Recognition: unknown
ARA: Agentic Reproducibility Assessment For Scalable Support Of Scientific Peer-Review
Pith reviewed 2026-05-08 01:41 UTC · model grok-4.3
The pith
An agentic system extracts directed workflow graphs from papers to score reproducibility at 61 percent accuracy.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
ARA formalizes reproducibility assessment as a structured reasoning task over scientific documents. Given a paper, ARA extracts a directed workflow graph linking sources, methods, experiments, and outputs, then evaluates its reconstructability using structural and content-based scores for reproducibility assessments. Experiments on 213 ReScience C articles demonstrate ARA's generalizability and consistent workflow reconstruction and assessment across LLMs, model temperatures, and scientific domains. ARA achieves the highest accuracy reported on ReproBench (60.71 percent versus 36.84 percent) and GoldStandardDB (61.68 percent versus 43.56 percent).
What carries the argument
The directed workflow graph that links sources, methods, experiments, and outputs, evaluated through structural and content-based reconstructability scores.
If this is right
- ARA produces consistent workflow reconstructions and scores regardless of the specific LLM or temperature setting used.
- The method outperforms prior approaches on the ReproBench and GoldStandardDB benchmarks.
- Workflow extraction and scoring generalize across multiple scientific domains in a 213-paper human-validated set.
- The approach supplies scalable, structured input that can complement human reviewers during peer review.
Where Pith is reading between the lines
- Journals could run ARA as an automated pre-filter to flag papers likely to have reproducibility issues before full human review.
- The graph-based representation might be adapted to capture non-computational experiments by adding new node and edge types.
- Hybrid human-AI pipelines could use ARA scores to allocate reviewer effort more efficiently on high-uncertainty submissions.
Load-bearing premise
LLM extraction of workflow graphs followed by structural and content scoring produces assessments that match human expert judgments without systematic distortion from hallucinations, incomplete text, or domain knowledge gaps.
What would settle it
A side-by-side test on papers with known reproduction outcomes where human experts independently score reproducibility and the ARA scores are checked for high correlation or predictive power.
Figures


read the original abstract
Scientific peer review increasingly struggles to assess reproducibility at the scale and complexity of modern research output. Evaluating reproducibility requires reconstructing experimental dependencies, methodological choices, data flows, and result-generating procedures, which often exceeds what human reviewers can provide. Agentic Reproducibility Assessment (ARA) formalizes reproducibility assessment as a structured reasoning task over scientific documents. Given a paper, ARA extracts a directed workflow graph linking sources, methods, experiments, and outputs, then evaluates its reconstructability using structural and content-based scores for reproducibility assessments. Experiments on 213 ReScience C articles - the largest cross-domain benchmark of human-validated computational reproducibility studies considered to date - demonstrate ARA's generalizability and consistent workflow reconstruction and assessment across LLMs, model temperatures, and scientific domains. ARA achieves ~61% accuracy on three benchmarks, and the highest accuracy reported on ReproBench (60.71% vs. 36.84%) and GoldStandardDB (61.68% vs. 43.56%), highlighting its potential to complement human review at scale and enabling next-generation peer review. Code and Data available: https://github.com/AndresLaverdeMarin/agentic_reproducibility_assessment.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper presents Agentic Reproducibility Assessment (ARA), which uses LLMs to extract a directed workflow graph from a scientific paper linking sources, methods, experiments, and outputs, followed by structural and content-based scoring to assess reproducibility. It reports experiments on 213 ReScience C articles showing generalizability across LLMs, temperatures, and domains, with ~61% accuracy on three benchmarks and superior performance on ReproBench (60.71% vs 36.84%) and GoldStandardDB (61.68% vs 43.56%).
