Recognition: 2 theorem links
· Lean TheoremFast Voxelwise SNR Estimation for Iterative MRI Reconstructions
Pith reviewed 2026-05-12 03:45 UTC · model grok-4.3
The pith
PICO recovers voxelwise noise variance in general MRI reconstructions by probing the image-domain covariance operator with random-phase vectors.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
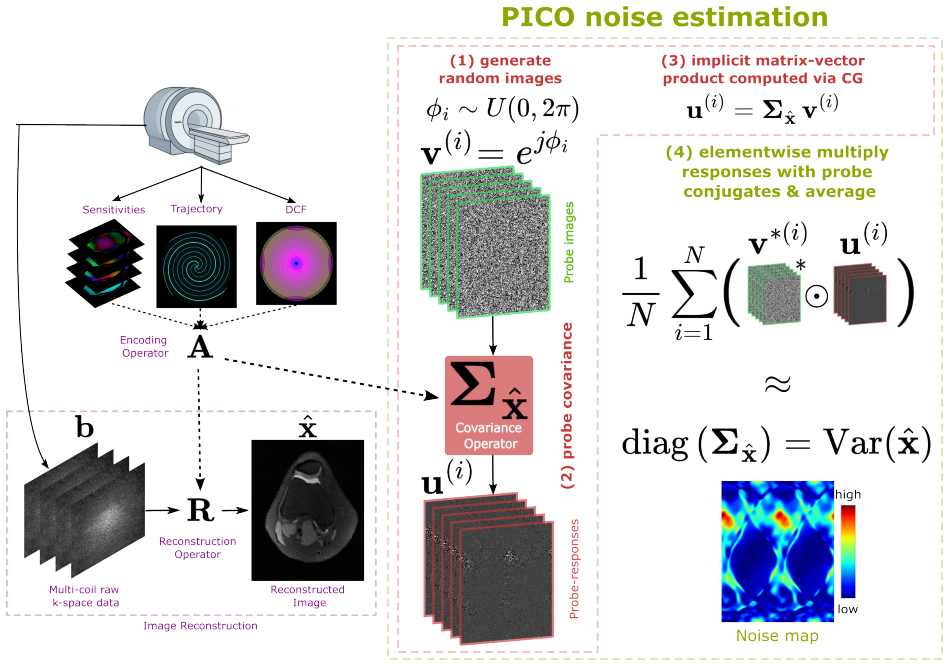
PICO estimates the image-domain noise variance by probing the noise covariance operator with complex random-phase vectors. These probes are shown to minimize estimator variance compared to Gaussian or real-valued alternatives. For nonlinear reconstructions, the Jacobian of the converged solution is probed instead. This yields accurate voxelwise SNR, g-factor, and related metrics without requiring closed-form expressions or large numbers of replica images.
What carries the argument
The probing of the image-domain noise covariance operator (or Jacobian) using complex random-phase probe images, which reuses existing reconstruction primitives to compute the variance estimates efficiently.
If this is right
- In Cartesian SENSE reconstructions, PICO reproduces analytical g-factor maps accurately.
- In non-Cartesian spiral imaging at R=2, it achieves 1% error in 64 seconds versus 462 seconds for PMR.
- For compressed-sensing knee reconstructions, Jacobian probing produces consistent noise maps faster than PMR.
- The method works across linear and nonlinear iterative reconstructions without additional calibration.
Where Pith is reading between the lines
- Integrating PICO into clinical reconstruction pipelines could make quantitative image quality assessment routine without extra scan time.
- Similar probing strategies might apply to noise estimation in other iterative reconstruction problems outside MRI, such as CT or ultrasound.
- Future work could explore optimal probe count or adaptive probing for even lower variance estimates.
Load-bearing premise
The image-domain noise covariance can be sufficiently well approximated by probing it with only a small number of random complex-phase vectors to give unbiased variance estimates.
What would settle it
A direct comparison on a new MRI dataset where PICO noise maps differ significantly from high-replica PMR references or analytical ground truth would show the estimator is inaccurate.
