Recognition: unknown
VERITAS: Verifiable Epistemic Reasoning for Image-Derived Hypothesis Testing via Agentic Systems
Pith reviewed 2026-05-10 14:51 UTC · model grok-4.3
The pith
A four-phase multi-agent system tests natural-language hypotheses on MRI datasets with 81.4 percent verdict accuracy and fully auditable evidence trails.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The paper claims that structured multi-agent decomposition substitutes for model scale while preserving the verifiability clinical research demands. VERITAS reaches 81.4 percent verdict accuracy with frontier models and 71.2 percent with locally-hosted open-weight models (8-30B parameters), outperforming all five single-model baselines, and produces the highest rate of independently verifiable statistical outputs at 86.6 percent so that even failures remain diagnosable through artifact inspection.
What carries the argument
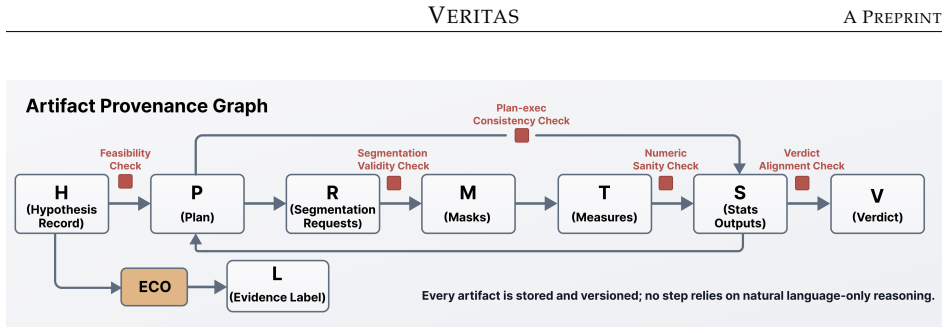
Four-phase agentic decomposition with role-specialized agents together with the epistemic evidence label framework that mechanically classifies outcomes as Supported, Refuted, Underpowered, or Invalid by jointly evaluating significance, effect direction, and study power.
If this is right
- Structured multi-agent systems can substitute for larger model scales in clinical reasoning tasks while maintaining auditability.
- The epistemic labeling distinguishes underpowered results from true absences of effect, which is common in medical imaging studies.
- Every statistical conclusion traces to inspectable executable outputs, enabling diagnosis of system failures through artifact review.
- Autonomous testing of natural-language hypotheses reduces the need to manually coordinate expertise across multiple domains.
- Performance advantages hold across both frontier and open-weight models on cardiac and glioma MRI data.
Where Pith is reading between the lines
- The same four-phase structure might transfer to hypothesis testing on genomic or electronic health record data with only modest adaptation.
- Adding optional human review at the planning phase could raise accuracy in regulated clinical environments without losing the audit trail.
- If the approach scales, it could increase the throughput of reproducible findings in radiology and neurology by lowering coordination costs.
- Future benchmarks on datasets with independently confirmed ground-truth outcomes would provide a stronger test of verdict reliability.
Load-bearing premise
The tiered benchmark of 64 hypotheses on two specific MRI datasets accurately represents real-world clinical hypothesis testing and the four-phase decomposition generalizes without human intervention.
What would settle it
Applying VERITAS unchanged to a new collection of 50 hypotheses drawn from a different clinical imaging modality such as CT or ultrasound and finding verdict accuracy below 65 percent or verifiable output rate below 75 percent would falsify the generalization claim.