Significance. If the core extraction and scoring pipeline can be shown to align with human expert judgment, ARA could provide a scalable complement to human peer review for assessing computational reproducibility at the volume of modern research output. The scale of the primary benchmark (213 human-validated articles across domains) and direct head-to-head comparisons against prior methods on ReproBench and GoldStandardDB are positive features.
major comments (3)
- [Methods (workflow extraction and graph construction)] The accuracy claims (~61% overall, 60.71% on ReproBench, 61.68% on GoldStandardDB) depend on the fidelity of the LLM-generated directed workflow graphs. The manuscript describes no human annotation, node/edge fidelity metrics, or error analysis of these graphs against the source papers or reproduction reports. Without such validation, it is unclear whether the reported numbers measure reproducibility or LLM extraction artifacts.
- [Experiments] The Experiments section provides no selection criteria for the 213 ReScience C articles, no exact definitions or formulas for the structural and content scores, and no ablation results on individual ARA components. These omissions make it impossible to assess the claimed cross-domain consistency and generalizability.
- [Results] The Results section reports benchmark accuracies without error bars, confidence intervals, or statistical significance tests for the improvements over baselines. Claims of consistency across LLMs and model temperatures are stated but not supported by quantitative tables or variance measures.
minor comments (2)
- [Abstract] The abstract states that code and data are available at a GitHub link; confirm that the repository contains the full evaluation pipeline and the 213-article dataset splits used for the reported numbers.
- Clarify the precise relationship among the 'three benchmarks' mentioned in the abstract and the two named datasets (ReproBench, GoldStandardDB) to avoid ambiguity in the performance claims.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed comments. These have highlighted important areas where additional clarity and supporting analyses will strengthen the manuscript. We address each major comment below and commit to the indicated revisions in the next version.
read point-by-point responses
-
Referee: The accuracy claims (~61% overall, 60.71% on ReproBench, 61.68% on GoldStandardDB) depend on the fidelity of the LLM-generated directed workflow graphs. The manuscript describes no human annotation, node/edge fidelity metrics, or error analysis of these graphs against the source papers or reproduction reports. Without such validation, it is unclear whether the reported numbers measure reproducibility or LLM extraction artifacts.
Authors: We agree that the absence of explicit graph fidelity validation makes it harder to rule out extraction artifacts as a contributor to the reported accuracies. The primary accuracy figures are computed by comparing ARA's final reproducibility scores against the human-provided ground truth in the ReScience C reproduction reports and the two external benchmarks; however, this does not isolate the quality of the intermediate workflow graphs. In the revision we will add a dedicated error-analysis subsection that (a) presents representative examples of node- and edge-level extraction errors, (b) reports precision/recall for key node categories on a manually inspected sample of 30 papers, and (c) discusses how the downstream structural and content scores are designed to be robust to certain classes of extraction noise. These additions will make the relationship between graph quality and final accuracy transparent. revision: yes
-
Referee: The Experiments section provides no selection criteria for the 213 ReScience C articles, no exact definitions or formulas for the structural and content scores, and no ablation results on individual ARA components. These omissions make it impossible to assess the claimed cross-domain consistency and generalizability.
Authors: The 213 articles comprise the complete set of ReScience C papers that possessed publicly available reproduction reports at the time of dataset construction; we will state this selection criterion explicitly. The structural score quantifies graph connectivity, completeness, and topological alignment with expected experimental flow, while the content score measures semantic overlap between extracted elements and the source text via embedding similarity; we will insert the precise mathematical definitions, weighting schemes, and pseudocode into the Methods section. We will also add an ablation study that reports accuracy when each major component (workflow extraction, structural scoring, content scoring) is removed or replaced, thereby quantifying their individual contributions to cross-domain performance. revision: yes
-
Referee: The Results section reports benchmark accuracies without error bars, confidence intervals, or statistical significance tests for the improvements over baselines. Claims of consistency across LLMs and model temperatures are stated but not supported by quantitative tables or variance measures.