Figures






read the original abstract
Purpose: To develop a fast, general-purpose framework for voxelwise noise characterization in linear and nonlinear iterative MRI reconstructions, recovering the image-domain noise variance from which SNR, $g$-factor, and related image-quality metrics are derived. The framework addresses both the intractability of closed-form formulas beyond Cartesian sampling and the long runtime of Pseudo Multiple Replica (PMR) methods. Methods: We propose PICO (Probing Image-space COvariance), an estimator that operates in the image domain by probing the image-domain noise covariance operator -- or, for nonlinear compressed-sensing reconstructions, the Jacobian of the converged solution -- with random probe images. Complex random-phase probes are shown theoretically and empirically to minimize estimator variance compared with Gaussian or real-valued alternatives. PICO was validated against analytical benchmarks and high-replica PMR references using retrospective Cartesian knee data ($R=2$), prospective non-Cartesian spiral brain phantom data ($R=2,3,4$), and compressed-sensing knee reconstructions ($R=2$). Results: In Cartesian experiments, PICO accurately reproduced analytical SENSE $g$-factor maps. In non-Cartesian spiral imaging ($R=2$), it achieved 1% estimation error in 64 s compared with 462 s for PMR (approximately 7.2x speedup), with the efficiency advantage persisting at higher acceleration. For nonlinear compressed sensing, the Jacobian-based estimator produced noise maps consistent with PMR while converging faster (52 s vs. 95 s; approximately 1.8x speedup). Conclusion: PICO provides a computationally efficient alternative to PMR for voxelwise noise and $g$-factor estimation across generalized iterative MRI reconstructions. By reusing existing reconstruction primitives, it enables voxelwise noise maps to be produced as a routine by-product of the reconstruction pipeline.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proposes PICO (Probing Image-space COvariance), a framework for fast voxelwise noise variance estimation in linear and nonlinear iterative MRI reconstructions. It recovers per-voxel noise statistics by applying a small number of complex random-phase probes to the image-domain noise covariance operator (or its Jacobian at convergence for compressed-sensing cases), derives SNR and g-factor maps from the resulting variance estimates, and reports substantial speedups over Pseudo Multiple Replica (PMR) while matching analytical SENSE references on Cartesian data and PMR on non-Cartesian spiral and CS knee data.
Significance. If the finite-probe estimator is unbiased and its variance remains low enough to preserve the reported accuracy, PICO would enable routine voxelwise noise mapping as a low-overhead byproduct of generalized iterative reconstructions, addressing a practical bottleneck in quantitative MRI. The explicit validation against both analytical g-factor maps and high-replica PMR on multiple sampling schemes, together with the reuse of existing reconstruction operators, strengthens its potential utility.
major comments (2)
- [Methods, PICO Estimator] Methods (PICO estimator derivation): the unbiasedness of the diagonal covariance estimate obtained from a modest number of complex random-phase probes is asserted on the basis of covariance-operator properties, yet the explicit expectation calculation showing E[probe estimate] equals the true per-voxel variance for finite probe count (rather than only asymptotically) is not provided; without it the 1 % error figures in the spiral experiments cannot be guaranteed independent of probe count.
- [Results, non-Cartesian and CS experiments] Results, non-Cartesian and CS sections: the reported speedups (7.2× for R=2 spiral, 1.8× for CS) and error levels presuppose that estimator variance with the chosen probe count stays below the threshold that would negate the advantage over PMR; no separate analysis or table quantifies the Monte-Carlo variance of the PICO estimator itself across repeated probe realizations.
minor comments (2)
- [Abstract and Methods] The abstract states that complex phases are 'theoretically and empirically optimal' for variance reduction; the corresponding variance formula or comparison table should be referenced in the main text for readers who do not consult the supplement.
- [Figures] Figure captions for the g-factor and noise maps should explicitly state the number of probes used in each PICO reconstruction so that the timing and accuracy numbers can be reproduced.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed review of our manuscript on the PICO framework. We address each major comment point by point below and have revised the manuscript to incorporate the requested clarifications.
read point-by-point responses
-
Referee: [Methods, PICO Estimator] Methods (PICO estimator derivation): the unbiasedness of the diagonal covariance estimate obtained from a modest number of complex random-phase probes is asserted on the basis of covariance-operator properties, yet the explicit expectation calculation showing E[probe estimate] equals the true per-voxel variance for finite probe count (rather than only asymptotically) is not provided; without it the 1 % error figures in the spiral experiments cannot be guaranteed independent of probe count.
Authors: We thank the referee for this observation. The manuscript asserts unbiasedness from the fact that complex random-phase probes p satisfy E[p p^H] = I exactly (due to uniform phase distribution over the unit circle), so that the element-wise estimator for the diagonal of the image-domain covariance operator C has expectation exactly equal to diag(C) for any finite N. The average over N probes is therefore an unbiased estimator, with variance decreasing as 1/N. We agree that an explicit derivation would strengthen the presentation and will add it to the Methods section in the revised manuscript, showing E[(C p) ⊙ conj(p)] = diag(C) step by step. This confirms that the reported 1% errors hold for the modest probe counts used and are not reliant on asymptotic arguments. revision: yes
-
Referee: [Results, non-Cartesian and CS experiments] Results, non-Cartesian and CS sections: the reported speedups (7.2× for R=2 spiral, 1.8× for CS) and error levels presuppose that estimator variance with the chosen probe count stays below the threshold that would negate the advantage over PMR; no separate analysis or table quantifies the Monte-Carlo variance of the PICO estimator itself across repeated probe realizations.
