Guo, Y.; Xiao, S.; others Programming colloidal bonding using DNA strand-displacement circuitry. Proceedings of the National Academy of Sciences 2020, 117, 5617--5623
2020
-
[23]
E.; Seyforth, H.; Jacobs, W
Hensley, A.; Videb k, T. E.; Seyforth, H.; Jacobs, W. M.; Rogers, W. B. Macroscopic photonic single crystals via seeded growth of DNA-coated colloids. Nature communications 2023, 14, 4237
2023
-
[24]
Y.; Winfree, E
Zhang, D. Y.; Winfree, E. Control of DNA strand displacement kinetics using toehold exchange. Journal of the American Chemical Society 2009, 131, 17303--17314
2009
-
[25]
Parolini, L.; Kotar, J.; Di Michele, L.; Mognetti, B. M. Controlling self-assembly kinetics of DNA-functionalized liposomes using toehold exchange mechanism. ACS nano 2016, 10, 2392--2398
2016
-
[26]
B.; Manoharan, V
Rogers, W. B.; Manoharan, V. N. Programming colloidal phase transitions with DNA strand displacement. Science 2015, 347, 639--642
2015
-
[27]
K.; Mognetti, B
Jana, P. K.; Mognetti, B. M. Self-assembly of finite-sized colloidal aggregates. Soft matter 2020, 16, 5915--5924
2020
-
[28]
K.; Bruylants, G.; Cicuta, P.; Mognetti, B
Lanfranco, R.; Jana, P. K.; Bruylants, G.; Cicuta, P.; Mognetti, B. M.; Di Michele, L. Adaptable DNA interactions regulate surface-triggered self-assembly. Nanoscale 2020, 12, 18616--18620
2020
-
[29]
Robust self-replication of combinatorial information via crystal growth and scission
Schulman, R.; Yurke, B.; Winfree, E. Robust self-replication of combinatorial information via crystal growth and scission. Proceedings of the national academy of sciences 2012, 109, 6405--6410
2012
-
[30]
D.; Schulman, R.; Rothemund, P
Barish, R. D.; Schulman, R.; Rothemund, P. W.; Winfree, E. An information-bearing seed for nucleating algorithmic self-assembly. Proceedings of the National Academy of Sciences 2009, 106, 6054--6059
2009
-
[31]
G.; O’Brien, J.; Winfree, E.; Murugan, A
Evans, C. G.; O’Brien, J.; Winfree, E.; Murugan, A. Pattern recognition in the nucleation kinetics of non-equilibrium self-assembly. Nature 2024, 625, 500--507
2024
-
[32]
S.; Elghanian, R.; Thomas, A
Thaxton, C. S.; Elghanian, R.; Thomas, A. D.; Stoeva, S. I.; Lee, J.-S.; Smith, N. D.; Schaeffer, A. J.; Klocker, H.; Horninger, W.; Bartsch, G.; Mirkin, C. A. Nanoparticle-based bio-barcode assay redefines ``undetectable''PSA and biochemical recurrence after radical prostatectomy. Proc.\ Natl.\ Acad.\ Sci.\ U.\ S.\ A. 2009, 106, 18437--18442
2009
-
[33]
Chapman, R.; Lin, Y.; Burnapp, M.; Bentham, A.; Hillier, D.; Zabron, A.; Khan, S.; Tyreman, M.; Stevens, M. M. Multivalent Nanoparticle Networks Enable Point-of-Care Detection of Human Phospholipase-A2 in Serum. ACS Nano 2015, 9, 2565--2573, PMID: 25756526
2015
-
[34]
J.; Frenkel, D
Martinez-Veracoechea, F. J.; Frenkel, D. Designing super selectivity in multivalent nano-particle binding. Proceedings of the National Academy of Sciences 2011, 108, 10963--10968
2011
-
[35]
A.; Tkachenko, A