Authors: We will augment the Results section with (i) error bars and 95 % confidence intervals computed via bootstrap resampling for all accuracy figures, (ii) paired statistical significance tests (McNemar’s test for binary reproducibility decisions and Wilcoxon signed-rank tests for score distributions) comparing ARA against the reported baselines, and (iii) a new table that lists mean accuracy and standard deviation across the LLMs and temperature settings examined. These quantitative measures will directly support the consistency claims. revision: yes
Circularity Check
No circularity: ARA accuracy claims derive from external human-validated benchmarks
full rationale
The paper defines ARA as an LLM-based extraction of directed workflow graphs from papers followed by structural/content scoring for reproducibility assessment. Reported results (~61% accuracy across three benchmarks, with specific gains on ReproBench and GoldStandardDB) are measured against independent, pre-existing human-validated datasets (213 ReScience C articles plus the other two benchmarks) and compared to prior methods. No equations, parameter fits, or self-citations reduce the central claims to tautological inputs or self-defined quantities. The derivation chain remains self-contained against external references.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption LLMs can extract accurate directed workflow graphs linking sources, methods, experiments, and outputs from arbitrary scientific documents
Reference graph
Works this paper leans on
-
[1]
G. Parchomovsky, “Publish or perish,”Michigan Law Review, vol. 98, no. 4, pp. 926–952, 2000. doi: 10.2307/1290335
-
[2]
Science in an exponential world,
A. Szalay and J. Gray, “Science in an exponential world,”Nature, vol. 440, no. 7083, pp. 413–414, 2006. doi: 10.1038/440413a
-
[3]
E. Mosca, M. H. I. Abdalla, P. Basso, M. Musumeci, and G. Groh, “Distinguishing fact from fiction: A benchmark dataset for identifying machine-generated scientific papers in the llm era.” inProceedings of the 3rd Workshop on Trustworthy Natural Language Processing (TrustNLP 2023), 2023, pp. 190–207. doi: 10.18653/v1/2023.trustnlp-1.17
-
[4]
H.-Z. Cheng, B. Sheng, A. Lee, V . Chaudhary, A. G. Atanasov, N. Liu, Y . Qiu, T. Y . Wong, Y .-C. Tham, and Y .-F. Zheng, “Have ai-generated texts from llm infiltrated the realm of sci- entific writing? a large-scale analysis of preprint platforms,”bioRxiv, pp. 2024–03, 2024. doi: 10.1101/2024.03.25.586710
-
[5]
Is llm a reliable reviewer? a comprehensive evaluation of llm on automatic paper reviewing tasks,
R. Zhou, L. Chen, and K. Yu, “Is llm a reliable reviewer? a comprehensive evaluation of llm on automatic paper reviewing tasks,” inProceedings of the 2024 joint international conference on computational linguistics, language resources and evaluation (LREC-COLING 2024), 2024, pp. 9340–9351. [Online]. Available: https://aclanthology.org/2024.lrec-main.816/
2024
-
[6]
G. Ellison, “Is peer review in decline?”Economic Inquiry, vol. 49, no. 3, pp. 635–657, 2011. doi: 10.1111/j.1465-7295.2010.00261.x
-
[7]
Q. Wei, S. Holt, J. Yang, M. Wulfmeier, and M. van der Schaar, “The ai imperative: Scaling high-quality peer review in machine learning,”arXiv preprint arXiv:2506.08134, 2025. doi: 10.48550/arXiv.2506.08134
-
[8]