Authors: We acknowledge that the manuscript does not provide a dedicated quantification of the Monte-Carlo variance of the PICO estimator across independent probe realizations. The 1% error values reflect agreement with high-replica PMR references (whose own variance is negligible), and the speedups are wall-clock comparisons at matched accuracy. To address the referee's concern directly, we will add a brief supplementary analysis (new table or paragraph in the revised Results) reporting the standard deviation of PICO estimates over 20 independent probe realizations for the spiral and CS cases. This will show that estimator variance remains low enough (relative standard deviation well below 1%) to preserve the accuracy and speedup claims. revision: yes
Circularity Check
No circularity; derivation from standard random probing of covariance operators
full rationale
The PICO estimator is constructed by applying random complex-phase vectors to the image-domain noise covariance operator (or Jacobian at convergence), with variance estimates obtained via the standard expectation property E[|probe^H * op * probe|] for the diagonal. This follows directly from linearity of expectation and properties of covariance operators without any fitted parameters, self-referential definitions, or load-bearing self-citations that reduce the result to its inputs. Validation against independent analytical SENSE g-factor maps and high-replica PMR references further confirms the chain is externally grounded rather than tautological. No equations in the provided description equate outputs to inputs by construction, and complex-phase optimality is shown via variance minimization rather than ansatz smuggling or renaming.
Axiom & Free-Parameter Ledger
free parameters (1)
- number of probes
axioms (2)
- domain assumption The reconstruction operator is differentiable or linear so that the Jacobian or covariance operator exists and can be probed.
- domain assumption Complex random-phase probes minimize estimator variance relative to Gaussian or real-valued probes.
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclear?
unclearRelation between the paper passage and the cited Recognition theorem.
Complex random-phase probes are shown theoretically and empirically to minimize estimator variance compared with Gaussian or real-valued alternatives... κ=1... unit-magnitude probes with uniformly distributed phase
-
IndisputableMonolith/Foundation/RealityFromDistinction.leanreality_from_one_distinction unclear?
unclearRelation between the paper passage and the cited Recognition theorem.
PICO... probing the image-domain noise covariance operator... Jacobian of the converged solution
What do these tags mean?
- matches
- The paper's claim is directly supported by a theorem in the formal canon.
- supports
- The theorem supports part of the paper's argument, but the paper may add assumptions or extra steps.
- extends
- The paper goes beyond the formal theorem; the theorem is a base layer rather than the whole result.
- uses
- The paper appears to rely on the theorem as machinery.
- contradicts
- The paper's claim conflicts with a theorem or certificate in the canon.
- unclear
- Pith found a possible connection, but the passage is too broad, indirect, or ambiguous to say the theorem truly supports the claim.
Reference graph
Works this paper leans on
-
[1]
Noise in mri.Magnetic Resonance in Medicine, 36(3):494–497, 1996
Albert Macovski. Noise in mri.Magnetic Resonance in Medicine, 36(3):494–497, 1996. doi: https://doi.org/10.1002/mrm.1910360327. URL https://onlinelibrary.wiley.com/doi/ abs/10.1002/mrm.1910360327
-
[3]
Olaf Dietrich, José G. Raya, Scott B. Reeder, Michael Ingrisch, Maximilian F. Reiser, and Stefan O. Schoenberg. Influence of multichannel combination, parallel imaging and other reconstruction techniques on mri noise characteristics.Magnetic Resonance Imaging, 26 (6):754–762, 2008. ISSN 0730-725X. doi: https://doi.org/10.1016/j.mri.2008.02.001. URL https:...