Figures




read the original abstract
Drawing meaningful conclusions from inherently multimodal clinical data (including medical imaging) requires coordinating expertise across the clinical specialty, radiology, programming, and biostatistics. This fragmented process bottlenecks discovery. We present VERITAS (Verifiable Epistemic Reasoning for Image-Derived Hypothesis Testing via Agentic Systems), a multi-agent system that autonomously tests natural-language hypotheses on multimodal clinical datasets while producing a fully auditable evidence trail: every statistical conclusion traces through inspectable, executable outputs from analysis plan to segmentation masks to statistical code to final verdict. VERITAS decomposes the workflow into four phases handled by role-specialized agents, and introduces an epistemic evidence label framework that mechanically classifies outcomes as Supported, Refuted, Underpowered, or Invalid by jointly evaluating significance, effect direction, and study power. This distinction is critical in medical imaging, where non-significant results often reflect insufficient sample size rather than absent effects. To evaluate the system, we construct a tiered benchmark of 64 hypotheses spanning six complexity levels across cardiac (ACDC, 150 subjects) and brain glioma (UCSF-PDGM, 501 subjects) MRI. VERITAS reaches 81.4% verdict accuracy with frontier models and 71.2% with locally-hosted open-weight models (8-30B), outperforming all five single-model baselines in both classes. It also produces the highest rate of independently verifiable statistical outputs (86.6%), so even its failures remain diagnosable through artifact inspection. Structured multi-agent decomposition thus substitutes for model scale while preserving the verifiability clinical research demands.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper presents VERITAS, a multi-agent system that decomposes natural-language hypothesis testing on multimodal clinical MRI data into four specialized phases, producing auditable statistical outputs. It introduces an epistemic evidence label framework classifying results as Supported, Refuted, Underpowered, or Invalid. On a constructed tiered benchmark of 64 hypotheses spanning six complexity levels from the ACDC (150 subjects) and UCSF-PDGM (501 subjects) datasets, VERITAS reports 81.4% verdict accuracy with frontier models and 71.2% with open-weight models (8-30B), outperforming five single-model baselines, along with 86.6% independently verifiable outputs.
Significance. If the benchmark is representative of real clinical workflows, the work demonstrates that structured agentic decomposition can substitute for raw model scale while delivering the auditability required for medical imaging research. The epistemic labeling approach usefully distinguishes underpowered from null results, a common issue in the domain. The emphasis on executable evidence trails is a concrete strength that could support reproducible discovery pipelines.
major comments (3)
- [Section 4] Benchmark construction (Section 4): The paper provides insufficient detail on how the 64 hypotheses were selected or generated, including whether they were synthesized to align with the system's statistical criteria or drawn from independent clinical sources. This directly affects the validity of the reported accuracy figures and the claim that the four-phase decomposition generalizes.
- [Section 5] Evaluation protocol (Section 5): No information is given on the ground-truth verdict adjudication process, inter-rater reliability for labels, statistical power calculations for the benchmark itself, or controls for confounds such as hypothesis phrasing bias. These omissions make it impossible to assess whether the 81.4%/71.2% outperformance is robust or artifactual.
- [Section 3.2] Epistemic framework implementation (Section 3.2): The mechanical rules for jointly evaluating significance, effect direction, and study power are described at a high level but lack explicit formulas or pseudocode for power estimation and label assignment, hindering independent reproduction and verification of the 86.6% verifiable-output rate.
minor comments (2)
- [Table 2] Table 2: The baseline comparison table would benefit from explicit reporting of per-complexity-level breakdowns to support the claim that multi-agent decomposition helps across all six tiers.
- [Figure 1] Figure 1: The agent workflow diagram uses abbreviations (e.g., 'EEL') without a legend in the caption, reducing immediate clarity for readers unfamiliar with the epistemic label framework.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed feedback on our manuscript describing VERITAS. The comments identify important areas where additional transparency will strengthen the paper's reproducibility and interpretability. We address each major comment below and have revised the manuscript accordingly to incorporate the requested details.
read point-by-point responses
-
Referee: [Section 4] Benchmark construction (Section 4): The paper provides insufficient detail on how the 64 hypotheses were selected or generated, including whether they were synthesized to align with the system's statistical criteria or drawn from independent clinical sources. This directly affects the validity of the reported accuracy figures and the claim that the four-phase decomposition generalizes.