K. Popper,Logik der Forschung. Vienna, Austria: Julius Springer Verlag GmbH, 1935. doi: 10.1007/978-3-7091-4177-9
-
[9]
London, UK: Hutchinson & Co., 1959
——,The Logic of Scientific Discovery. London, UK: Hutchinson & Co., 1959. doi: 10.2307/2412687
-
[10]
The replicability crisis and public trust in psychological sci- ence,
F. Anvari and D. Lakens, “The replicability crisis and public trust in psychological sci- ence,”Comprehensive Results in Social Psychology, vol. 3, no. 3, pp. 266–286, 2018. doi: 10.1080/23743603.2019.1684822
-
[11]
An open investi- gation of the reproducibility of cancer biology research,
T. M. Errington, E. Iorns, W. Gunn, F. E. Tan, J. Lomax, and B. A. Nosek, “An open investi- gation of the reproducibility of cancer biology research,”Elife, vol. 3, p. e04333, 2014. doi: 10.7554/eLife.04333
-
[12]
The reproducibility crisis in the age of digital medicine,
A. Stupple, D. Singerman, and L. A. Celi, “The reproducibility crisis in the age of digital medicine,”NPJ digital medicine, vol. 2, no. 1, p. 2, 2019. doi: 10.1038/s41746-019-0079-z
-
[13]
No raw data, no science: another possible source of the reproducibility crisis,
T. Miyakawa, “No raw data, no science: another possible source of the reproducibility crisis,” p. 24, 2020. doi: 10.1186/s13041-020-0552-2
-
[14]
Repro- ducibility in management science,
M. Fišar, B. Greiner, C. Huber, E. Katok, A. I. Ozkes, and M. S. R. Collaboration, “Repro- ducibility in management science,”Management Science, vol. 70, no. 3, pp. 1343–1356, 2024. doi: 10.1287/mnsc.2023.03556
-
[15]
Investigating the replicability of the social and behavioural sciences,
A. H. Tyner, A. L. Abatayo, M. Daley, S. Field, N. Fox, N. A. Haber, K. M. Hahn, M. K. Struhl, B. Mawhinney, O. Miskeet al., “Investigating the replicability of the social and behavioural sciences,”Nature, vol. 652, no. 8108, pp. 143–150, 2026. doi: 10.1038/s41586-025-10078-y
-
[16]
Artificial intelligence faces reproducibility crisis
M. Hutson, “Artificial intelligence faces reproducibility crisis,” 2018. doi: 10.1126/science.359.6377.725
-
[17]
Revisiting reproducibility in transportation simulation studies,
K. Riehl, A. Kouvelas, and M. A. Makridis, “Revisiting reproducibility in transportation simulation studies,”European Transport Research Review, vol. 17, no. 1, p. 22, 2025. doi: 10.1186/s12544-025-00718-9 . 12
-
[18]
M. Baker, “Reproducibility crisis,”nature, vol. 533, no. 26, pp. 353–66, 2016. doi: 10.1038/533437a
-
[19]
Is science really facing a reproducibility crisis, and do we need it to?
D. Fanelli, “Is science really facing a reproducibility crisis, and do we need it to?”Pro- ceedings of the National Academy of Sciences, vol. 115, no. 11, pp. 2628–2631, 2018. doi: 10.1073/pnas.1708272114
-
[20]
Before reproducibility must come preproducibility
P. B. Stark, “Before reproducibility must come preproducibility.”Nature, vol. 557, no. 7706, pp. 613–614, 2018. doi: 10.1038/d41586-018-05256-0
-
[21]
N. A. o. S. NAS,Reproducibility and replicability in science. National Academies Press, 2019. doi: 10.17226/25303
-
[22]
Agentic ai: Autonomous intelligence for com- plex goals—a comprehensive survey,
D. B. Acharya, K. Kuppan, and B. Divya, “Agentic ai: Autonomous intelligence for com- plex goals—a comprehensive survey,”IEEE Access, vol. 13, pp. 18 912–18 936, 2025. doi: 10.1109/ACCESS.2025.3532853
-
[23]