-
[4]
Olaf Dietrich, José G. Raya, Scott B. Reeder, Maximilian F. Reiser, and Stefan O. Schoenberg. Measurement of signal-to-noise ratios in mr images: Influence of multichannel coils, parallel imaging, and reconstruction filters.Journal of Magnetic Resonance Imaging, 26:375–385, 8
- [5]
-
[6]
Peter Kellman and Elliot R. McVeigh. Image reconstruction in snr units: A general method for snr measurement.Magnetic Resonance in Medicine, 54:1439–1447, 2005. ISSN 07403194. doi: 10.1002/mrm.20713
-
[7]
James D. Rubenstein, John M. Brown, John C. Kohn, and Steven P. Arnoczky. Cartilage invasion by fat: A possible mechanism of steroid-induced osteonecrosis?American Journal 18 of Roentgenology, 169(6):1439–1441, 1997. doi: 10.2214/ajr.169.6.9393162. URL https: //www.ajronline.org/doi/10.2214/ajr.169.6.9393162
-
[8]
R. A. Lerski and J. D. de Certaines. Performance assessment and quality control in mri by eurospin test objects and protocols.Magnetic Resonance Imaging, 11(6):817–833, 1993. doi: 10.1016/0730-725X(93)90199-N. URLhttps://pubmed.ncbi.nlm.nih.gov/8371637/
-
[9]
Deep learning: A primer for radiologists.RadioGraphics, 43(1):e1–e15, 2023
Shingo Kiryu, Hiroyuki Abe, Yusuke Hara, Takayuki Shimizu, Yoshitaka Narita, and Hiroshi Fujita. Deep learning: A primer for radiologists.RadioGraphics, 43(1):e1–e15, 2023. doi: 10.1148/rg.211719. URLhttps://pubs.rsna.org/doi/10.1148/rg.211719
-
[11]
Alexander R. Toews, Philip K. Lee, Krishna S. Nayak, and Brian A. Hargreaves. Comprehen- sive assessment of nonuniform image quality: Application to imaging near metal.Magnetic Resonance in Medicine, 92(6):2358–2372, 2024. doi: https://doi.org/10.1002/mrm.30222. URL https://onlinelibrary.wiley.com/doi/abs/10.1002/mrm.30222
-
[13]
Daniel K. Sodickson, Mark A. Griswold, Peter M. Jakob, Robert R. Edelman, and Warren J. Manning. Signal-to-noise ratio and signal-to-noise efficiency in smash imaging.Magnetic Reso- nance in Medicine, 41:1009–1022, 1999. ISSN 07403194. doi: 10.1002/(SICI)1522-2594(199905) 41:5<1009::AID-MRM21>3.0.CO;2-4
-
[14]
Felix A. Breuer, Stephan A.R. Kannengiesser, Martin Blaimer, Nicole Seiberlich, Peter M. Jakob, and Mark A. Griswold. General formulation for quantitative g-factor calculation in GRAPPA reconstructions.Magnetic Resonance in Medicine, 62(3):739–746, 2009. doi: 10.1002/mrm.22066
-
[15]
Yakushiji, Ichiro Tani, Yasuo Nakajima, and Marc Van Cauteren
Yasuyuki Kurihara, Yoshiko K. Yakushiji, Ichiro Tani, Yasuo Nakajima, and Marc Van Cauteren. Coil sensitivity encoding in mr imaging.American Journal of Roentgenology, 178(5):1087–1091,
-
[16]
URL https://doi.org/10.2214/ajr.178.5.1781087
doi: 10.2214/ajr.178.5.1781087. URL https://doi.org/10.2214/ajr.178.5.1781087. PMID: 11959706
-
[17]
Frank L. Goerner and Geoffrey D. Clarke. Measuring signal-to-noise ratio in partially parallel imaging mri.Medical Physics, 38:5049–5057, 2011. ISSN 00942405. doi: 10.1118/1.3618730
-
[18]
Pruessmann, Markus Weiger, Peter Börnert, and Peter Boesiger
Klaas P. Pruessmann, Markus Weiger, Peter Börnert, and Peter Boesiger. Advances in sensitivity encoding with arbitrary k-space trajectories.Magnetic Resonance in Medicine, 46:638–651,
- [19]
-
[20]
Roland Bammer, Murat Aksoy, and Chunlei Liu. Augmented generalized sense reconstruction to correct for rigid body motion.Magnetic Resonance in Medicine, 57(1):90–102, 2007. doi: 19 https://doi.org/10.1002/mrm.21106. URL https://onlinelibrary.wiley.com/doi/abs/10. 1002/mrm.21106
-
[21]
Oliver Maier, Steven Hubert Baete, Alexander Fyrdahl, Kerstin Hammernik, Seb Harrevelt, Lars Kasper, Agah Karakuzu, Michael Loecher, Franz Patzig, Ye Tian, Ke Wang, Daniel Gallichan, Martin Uecker, and Florian Knoll. Cg-sense revisited: Results from the first ismrm reproducibility challenge.Magnetic Resonance in Medicine, 85(4):1821–1839, 2021. doi: https...