Authors: We agree that the original description of hypothesis generation lacked sufficient specificity. The 64 hypotheses were developed in collaboration with clinical experts to mirror realistic research questions drawn from the ACDC and UCSF-PDGM datasets, spanning six complexity tiers based on factors such as number of modalities, statistical operations required, and clinical relevance; they were not synthesized post hoc to match the system's output criteria. To address this, we have added a new subsection (4.1) and Appendix B that provides the complete list of hypotheses, the generation protocol (including input from independent clinicians), tiering rationale, and explicit confirmation that selection was independent of the VERITAS statistical pipeline. These additions directly support the generalizability claim. revision: yes
-
Referee: [Section 5] Evaluation protocol (Section 5): No information is given on the ground-truth verdict adjudication process, inter-rater reliability for labels, statistical power calculations for the benchmark itself, or controls for confounds such as hypothesis phrasing bias. These omissions make it impossible to assess whether the 81.4%/71.2% outperformance is robust or artifactual.
Authors: We acknowledge this as a valid criticism that limits assessment of robustness. Ground-truth verdicts were obtained via independent review by two board-certified radiologists and one biostatistician, with disagreements resolved through consensus discussion; inter-rater reliability was quantified using Fleiss' kappa. Post-hoc power calculations for the benchmark were performed using standard formulas for the observed effect sizes and sample sizes in each dataset. To mitigate phrasing bias, hypotheses were generated from standardized clinical templates. We have expanded Section 5 with a new subsection (5.1) detailing the full adjudication protocol, reliability metrics, power analysis, and bias controls, along with the raw agreement statistics. revision: yes
-
Referee: [Section 3.2] Epistemic framework implementation (Section 3.2): The mechanical rules for jointly evaluating significance, effect direction, and study power are described at a high level but lack explicit formulas or pseudocode for power estimation and label assignment, hindering independent reproduction and verification of the 86.6% verifiable-output rate.
Authors: We agree that the high-level description impedes reproduction. The label assignment follows a deterministic decision tree: (1) compute p-value via the appropriate test (t-test, chi-square, or regression as selected by the analysis agent); (2) check effect direction consistency against the hypothesis; (3) estimate power using the formula for the given test, sample size, and observed effect size (implemented via scipy.stats.power or equivalent); (4) assign Supported if significant and powered, Refuted if significant but opposite direction, Underpowered if non-significant but power < 0.8, or Invalid for other failures. We have inserted explicit formulas, a pseudocode listing of the full decision procedure, and threshold values (e.g., alpha=0.05, power threshold=0.8) into Section 3.2 plus a new Appendix C, enabling direct verification of the 86.6% rate. revision: yes
Circularity Check
No significant circularity detected; evaluation is empirical on external benchmarks.
full rationale