From Generation to Judgment: Opportunities and Challenges of LLM -as-a-judge
D. Li, B. Jiang, L. Huang, A. Beigi, C. Zhao, Z. Tan, A. Bhattacharjee, Y . Jiang, C. Chen, T. Wuet al., “From generation to judgment: Opportunities and challenges of llm-as-a-judge,” in Proceedings of the 2025 Conference on Empirical Methods in Natural Language Processing, 2025, pp. 2757–2791. doi: 10.18653/v1/2025.emnlp-main.138
-
[24]
Agentreview: Exploring peer review dynamics with llm agents,
Y . Jin, Q. Zhao, Y . Wang, H. Chen, K. Zhu, Y . Xiao, and J. Wang, “Agentreview: Exploring peer review dynamics with llm agents,” inProceedings of the 2024 Conference on Empirical Methods in Natural Language Processing, 2024, pp. 1208–1226. doi: 10.18653/v1/2024.emnlp-main.70
-
[25]
Can llm feedback enhance review quality? a randomized study of 20k reviews at iclr 2025,
N. Thakkar, M. Yuksekgonul, J. Silberg, A. Garg, N. Peng, F. Sha, R. Yu, C. V ondrick, and J. Zou, “Can llm feedback enhance review quality? a randomized study of 20k reviews at iclr 2025,”arXiv preprint arXiv:2504.09737, 2025. doi: 10.48550/arXiv.2504.09737
-
[26]
W. Liang, Y . Zhang, H. Cao, B. Wang, D. Y . Ding, X. Yang, K. V odrahalli, S. He, D. S. Smith, Y . Yinet al., “Can large language models provide useful feedback on research pa- pers? a large-scale empirical analysis,”NEJM AI, vol. 1, no. 8, p. AIoa2400196, 2024. doi: 10.1056/AIoa2400196
-
[27]
Llms as meta-reviewers’ assistants: A case study,
E. Hossain, S. K. Sinha, N. Bansal, R. A. Knipper, S. Sarkar, J. Salvador, Y . Mahajan, S. R. P. K. Guttikonda, M. Akter, M. M. Hassanet al., “Llms as meta-reviewers’ assistants: A case study,” inProceedings of the 2025 Conference of the Nations of the Americas Chapter of the Association for Computational Linguistics: Human Language Technologies (Volume 1...
-
[28]
Large language models for automated scholarly paper review: A survey,
Z. Zhuang, J. Chen, H. Xu, Y . Jiang, and J. Lin, “Large language models for automated scholarly paper review: A survey,”Information Fusion, vol. 124, p. 103332, 2025. doi: 10.1016/j.inffus.2025.103332
-
[29]
Repro-bench: Can agentic ai systems assess the reproducibility of social science research?
C. Hu, L. Zhang, Y . Lim, A. Wadhwani, A. Peters, and D. Kang, “Repro-bench: Can agentic ai systems assess the reproducibility of social science research?” inFindings of the Association for Computational Linguistics: ACL 2025, 2025, pp. 23 616–23 626. doi: 10.18653/v1/2025.findings-acl.1210
-
[30]
ReplicatorBench: Benchmarking LLM Agents for Replicability in Social and Behavioral Sciences
B. Nguyen, D. Soós, Q. Ma, R. R. Obadage, Z. Ranjan, S. Koneru, T. M. Errington, S. Ne- matova, S. Rajtmajer, J. Wuet al., “Replicatorbench: Benchmarking llm agents for repli- cability in social and behavioral sciences,”arXiv preprint arXiv:2602.11354, 2026. doi: 10.48550/arXiv.2602.11354
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2602.11354 2026
-
[31]
Miller, Tatiana Shavrina, Jakob N
A. Lupidi, B. Gauri, T. S. Foster, B. A. Omari, D. Magka, A. Pepe, A. Audran-Reiss, M. Aghamelu, N. Baldwin, L. Cipolina-Kun, J.-C. Gagnon-Audet, C. H. Leow, S. Lefdal, H. Mossalam, A. Moudgil, S. Nazir, E. Tewolde, I. Urrego, J. A. Estape, A. Budhiraja, G. Chaura- sia, A. Charnalia, D. Dunfield, K. Hambardzumyan, D. Izcovich, M. Josifoski, I. Mediratta, ...