Licata, N. A.; Tkachenko, A. V. Kinetic limitations of cooperativity-based drug delivery systems. Physical review letters 2008, 100, 158102
2008
-
[36]
Self-assembly of DNA origami for nanofabrication, biosensing, drug delivery, and computational storage
He, Z.; Shi, K.; Li, J.; Chao, J. Self-assembly of DNA origami for nanofabrication, biosensing, drug delivery, and computational storage. Iscience 2023, 26
2023
-
[37]
Advances in programmable DNA nanostructures enabling stimuli-responsive drug delivery and multimodal biosensing
Hong, Y.; Ma, W.; Wang, M.; Wang, H.-H. Advances in programmable DNA nanostructures enabling stimuli-responsive drug delivery and multimodal biosensing. RSC Chemical Biology 2025, 6, 1366--1385
2025
-
[38]
van der Meulen, S. A. J.; Leunissen, M. E. Solid Colloids with Surface-Mobile DNA Linkers. J. Am. Chem. Soc. 2013, 135, 15129--15134
2013
-
[39]
W.; Chakraborty, I.; Kraft, D
Rinaldin, M.; Verweij, R. W.; Chakraborty, I.; Kraft, D. J. Colloid supported lipid bilayers for self-assembly. Soft Matter 2019,
2019
-
[40]
A.; Vanderlick, T
Beales, P. A.; Vanderlick, T. K. Specific Binding of Different Vesicle Populations by the Hybridization of Membrane-Anchored DNA . J. Phys. Chem. A 2007, 111, 12372--12380
2007
-
[41]
T.; Fellermann, H.; Eggenberger Hotz, P.; Hanczyc, M
Hadorn, M.; Boenzli, E.; S rensen, K. T.; Fellermann, H.; Eggenberger Hotz, P.; Hanczyc, M. M. Specific and reversible DNA -directed self-assembly of oil-in-water emulsion droplets. Proc.\ Natl.\ Acad.\ Sci.\ U.\ S.\ A. 2012, 109, 20320--20325
2012
-
[42]
M.; Kotar, J.; Eiser, E.; Cicuta, P.; Di Michele, L
Parolini, L.; Mognetti, B. M.; Kotar, J.; Eiser, E.; Cicuta, P.; Di Michele, L. Volume and porosity thermal regulation in lipid mesophases by coupling mobile ligands to soft membranes. Nat.\ Commun. 2015, 6, 5948
2015
-
[43]
M.; Frenkel, D
Angioletti-Uberti, S.; Varilly, P.; Mognetti, B. M.; Frenkel, D. Mobile linkers on DNA-coated colloids: valency without patches. Phys.\ Rev.\ Lett. 2014, 113, 128303--128306
2014
-
[44]
M.; Cicuta, P.; Di Michele, L
Mognetti, B. M.; Cicuta, P.; Di Michele, L. Programmable interactions with biomimetic DNA linkers at fluid membranes and interfaces. Reports on progress in physics 2019, 82, 116601
2019
-
[45]
J.; Kotar, J.; Parolini, L.; S ari \'c , A.; Cicuta, P.; Di Michele, L.; Mognetti, B
Bachmann, S. J.; Kotar, J.; Parolini, L.; S ari \'c , A.; Cicuta, P.; Di Michele, L.; Mognetti, B. M. Melting transition in lipid vesicles functionalised by mobile DNA linkers. Soft Matter 2016, 12, 7804--7817
2016
-
[46]
Sciortino, F.; Zhang, Y.; Gang, O.; Kumar, S. K. Combinatorial-entropy-driven aggregation in DNA-grafted nanoparticles. ACS nano 2020, 14, 5628--5635
2020
-
[47]
A.; Mahmoudabadi, G.; Milam, V
Baker, B. A.; Mahmoudabadi, G.; Milam, V. T. Strand displacement in DNA-based materials systems. Soft Matter 2013, 9, 11160--11172
2013
-
[48]
K.; Milam, V
Tison, C. K.; Milam, V. T. Programming the kinetics and extent of colloidal disassembly using a DNA trigger. Soft Matter 2010, 6, 4446--4453
2010
-
[49]