The paper presents an agentic system with a four-phase decomposition and an epistemic labeling framework (Supported/Refuted/Underpowered/Invalid) that classifies outcomes by significance, effect, and power. It evaluates this on a constructed tiered benchmark of 64 hypotheses using public external datasets (ACDC with 150 subjects, UCSF-PDGM with 501 subjects) and compares against five independent single-model baselines. Reported accuracies (81.4% frontier, 71.2% open-weight) and verifiable output rate (86.6%) are direct empirical measurements, not reductions of any claimed derivation or prediction to fitted inputs or self-definitions. No equations, self-citation load-bearing premises, uniqueness theorems, or ansatzes are invoked that would make the central performance claims tautological by construction. The chain is self-contained as a system description plus benchmark evaluation against external data.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Multi-agent systems can decompose complex multimodal analysis workflows into verifiable executable steps
- domain assumption Outcomes can be mechanically classified by jointly evaluating statistical significance, effect direction, and study power
invented entities (1)
-
Epistemic evidence label framework
no independent evidence
Reference graph
Works this paper leans on
-
[1]
Deep learning techniques for automatic mri cardiac multi-structures segmentation and diagnosis: is the problem solved?IEEE transactions on medical imaging, 37(11):2514–2525, 2018
Olivier Bernard, Alain Lalande, Clement Zotti, Frederick Cervenansky, Xin Yang, Pheng-Ann Heng, Irem Cetin, Karim Lekadir, Oscar Camara, Miguel Angel Gonzalez Ballester, et al. Deep learning techniques for automatic mri cardiac multi-structures segmentation and diagnosis: is the problem solved?IEEE transactions on medical imaging, 37(11):2514–2525, 2018
2018
-
[2]
The university of california san francisco preoperative diffuse glioma mri dataset.Radiology: Artificial Intelligence, 4(6):e220058, 2022
Evan Calabrese, Javier E Villanueva-Meyer, Jeffrey D Rudie, Andreas M Rauschecker, Ujjwal Baid, Spyridon Bakas, Soonmee Cha, John T Mongan, and Christopher P Hess. The university of california san francisco preoperative diffuse glioma mri dataset.Radiology: Artificial Intelligence, 4(6):e220058, 2022
2022
-
[3]
Segment anything
Alexander Kirillov, Eric Mintun, Nikhila Ravi, Hanzi Mao, Chloe Rolland, Laura Gustafson, Tete Xiao, Spencer Whitehead, Alexander C Berg, Wan-Yen Lo, et al. Segment anything. InProceedings of the IEEE/CVF international conference on computer vision, pages 4015–4026, 2023
2023
-
[4]
SAM 2: Segment Anything in Images and Videos
Nikhila Ravi, Valentin Gabeur, Yuan-Ting Hu, Ronghang Hu, Chaitanya Ryali, Tengyu Ma, Haitham Khedr, Roman Rädle, Chloe Rolland, Laura Gustafson, et al. Sam 2: Segment anything in images and videos.arXiv preprint arXiv:2408.00714, 2024
work page internal anchor Pith review Pith/arXiv arXiv 2024
-
[5]
SAM 3: Segment Anything with Concepts
Nicolas Carion, Laura Gustafson, Yuan-Ting Hu, Shoubhik Debnath, Ronghang Hu, Didac Suris, Chaitanya Ryali, Kalyan Vasudev Alwala, Haitham Khedr, Andrew Huang, et al. Sam 3: Segment anything with concepts.arXiv preprint arXiv:2511.16719, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[6]
Segment anything in medical images
Jun Ma, Yuting He, Feifei Li, Lin Han, Chenyu You, and Bo Wang. Segment anything in medical images. Nature communications, 15(1):654, 2024. 12 VERITASA PREPRINT
2024
-
[7]
MedSAM2: Segment anything in 3d medical images and videos, 2025
Jun Ma, Zongxin Yang, Sumin Kim, Bihui Chen, Mohammed Baharoon, Adibvafa Fallahpour, Reza Asakereh, Hongwei Lyu, and Bo Wang. Medsam2: Segment anything in 3d medical images and videos. arXiv preprint arXiv:2504.03600, 2025