-
[32]
From reproduction to replication: Evaluating research agents with progressive code masking,
G. J. Kim, A. Wilf, L.-P. Morency, and D. Fried, “From reproduction to replication: Evaluating research agents with progressive code masking,”arXiv preprint arXiv:2506.19724, 2025. doi: 10.48550/arXiv.2506.19724
-
[33]
M. Seo, J. Baek, S. Lee, and S. J. Hwang, “Paper2code: Automating code generation from scientific papers in machine learning,”arXiv preprint arXiv:2504.17192, 2025. doi: 10.48550/arXiv.2504.17192
-
[34]
Rescience c: a journal for reproducible replications in computa- tional science,
N. P. Rougier and K. Hinsen, “Rescience c: a journal for reproducible replications in computa- tional science,” inInternational Workshop on Reproducible Research in Pattern Recognition. Springer, 2018, pp. 150–156. doi: 10.1007/978-3-030-23987-9_14
-
[35]
Assessing data availability and research reproducibility in hydrology and water resources,
J. H. Stagge, D. E. Rosenberg, A. M. Abdallah, H. Akbar, N. A. Attallah, and R. James, “Assessing data availability and research reproducibility in hydrology and water resources,” Scientific data, vol. 6, no. 1, p. 190030, 2019. doi: 10.1038/sdata.2019.30
-
[36]
Reliability: on the reproducibility of assessment data,
S. M. Downing, “Reliability: on the reproducibility of assessment data,”Medical education, vol. 38, no. 9, pp. 1006–1012, 2004. doi: 10.1111/j.1365-2929.2004.01932.x
-
[37]
A. Bazzocchi, G. Filonzi, F. Ponti, C. Sassi, E. Salizzoni, G. Battista, and R. Canini, “Accuracy, reproducibility and repeatability of ultrasonography in the assessment of abdominal adiposity,” Academic radiology, vol. 18, no. 9, pp. 1133–1143, 2011. doi: 10.1016/j.acra.2011.04.014
-
[38]
A. L. Crowley, E. Yow, H. X. Barnhart, M. A. Daubert, R. Bigelow, D. C. Sullivan, M. Pencina, and P. S. Douglas, “Critical review of current approaches for echocardiographic reproducibility and reliability assessment in clinical research,”Journal of the American Society of Echocardiog- raphy, vol. 29, no. 12, pp. 1144–1154, 2016. doi: 10.1016/j.echo.2016.08.006
-
[39]
A practical guide to assess the reproducibility of echocardiographic measurements,
K. V . Bunting, R. P. Steeds, K. Slater, J. K. Rogers, G. V . Gkoutos, and D. Kotecha, “A practical guide to assess the reproducibility of echocardiographic measurements,”Journal of the American Society of Echocardiography, vol. 32, no. 12, pp. 1505–1515, 2019. doi: 10.1016/j.echo.2019.08.015
-
[40]
Statistical methods for replicability assessment,
K. Hung and W. Fithian, “Statistical methods for replicability assessment,”The Annals of Applied Statistics, vol. 14, no. 3, pp. 1063–1087, 2020. doi: 10.1214/20-AOAS1336
-
[41]
The assessment of replicability using the sum of p-values,
L. Held, S. Pawel, and C. Micheloud, “The assessment of replicability using the sum of p-values,” Royal Society Open Science, vol. 11, no. 8, 2024. doi: 10.1098/rsos.240149
-
[42]
M. Arroyo-Araujo, B. V oelkl, C. Laloux, J. Novak, B. Koopmans, A.-M. Waldron, I. Seiffert, H. Stirling, K. Aulehner, S. K. Janhunenet al., “Systematic assessment of the replicability and generalizability of preclinical findings: Impact of protocol harmonization across laboratory sites,”PLoS biology, vol. 20, no. 11, p. e3001886, 2022. doi: 10.1371/journa...