C.; Brujic, J.; Chaikin, P
Zhang, Y.; McMullen, A.; Pontani, L.-L.; He, X.; Sha, R.; Seeman, N. C.; Brujic, J.; Chaikin, P. M. Sequential self-assembly of DNA functionalized droplets. Nat.\ Commun. 2017, 8, 21
2017
-
[50]
Designing 3D multicomponent self-assembling systems with signal-passing building blocks
Evans, J.; S ulc, P. Designing 3D multicomponent self-assembling systems with signal-passing building blocks. The Journal of Chemical Physics 2024, 160
2024
-
[51]
D.; Tkachenko, A
Halverson, J. D.; Tkachenko, A. V. Sequential programmable self-assembly: Role of cooperative interactions. The Journal of chemical physics 2016, 144
2016
-
[52]
S.; Petitzon, M.; Mognetti, B
Bachmann, J. S.; Petitzon, M.; Mognetti, B. M. Bond formation kinetics affects self-assembly directed by ligand-receptor interactions. Soft Matter 2016, 12, 9585--9592
2016
-
[53]
Ranganathan, S.; Shakhnovich, E. I. Dynamic metastable long-living droplets formed by sticker-spacer proteins. Elife 2020, 9, e56159
2020
-
[54]
Gillespie, D. T. Exact stochastic simulation of coupled chemical reactions. The journal of physical chemistry 1977, 81, 2340--2361
1977
-
[55]
E.; S ulc, P.; Schaeffer, J
Srinivas, N.; Ouldridge, T. E.; S ulc, P.; Schaeffer, J. M.; Yurke, B.; Louis, A. A.; Doye, J. P.; Winfree, E. On the biophysics and kinetics of toehold-mediated DNA strand displacement. Nucleic acids research 2013, 41, 10641--10658
2013
-
[56]
L.; Dehmelt, F
Ho, D.; Zimmermann, J. L.; Dehmelt, F. A.; Steinbach, U.; Erdmann, M.; Severin, P.; Falter, K.; Gaub, H. E. Force-driven separation of short double-stranded DNA. Biophysical journal 2009, 97, 3158--3167
2009
-
[57]
Mitra, G.; Chang, C.; McMullen, A.; Puchall, D.; Brujic, J.; Hocky, G. M. A coarse-grained simulation model for colloidal self-assembly via explicit mobile binders. Soft Matter 2023, 19, 4223--4236
2023
-
[58]
P.; Tildesley, D
Allen, M. P.; Tildesley, D. J. Computer simulation of liquids; Oxford university press, 2017
2017
-
[59]
Propagation of large concentration changes in reversible protein-binding networks
Maslov, S.; Ispolatov, I. Propagation of large concentration changes in reversible protein-binding networks. Proceedings of the National Academy of Sciences 2007, 104, 13655--13660
2007
-
[60]
J.; Parolini, L.; Mognetti, B
Di Michele, L.; Bachmann, S. J.; Parolini, L.; Mognetti, B. M. Communication: free energy of ligand-receptor systems forming multimeric complexes. The Journal of Chemical Physics 2016, 144
2016
-
[61]
Understanding molecular simulation: from algorithms to applications; elsevier, 2023
Frenkel, D.; Smit, B. Understanding molecular simulation: from algorithms to applications; elsevier, 2023
2023
-
[62]
E.; Sha, R.; Tkachenko, A
Dreyfus, R.; Leunissen, M. E.; Sha, R.; Tkachenko, A. V.; Seeman, N. C.; Pine, D. J.; Chaikin, P. M. Simple Quantitative Model for the Reversible Association of DNA Coated Colloids. Phys. Rev. Lett. 2009, 102, 048301 mcitethebibliography document
2009
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.