-
[22]
Oren N Jaspan, Roman Fleysher, and Michael L Lipton. Compressed sensing mri: a review of the clinical literature.British Journal of Radiology, 88(1056):20150487, 10 2015. ISSN 0007-1285. doi: 10.1259/bjr.20150487. URLhttps://doi.org/10.1259/bjr.20150487
-
[23]
Shreyas S. Vasanawala, Marcus T. Alley, Brian A. Hargreaves, Richard A. Barth, John M. Pauly, and Michael Lustig. Improved pediatric mr imaging with compressed sensing.Radiology, 256(2):607–616, 2010. doi: 10.1148/radiol.10091218. URLhttps://doi.org/10.1148/radiol. 10091218. PMID: 20529991
-
[24]
Kerstin Hammernik, Thomas Küstner, Burhaneddin Yaman, Zhengnan Huang, Daniel Rueckert, Florian Knoll, and Mehmet Akçakaya. Physics-driven deep learning for computational magnetic resonance imaging: Combining physics and machine learning for improved medical imaging. IEEE Signal Processing Magazine, 40(1):98–114, 2023. doi: 10.1109/MSP.2022.3215288
-
[25]
Reinhard Heckel, Mathews Jacob, Akshay Chaudhari, Or Perlman, and Efrat Shimron. Deep learning for accelerated and robust MRI reconstruction.Magnetic Resonance Materials in Physics, Biology and Medicine, 37(3):335–368, July 2024. doi: 10.1007/s10334-024-01173-8. URLhttps://doi.org/10.1007/s10334-024-01173-8
-
[27]
J.A. Fessler, Sangwoo Lee, V.T. Olafsson, H.R. Shi, and D.C. Noll. Toeplitz-based iterative im- age reconstruction for mri with correction for magnetic field inhomogeneity.IEEE Transactions on Signal Processing, 53(9):3393–3402, 2005. doi: 10.1109/TSP.2005.853152
-
[28]
Per Thunberg and Per Zetterberg. Noise distribution in sense- and grappa-reconstructed images: a computer simulation study.Magnetic Resonance Imaging, 25:1089–1094, 9 2007. ISSN 0730725X. doi: 10.1016/j.mri.2006.11.003
-
[29]
Eros Montin and Riccardo Lattanzi. Seeking a widely adoptable practical standard to estimate signal-to-noise ratio in magnetic resonance imaging for multiple-coil reconstructions.J. Magn. Reson. Imaging, 54(6):1952–1964, 2021. doi: 10.1002/jmri.27816
-
[31]
Noise-induced variability quantification in deep learning-based mri reconstructions
Onat Dalmaz, Arjun Divyang Desai, Akshay Chaudhari, and Brian Hargreaves. Noise-induced variability quantification in deep learning-based mri reconstructions. In2024 ISMRM and ISMRT Annual Meeting and Exhibition, Singapore, Singapore, May 2024. ISMRM. URLhttp: //echo.ismrm.org/abstracts/view/4d1e145f-9b3a-4949-9121-1f5d15f99c48. Program Number: 2785
work page 2024
-
[32]
Mehmet Akcakaya, Tamer A. Basha, Warren J. Manning, and Reza Nezafat. Efficient calculation of g-factors for cg-sense in high dimensions: noise amplification in random undersampling. Journal of Cardiovascular Magnetic Resonance, 16, 1 2014. ISSN 1532429X. doi: 10.1186/ 1532-429X-16-S1-W28
work page 2014
-
[33]
Kerr, Petter Dyverfeldt, Charles J
Pierre Daudé, Susan Schnell, Caroline Wu, Aurelien B. Kerr, Petter Dyverfeldt, Charles J. François, Michael Markl, Brian A. Hargreaves, and Krishna S. Nayak. Inline automatic quality control of 2D phase-contrast flow MR imaging for subject-specific scan time adaptation.Magn. Reson. Med., 92(2):908–922, 2024. doi: 10.1002/mrm.30083
-
[34]
Curtis N. Wiens, Shawn J. Kisch, Jacob D. Willig-Onwuachi, and Charles A. McKenzie. Computationally rapid method of estimating signal-to-noise ratio for phased array image reconstructions.Magn. Reson. Med., 66(4):1192–1197, 2011. doi: 10.1002/mrm.22893
-
[35]
Efficient noise calculation in deep learning-based MRI reconstructions
Onat Dalmaz, Arjun D Desai, Reinhard Heckel, Tolga Cukur, Akshay S Chaudhari, and Brian Hargreaves. Efficient noise calculation in deep learning-based MRI reconstructions. In Aarti Singh, Maryam Fazel, Daniel Hsu, Simon Lacoste-Julien, Felix Berkenkamp, Tegan Maharaj, Kiri Wagstaff, and Jerry Zhu, editors,Proceedings of the 42nd International Conference o...