-
[8]
Medical sam 2: Segment medical images as video via segment anything model 2,
Jiayuan Zhu, Abdullah Hamdi, Yunli Qi, Yueming Jin, and Junde Wu. Medical sam 2: Segment medical images as video via segment anything model 2.arXiv preprint arXiv:2408.00874, 2024
-
[9]
Large-vocabulary segmentation for medical images with text prompts.NPJ Digital Medicine, 8(1): 566, 2025
Ziheng Zhao, Yao Zhang, Chaoyi Wu, Xiaoman Zhang, Xiao Zhou, Ya Zhang, Yanfeng Wang, and Weidi Xie. Large-vocabulary segmentation for medical images with text prompts.NPJ Digital Medicine, 8(1): 566, 2025
2025
-
[10]
arXiv preprint arXiv:2511.11450, 2025
Maximilian Rokuss, Moritz Langenberg, Yannick Kirchhoff, Fabian Isensee, Benjamin Hamm, Con- stantin Ulrich, Sebastian Regnery, Lukas Bauer, Efthimios Katsigiannopulos, Tobias Norajitra, et al. Voxtell: Free-text promptable universal 3d medical image segmentation.arXiv preprint arXiv:2511.11450, 2025
-
[11]
nnu-net: a self-configuring method for deep learning-based biomedical image segmentation.Nature methods, 18(2): 203–211, 2021
Fabian Isensee, Paul F Jaeger, Simon AA Kohl, Jens Petersen, and Klaus H Maier-Hein. nnu-net: a self-configuring method for deep learning-based biomedical image segmentation.Nature methods, 18(2): 203–211, 2021
2021
-
[12]
Learning to exploit temporal structure for biomedical vision-language processing
Shruthi Bannur, Stephanie Hyland, Qianchu Liu, Fernando Perez-Garcia, Maximilian Ilse, Daniel C Castro, Benedikt Boecking, Harshita Sharma, Kenza Bouzid, Anja Thieme, et al. Learning to exploit temporal structure for biomedical vision-language processing. InProceedings of the IEEE/CVF conference on computer vision and pattern recognition, pages 15016–15027, 2023
2023
-
[13]
Llava-med: Training a large language-and-vision assistant for biomedicine in one day.Advances in Neural Information Processing Systems, 36:28541–28564, 2023
Chunyuan Li, Cliff Wong, Sheng Zhang, Naoto Usuyama, Haotian Liu, Jianwei Yang, Tristan Naumann, Hoifung Poon, and Jianfeng Gao. Llava-med: Training a large language-and-vision assistant for biomedicine in one day.Advances in Neural Information Processing Systems, 36:28541–28564, 2023
2023
-
[14]
Theodore Zhao, Yu Gu, Jianwei Yang, Naoto Usuyama, Ho Hin Lee, Tristan Naumann, Jianfeng Gao, Angela Crabtree, Jacob Abel, Christine Moung-Wen, et al. Biomedparse: a biomedical foundation model for image parsing of everything everywhere all at once.arXiv preprint arXiv:2405.12971, 2024
-
[15]
Andrew Hoopes, Neel Dey, Victor Ion Butoi, John V Guttag, and Adrian V Dalca. Voxelprompt: A vision agent for end-to-end medical image analysis.arXiv preprint arXiv:2410.08397, 2024
-
[16]
Introducing gpt-5.2, 12 2025
OpenAI. Introducing gpt-5.2, 12 2025. URL https://openai.com/index/ introducing-gpt-5-2/. Accessed: 2026-03-03
2025
-
[17]
The claude 3 model family: Opus, sonnet, haiku.Anthropic, 2024
Anthropic. The claude 3 model family: Opus, sonnet, haiku.Anthropic, 2024
2024
-
[18]
Gheorghe Comanici, Eric Bieber, Mike Schaekermann, Ice Pasupat, Noveen Sachdeva, Inderjit Dhillon, Marcel Blistein, Ori Ram, Dan Zhang, Evan Rosen, et al. Gemini 2.5: Pushing the frontier with advanced reasoning, multimodality, long context, and next generation agentic capabilities.arXiv preprint arXiv:2507.06261, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[19]