-
[43]
https: //arxiv.org/abs/2512.07921
Z. Li, Z. Li, Z. Guo, X. Ren, and C. Huang, “Deepcode: Open agentic coding,”arXiv preprint arXiv:2512.07921, 2025. doi: 10.48550/arXiv.2512.07921
-
[44]
C. Ye, S. Yuan, S. Cooray, S. Dillmann, I. L. Roque, D. Baron, P. Frank, S. Martin-Alvarez, N. Koblischke, F. J. Quet al., “Replicationbench: Can ai agents replicate astrophysics research papers?”arXiv preprint arXiv:2510.24591, 2025. doi: 10.48550/arXiv.2510.24591
-
[45]
Paperbench: Evaluating ai’s ability to replicate ai research, 2025
G. Starace, O. Jaffe, D. Sherburn, J. Aung, J. S. Chan, L. Maksin, R. Dias, E. Mays, B. Kinsella, W. Thompsonet al., “Paperbench: Evaluating ai’s ability to replicate ai research,”arXiv preprint arXiv:2504.01848, 2025. doi: 10.48550/arXiv.2504.01848
-
[46]
Llm-assisted repli- cation for quantitative social science,
S. Kubota, H. Yakura, S. Coavoux, S. Yamada, and Y . Nakamura, “Llm-assisted repli- cation for quantitative social science,”arXiv preprint arXiv:2602.18453, 2026. doi: 10.48550/arXiv.2602.18453
-
[47]
Llm-assisted replication as scientific infrastructure,
S. Kubota, H. Yakura, S. Yamada, Y . Nakamura, T. Werner, and S. Coavoux, “Llm-assisted replication as scientific infrastructure,”Open Science Framework, 2026
2026
-
[48]
Ai-driven review systems: evaluating llms in scalable and bias-aware academic reviews,
K. Tyser, B. Segev, G. Longhitano, X.-Y . Zhang, Z. Meeks, J. Lee, U. Garg, N. Belsten, A. Sh- porer, M. Udellet al., “Ai-driven review systems: evaluating llms in scalable and bias-aware academic reviews,”arXiv preprint arXiv:2408.10365, 2024. doi: 10.48550/arXiv.2408.10365 . 14
-
[49]
From replication to redesign: Exploring pairwise comparisons for llm-based peer review,
Y . Zhang, H. Zhang, W. Ji, T. Hua, N. Haber, H. Cao, and W. Liang, “From replication to redesign: Exploring pairwise comparisons for llm-based peer review,”arXiv preprint arXiv:2506.11343, 2025. doi: 10.48550/arXiv.2506.11343
-
[50]
Ai is transforming peer review—and many scientists are worried,
M. Naddaf, “Ai is transforming peer review—and many scientists are worried,”Nature, vol. 639, no. 8056, pp. 852–854, 2025. doi: 10.1038/d41586-025-00894-7
-
[51]
More than half of researchers now use ai for peer review—often against guidance,
——, “More than half of researchers now use ai for peer review—often against guidance,” Nature, vol. 649, no. 8096, pp. 273–274, 2026. doi: 10.1038/d41586-025-04066-5
-
[52]
A. Bhaskar and V . Stodden, “Reproscreener: Leveraging llms for assessing computational reproducibility of machine learning pipelines,” inProceedings of the 2nd ACM Conference on Reproducibility and Replicability, 2024, pp. 101–109. doi: 10.1145/3641525.3663629