work page 2025
-
[36]
P. Dawood, F. Breuer, M. Gram, I. Homolya, P. M. Jakob, M. Zaiss, and M. Blaimer. Image space formalism of convolutional neural networks for k-space interpolation.Magnetic Resonance in Medicine, 94(6):2680–2701, 2025. doi: https://doi.org/10.1002/mrm.70002. URL https://onlinelibrary.wiley.com/doi/abs/10.1002/mrm.70002
-
[37]
Istvan Homolya, Jannik Stebani, Felix Breuer, Grit Hein, Matthias Gamer, Florian Knoll, and Martin Blaimer. Controlling spatial correlation in k-space interpolation networks for MRI reconstruction: denoising versus apparent blurring.arXiv preprint arXiv:2505.11155, 2025. doi: 10.48550/arXiv.2505.11155
-
[38]
Michael F. Hutchinson. A stochastic estimator of the trace of the influence matrix for laplacian smoothing splines.J. Commun. Statist. Simula., 19(2):433–450, 1990. URLhttps://cir.nii. ac.jp/crid/1573668925763299072
-
[39]
C. Bekas, E. Kokiopoulou, and Y. Saad. An estimator for the diagonal of a matrix.Applied Numerical Mathematics, 57(11):1214–1229, 2007. ISSN 0168-9274. doi: https://doi.org/10. 1016/j.apnum.2007.01.003. URL https://www.sciencedirect.com/science/article/pii/ S0168927407000244. Numerical Algorithms, Parallelism and Applications (2)
work page 2007
-
[40]
Michael S. Hansen and Peter Kellman. Image reconstruction: an overview for clinicians.J. Magn. Reson. Imaging, 41(3):573–585, 2015. doi: 10.1002/jmri.24687
-
[41]
Kylie Yeung, Christine Tobler, Rolf F Schulte, Benjamin White, Anthony McIntyre, Sébastien Serres, Peter Morris, Dorothee Auer, Fergus V Gleeson, Damian J Tyler, et al. Algebraic 21 methods and computational strategies for pseudoinverse-based mr image reconstruction (pinv- recon).Scientific Reports, 15(1):37997, 2025
work page 2025
-
[42]
Michael Lustig, David L. Donoho, Juan M. Santos, and John M. Pauly. Compressed sensing mri.IEEE Signal Processing Magazine, 25(2):72–82, 2008. doi: 10.1109/MSP.2007.914728
-
[43]
Creation of fully sampled mr data repository for compressed sensing of the knee
Kevin Epperson, Anne Marie Sawyer, Michael Lustig, Marcus Alley, Martin Uecker, Patrick Virtue, Peng Lai, and Shreyas Vasanawala. Creation of fully sampled mr data repository for compressed sensing of the knee. ISMRM 2013 Meeting Proceedings. Section for Magnetic Resonance Technologists, 2013
work page 2013
-
[44]
Leslie Ying and Jinhua Sheng. Joint image reconstruction and sensitivity estimation in sense (jsense).Magnetic Resonance in Medicine, 57:1196–1202, 2007. ISSN 15222594. doi: 10.1002/mrm.21245
-
[45]
Murphy, Patrick Virtue, Michael Elad, John M
Martin Uecker, Peng Lai, Mark J. Murphy, Patrick Virtue, Michael Elad, John M. Pauly, Shreyas S. Vasanawala, and Michael Lustig. Espirit - an eigenvalue approach to autocalibrating parallel mri: Where sense meets grappa.Magnetic Resonance in Medicine, 71:990–1001, 2014. ISSN 15222594. doi: 10.1002/mrm.24751
-
[46]
Muckley, Mary Bruno, Aaron Defazio, Marc Parente, Krzysztof J
Florian Knoll, Jure Zbontar, Anuroop Sriram, Matthew J. Muckley, Mary Bruno, Aaron Defazio, Marc Parente, Krzysztof J. Geras, Joe Katsnelson, Hersh Chandarana, Zizhao Zhang, Michal Drozdzalv, Adriana Romero, Michael Rabbat, Pascal Vincent, James Pinkerton, Duo Wang, Nafissa Yakubova, Erich Owens, C. Lawrence Zitnick, Michael P. Recht, Daniel K. Sodickson,...
-
[47]
A fast iterative shrinkage-thresholding algorithm for linear inverse problems
AmirBeckandMarcTeboulle. Afastiterativeshrinkage-thresholdingalgorithmforlinearinverse problems.SIAM Journal on Imaging Sciences, 2(1):183–202, 2009. doi: 10.1137/080716542
-
[48]
Florian Knoll, Kristian Bredies, Thomas Pock, and Rudolf Stollberger. Second order total generalized variation (tgv) for mri.Magnetic Resonance in Medicine, 65(2):480–491, 2011. doi: https://doi.org/10.1002/mrm.22595. URL https://onlinelibrary.wiley.com/doi/abs/10. 1002/mrm.22595
-
[49]
Curran Associates Inc., Red Hook, NY, USA, 2019
Adam Paszke, Sam Gross, Francisco Massa, Adam Lerer, James Bradbury, Gregory Chanan, Trevor Killeen, Zeming Lin, Natalia Gimelshein, Luca Antiga, Alban Desmaison, Andreas Köpf, Edward Yang, Zach DeVito, Martin Raison, Alykhan Tejani, Sasank Chilamkurthy, Benoit Steiner, Lu Fang, Junjie Bai, and Soumith Chintala.PyTorch: an imperative style, high-performan...