Jiaqi Wei, Yuejin Yang, Xiang Zhang, Yuhan Chen, Xiang Zhuang, Zhangyang Gao, Dongzhan Zhou, Guangshuai Wang, Zhiqiang Gao, Juntai Cao, et al. From ai for science to agentic science: A survey on autonomous scientific discovery.arXiv preprint arXiv:2508.14111, 2025
-
[20]
Empowering biomedical discovery with ai agents.Cell, 187(22):6125–6151, 2024
Shanghua Gao, Ada Fang, Yepeng Huang, Valentina Giunchiglia, Ayush Noori, Jonathan Richard Schwarz, Yasha Ektefaie, Jovana Kondic, and Marinka Zitnik. Empowering biomedical discovery with ai agents.Cell, 187(22):6125–6151, 2024
2024
-
[21]
The AI Scientist: Towards Fully Automated Open-Ended Scientific Discovery
Chris Lu, Cong Lu, Robert Tjarko Lange, Jakob Foerster, Jeff Clune, and David Ha. The ai scientist: Towards fully automated open-ended scientific discovery.arXiv preprint arXiv:2408.06292, 2024
work page internal anchor Pith review arXiv 2024
-
[22]
Juraj Gottweis, Wei-Hung Weng, Alexander Daryin, Tao Tu, Anil Palepu, Petar Sirkovic, Artiom Myaskovsky, Felix Weissenberger, Keran Rong, Ryutaro Tanno, et al. Towards an ai co-scientist.arXiv preprint arXiv:2502.18864, 2025
work page internal anchor Pith review arXiv 2025
-
[23]
Ali Essam Ghareeb, Benjamin Chang, Ludovico Mitchener, Angela Yiu, Caralyn J Szostkiewicz, Jon M Laurent, Muhammed T Razzak, Andrew D White, Michaela M Hinks, and Samuel G Rodriques. Robin: A multi-agent system for automating scientific discovery.arXiv preprint arXiv:2505.13400, 2025
-
[24]
Disciple: Learning interpretable programs for scientific visual discovery
Utkarsh Mall, Cheng Perng Phoo, Mia Chiquier, Bharath Hariharan, Kavita Bala, and Carl Vondrick. Disciple: Learning interpretable programs for scientific visual discovery. InProceedings of the Computer Vision and Pattern Recognition Conference, pages 29258–29267, 2025
2025
-
[25]
POPPER: Agentic fal- sification of free-form hypotheses.arXiv preprint arXiv:2502.09858, 2025
Kexin Huang, Ying Jin, Ryan Li, Michael Y Li, Emmanuel Candès, and Jure Leskovec. Automated hypothesis validation with agentic sequential falsifications.arXiv preprint arXiv:2502.09858, 2025. 13 VERITASA PREPRINT
-
[26]
Toolformer: Language models can teach them- selves to use tools.Advances in neural information processing systems, 36:68539–68551, 2023
Timo Schick, Jane Dwivedi-Yu, Roberto Dessì, Roberta Raileanu, Maria Lomeli, Eric Hambro, Luke Zettlemoyer, Nicola Cancedda, and Thomas Scialom. Toolformer: Language models can teach them- selves to use tools.Advances in neural information processing systems, 36:68539–68551, 2023
2023
-
[27]
Vipergpt: Visual inference via python execution for reasoning
Dídac Surís, Sachit Menon, and Carl Vondrick. Vipergpt: Visual inference via python execution for reasoning. InProceedings of the IEEE/CVF international conference on computer vision, pages 11888–11898, 2023
2023
-
[28]
ToolLLM: Facilitating Large Language Models to Master 16000+ Real-world APIs
Yujia Qin, Shihao Liang, Yining Ye, Kunlun Zhu, Lan Yan, Yaxi Lu, Yankai Lin, Xin Cong, Xiangru Tang, Bill Qian, et al. Toolllm: Facilitating large language models to master 16000+ real-world apis.arXiv preprint arXiv:2307.16789, 2023
work page internal anchor Pith review Pith/arXiv arXiv 2023
-
[29]
Executable code actions elicit better llm agents
Xingyao Wang, Yangyi Chen, Lifan Yuan, Yizhe Zhang, Yunzhu Li, Hao Peng, and Heng Ji. Executable code actions elicit better llm agents. InForty-first International Conference on Machine Learning, 2024
2024
-
[30]
Gorilla: Large language model connected with massive apis.Advances in Neural Information Processing Systems, 37:126544–126565, 2024