-
[53]
Paper-snitch: A practical tool for evidence-based reproducibility assessment,
D. Santoli and F. Bolelli, “Paper-snitch: A practical tool for evidence-based reproducibility assessment,” Master’s thesis, University of Modena and Reggio Emilia, 2024. [Online]. Available: https://federicobolelli.it/media/supervision_pdfs/LM_Davide_Santoli.pdf
2024
-
[54]
F. Da Ros, T. Za ˇciragi´c, A. Plaat, T. Bäck, and N. van Stein, “Assessing reproducibility in evolutionary computation: A case study using human-and llm-based assessment,”arXiv preprint arXiv:2602.07059, 2026. doi: 10.48550/arXiv.2602.07059
-
[55]
Auto-metrics: Llm-assisted scientific quality control for radiomics research,
J. G. de Almeida and N. Papanikolaou, “Auto-metrics: Llm-assisted scientific quality control for radiomics research,”European Journal of Radiology, p. 112358, 2025. doi: 10.1016/j.ejrad.2025.112358
-
[56]
Mass reproducibility and replicability: A new hope,
A. Brodeur, D. Mikola, and N. Cook, “Mass reproducibility and replicability: A new hope,” JSTOR, Tech. Rep., 2024. [Online]. Available: https://www.jstor.org/stable/pdf/resrep58994. pdf?acceptTC=true&coverpage=false&addFooter=false
2024
-
[57]
State of the art: Reproducibility in artificial intelligence
O. E. Gundersen and S. Kjensmo, “State of the art: Reproducibility in artificial intelligence,” inProceedings of the AAAI conference on artificial intelligence, vol. 32, no. 1, 2018. doi: 10.1609/aaai.v32i1.11503
-
[58]
I4r discussion paper series, the institute for replication (i4r),
A. Brodeur, “I4r discussion paper series, the institute for replication (i4r),” Institute for Replica- tion (I4R), Tech. Rep., 2024
2024
-
[59]
Retraction watch – tracking retractions as a window into the scientific process,
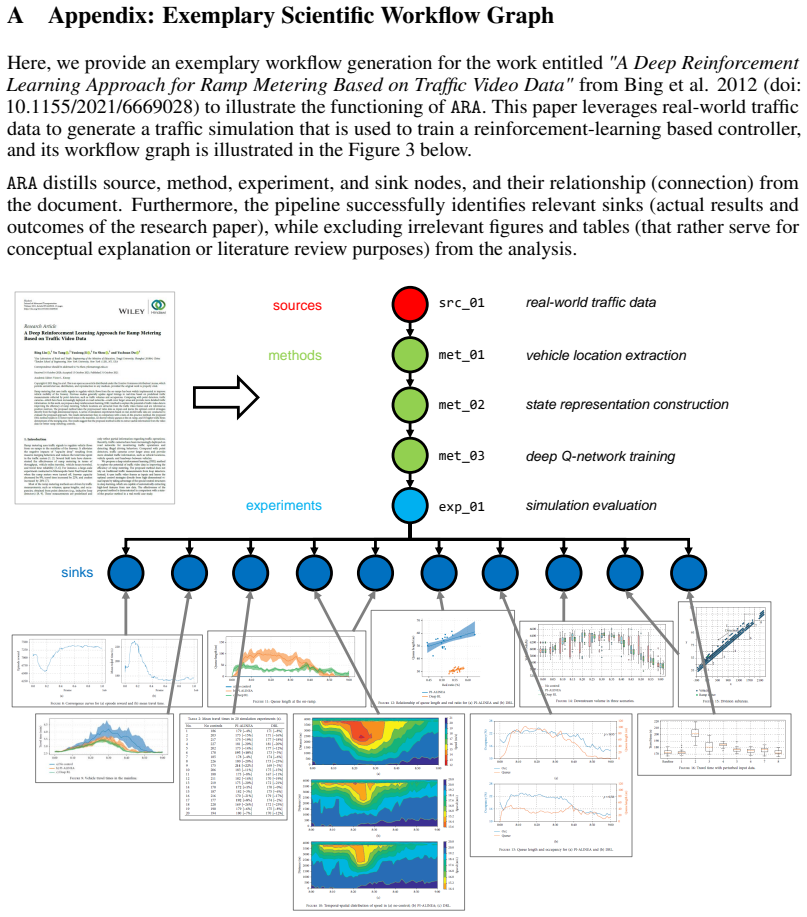
A. Marcus and I. Oransky, “Retraction watch – tracking retractions as a window into the scientific process,” Retraction Watch, Tech. Rep., 2024. [Online]. Available: https://retractionwatch.com/ 15 A Appendix: Exemplary Scientific Workflow Graph Here, we provide an exemplary workflow generation for the work entitled"A Deep Reinforcement Learning Approach ...
-
[60]
Guidelines: • The answer [N/A] means that the paper does not involve crowdsourcing nor research with human subjects
Institutional review board (IRB) approvals or equivalent for research with human subjects 33 Question: Does the paper describe potential risks incurred by study participants, whether such risks were disclosed to the subjects, and whether Institutional Review Board (IRB) approvals (or an equivalent approval/review based on the requirements of your country ...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.