work page 2019
-
[50]
Sigpy: A python package for high performance iterative reconstruction.Proc
Frank Ong and Michael Lustig. Sigpy: A python package for high performance iterative reconstruction.Proc. Intl. Soc. Mag. Reson. Med., 27:4819, 2019
work page 2019
-
[51]
R.W. Brown, Y.C.N. Cheng, E.M. Haacke, M.R. Thompson, and R. Venkatesan.Magnetic Resonance Imaging: Physical Principles and Sequence Design. Wiley, 2014. ISBN9781118633977. URLhttps://books.google.com/books?id=rQGCAwAAQBAJ
work page 2014
-
[52]
Wiley-Blackwell, 5th edition, 2018
Catherine Westbrook, Carolyn Kaut Roth, and John Talbot.MRI in Practice. Wiley-Blackwell, 5th edition, 2018. ISBN 978-1119391968. 22
work page 2018
-
[53]
Haim Avron and Sivan Toledo. Randomized algorithms for estimating the trace of an implicit symmetric positive semi-definite matrix.Journal of the ACM, 58(2):8, 2011. doi: 10.1145/ 1944345.1944349
-
[54]
Philip M. Robson, Aaron K. Grant, Ananth J. Madhuranthakam, Riccardo Lattanzi, Daniel K. Sodickson, and Charles A. McKenzie. Comprehensive quantification of signal-to-noise ratio and g-factor for image-based and k-space-based parallel imaging reconstructions.Magnetic Resonance in Medicine, 60(4):895–907, 2008. doi: 10.1002/mrm.21728
-
[55]
Sodickson, and Mehmet Akcakaya
Florian Knoll, Kerstin Hammernik, Chi Zhang, Steen Moeller, Thomas Pock, Daniel K. Sodickson, and Mehmet Akcakaya. Deep-learning methods for parallel magnetic resonance imaging reconstruction: a survey of the current approaches, trends, and issues.IEEE Signal Process. Mag., 37(1):128–140, 2020. doi: 10.1109/MSP.2019.2950640
-
[56]
Data-driven regularization parameter selection in dynamic MRI.J
Matti Hanhela, Mikael Delić, Aku Nissinen, and Ville Kolehmainen. Data-driven regularization parameter selection in dynamic MRI.J. Imaging, 7(2):38, 2021. doi: 10.3390/jimaging7020038
-
[57]
Mathieu Boudreau, Agah Karakuzu, Julien Cohen-Adad, Ecem Bozkurt, Madeline Carr, Marco Castellaro, Luis Concha, Mariya Doneva, Seraina A. Dual, Alex Ensworth, Alexandru Foias, Véronique Fortier, Refaat E. Gabr, Guillaume Gilbert, Carri K. Glide-Hurst, Matthew Grech-Sollars, Siyuan Hu, Oscar Jalnefjord, Jorge Jovicich, Kübra Keskin, Peter Koken, Anastasia ...
work page doi:10.1002/mrm 2024
-
[58]
Siria Pasini, Steffen Ringgaard, Tau Vendelboe, Leyre Garcia-Ruiz, Anika Strittmatter, Giulia Villa, Anish Raj, Rebeca Echeverria-Chasco, Michela Bozzetto, Paolo Brambilla, Malene Aastrup, Esben S. S. Hansen, Luisa Pierotti, Matteo Renzulli, Susan T. Francis, Frank G. Zöllner, Christoffer Laustsen, Maria A. Fernandez-Seara, and Anna Caroli. Multi-center a...
-
[59]
Eros Montin, Xuan Thao Nguyen, and Riccardo Lattanzi. MR Optimum: A web-based open- source tool for standardized signal-to-noise ratio evaluation in MRI.Computer Methods and Programs in Biomedicine Update, page 100235, 2026. doi: 10.1016/j.cmpbup.2026.100235
-
[60]
Jeremiah J. Hess, Catherine J. Moran, Preya Shah, Jana Vincent, Fraser J. L. Robb, Bruce L. Daniel, and Brian A. Hargreaves. Relative SNR measurements in supine vs. prone breast MRI. Magnetic Resonance in Medicine, 95(5):2718–2725, 2026. doi: 10.1002/mrm.70217
-
[61]
Gaël Varoquaux, Flore Baronnet, Andreas Kleinschmidt, Pierre Fillard, and Bertrand Thirion. Detection of brain functional-connectivity difference in post-stroke patients using group-level covariance modeling. In Tianzi Jiang, Nassir Navab, Josien P. W. Pluim, and Max A. Viergever, 23 editors,Medical Image Computing and Computer-Assisted Intervention – MIC...
work page 2010
-
[62]
Ali Behrouz, Parsa Delavari, and Farnoosh Hashemi. Unsupervised representation learning of brain activity via bridging voxel activity and functional connectivity. InProceedings of the 41st International Conference on Machine Learning (ICML), volume 235 ofProceedings of Machine Learning Research, pages 3347–3381, 2024
work page 2024
-
[63]
Brian Toner. Methods for uncertainty quantification in dictionary matching to advance interpretable quantitative MRI. Sedona, AZ, United States of America, January 2026. Proc. Intl. Soc. Mag. Reson. Med. Workshop on Data Sampling and Image Reconstruction. URL https://echo.ismrm.org/program/SEDONA26. Abstract #00187
work page 2026
-
[64]
Vandana V. Hanchate and Kalyani R. Joshi. Mri denoising using bm3d equipped with noise invalidation denoising technique and vst for improved contrast.SN Applied Sciences, 2(2):234, 2020
work page 2020
-
[65]
Zhoubo Li, Lifeng Yu, Joshua D. Trzasko, David S. Lake, Daniel J. Blezek, Joel G. Fletcher, Cynthia H. McCollough, and Armando Manduca. Adaptive nonlocal means filtering based on local noise level for ct denoising.Medical Physics, 41(1):011908, 2014. doi: 10.1118/1.4851635
-
[66]
Sai Gokul Hariharan, Norbert Strobel, Christian Kaethner, Markus Kowarschik, Rebecca Fahrig, and Nassir Navab. Data-driven estimation of noise variance stabilization parameters for low-dose x-ray images.Physics in Medicine and Biology, 65(22):225027, 2020. doi: 10.1088/ 1361-6560/abbc82
work page 2020
-
[67]
Yongjian Yu and Scott T. Acton. Speckle reducing anisotropic diffusion.IEEE Transactions on Image Processing, 11(11):1260–1270, 2002. doi: 10.1109/TIP.2002.804276
-
[68]
Quan Dou, Zhixing Wang, Xue Feng, Adrienne E. Campbell-Washburn, John P. Mugler, and Craig H. Meyer. Mri denoising with a non-blind deep complex-valued convolutional neural network.NMR in Biomedicine, 38(1):e5291, 2025. doi: 10.1002/nbm.5291
-
[69]
U-medsam: Uncertainty-aware medsam for medical image segmentation
Xin Wang, Xiaoyu Liu, Peng Huang, Pu Huang, Shu Hu, and Hongtu Zhu. U-medsam: Uncertainty-aware medsam for medical image segmentation. In Jun Ma, Yuyin Zhou, and Bo Wang, editors,Medical Image Segmentation Foundation Models. CVPR 2024 Challenge: Segment Anything in Medical Images on Laptop, pages 206–217, Cham, 2025. Springer Nature Switzerland. ISBN 978-...
work page 2024
-
[70]
Hui Xue, Sarah M. Hooper, Iain Pierce, Rhodri H. Davies, John Stairs, Joseph Naegele, Adrienne E. Campbell-Washburn, Charlotte Manisty, James C. Moon, Thomas A. Treibel, Michael S. Hansen, and Peter Kellman. SNRAware: Improved deep learning MRI denoising with signal-to-noise ratio unit training and g-factor map augmentation.Radiology: Artificial Intellige...
-
[71]
Hui Xue, Sarah Hooper, Azaan Rehman, Iain Pierce, Thomas Treibel, Rhodri Davies, W. Pa- tricia Bandettini, Rajiv Ramasawmy, Ahsan Javed, Zheren Zhu, Yang Yang, James Moon, Adrienne Campbell, and Peter Kellman. Imaging transformer for MRI denoising with the SNR unit training: enabling generalization across field-strengths, imaging contrasts, and anatomy,
- [72]
-
[73]
Pruessmann, Markus Weiger, Markus B
Klaas P. Pruessmann, Markus Weiger, Markus B. Scheidegger, and Peter Boesiger. Sense: Sensitivity encoding for fast mri.Magnetic Resonance in Medicine, 42(5):952–962, 1999. doi: 10.1002/(SICI)1522-2594(199911)42:5<952::AID-MRM16>3.0.CO;2-S
-
[74]
Harrison H. Barrett. Task-based measures of image quality and their relation to radiation dose and patient risk.Physics in Medicine and Biology, 60(2):R1–R75, 2015. doi: 10.1088/ 0031-9155/60/2/R1
work page 2015
- [75]
-
[76]
Rad-IQMRI: A benchmark for MRI image quality assessment.Neurocomputing, 602:128292, 2024
Yueran Ma, Jianxun Lou, Jean-Yves Tanguy, Padraig Corcoran, and Hantao Liu. Rad-IQMRI: A benchmark for MRI image quality assessment.Neurocomputing, 602:128292, 2024. doi: 10.1016/j.neucom.2024.128292
-
[77]
Eisenmenger, Leonardo Rivera-Rivera, Eugene Huo, Jacqueline C
Chenwei Tang, Laura B. Eisenmenger, Leonardo Rivera-Rivera, Eugene Huo, Jacqueline C. Junn, Anthony D. Kuner, Thekla H. Oechtering, Anthony Peret, Jitka Starekova, and Kevin M. Johnson. Incorporating radiologist knowledge into MRI quality metrics for machine learning using rank-based ratings.Journal of Magnetic Resonance Imaging, 61(6):2572–2584, 2025. do...
-
[78]
Cambridge Series in Statistical and Probabilistic Mathematics
Roman Vershynin.High-Dimensional Probability: An Introduction with Applications in Data Science. Cambridge Series in Statistical and Probabilistic Mathematics. Cambridge University Press, Cambridge, 2 edition, 2026. 25 Algorithm 1Probing Image-space COvariance (PICO) Require: Encoding operatorA(and adjointA H); regularization parameterλ≥0; number of probe...
work page 2026
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.