Shishir G Patil, Tianjun Zhang, Xin Wang, and Joseph E Gonzalez. Gorilla: Large language model connected with massive apis.Advances in Neural Information Processing Systems, 37:126544–126565, 2024
2024
-
[31]
React: Synergizing reasoning and acting in language models
Shunyu Yao, Jeffrey Zhao, Dian Yu, Nan Du, Izhak Shafran, Karthik R Narasimhan, and Yuan Cao. React: Synergizing reasoning and acting in language models. InThe eleventh international conference on learning representations, 2022
2022
-
[32]
Wenhu Chen, Xueguang Ma, Xinyi Wang, and William W Cohen. Program of thoughts prompting: Dis- entangling computation from reasoning for numerical reasoning tasks.arXiv preprint arXiv:2211.12588, 2022
work page internal anchor Pith review arXiv 2022
-
[33]
Pal: Program-aided language models
Luyu Gao, Aman Madaan, Shuyan Zhou, Uri Alon, Pengfei Liu, Yiming Yang, Jamie Callan, and Graham Neubig. Pal: Program-aided language models. InInternational conference on machine learning, pages 10764–10799. PMLR, 2023
2023
-
[34]
arXiv preprint arXiv:2506.14142 (2025)
Wenting Chen, Yi Dong, Zhaojun Ding, Yucheng Shi, Yifan Zhou, Fang Zeng, Yijun Luo, Tianyu Lin, Yihang Su, Yichen Wu, et al. Radfabric: Agentic ai system with reasoning capability for radiology.arXiv preprint arXiv:2506.14142, 2025
-
[35]
Medagents: Large language models as collaborators for zero-shot medical reasoning
Xiangru Tang, Anni Zou, Zhuosheng Zhang, Ziming Li, Yilun Zhao, Xingyao Zhang, Arman Cohan, and Mark Gerstein. Medagents: Large language models as collaborators for zero-shot medical reasoning. InFindings of the Association for Computational Linguistics: ACL 2024, pages 599–621, 2024
2024
-
[36]
Medagent-pro: Towards evidence-based multi-modal medical diagnosis via reasoning agentic workflow,
Ziyue Wang, Junde Wu, Linghan Cai, Chang Han Low, Xihong Yang, Qiaxuan Li, and Yueming Jin. Medagent-pro: Towards evidence-based multi-modal medical diagnosis via reasoning agentic workflow. arXiv preprint arXiv:2503.18968, 2025
-
[37]
Mdagents: An adaptive collaboration of llms for medical decision-making.Advances in Neural Information Processing Systems, 37:79410–79452, 2024
Yubin Kim, Chanwoo Park, Hyewon Jeong, Yik S Chan, Xuhai Xu, Daniel McDuff, Hyeonhoon Lee, Marzyeh Ghassemi, Cynthia Breazeal, and Hae W Park. Mdagents: An adaptive collaboration of llms for medical decision-making.Advances in Neural Information Processing Systems, 37:79410–79452, 2024
2024
-
[38]
Agentmd: Empowering language agents for risk prediction with large-scale clinical tool learning.Nature Communications, 16(1):9377, 2025
Qiao Jin, Zhizheng Wang, Yifan Yang, Qingqing Zhu, Donald Wright, Thomas Huang, Nikhil Khandekar, Nicholas Wan, Xuguang Ai, W John Wilbur, et al. Agentmd: Empowering language agents for risk prediction with large-scale clinical tool learning.Nature Communications, 16(1):9377, 2025
2025
-
[39]
Junkai Li, Yunghwei Lai, Weitao Li, Jingyi Ren, Meng Zhang, Xinhui Kang, Siyu Wang, Peng Li, Ya-Qin Zhang, Weizhi Ma, et al. Agent hospital: A simulacrum of hospital with evolvable medical agents. arXiv preprint arXiv:2405.02957, 2024
-
[40]
arXiv preprint arXiv:2506.22405 , year=
Harsha Nori, Mayank Daswani, Christopher Kelly, Scott Lundberg, Marco Tulio Ribeiro, Marc Wilson, Xiaoxuan Liu, Viknesh Sounderajah, Jonathan Carlson, Matthew P Lungren, et al. Sequential diagnosis with language models.arXiv preprint arXiv:2506.22405, 2025
-
[41]
The virtual lab of ai agents designs new sars-cov-2 nanobodies.Nature, 646(8085):716–723, 2025
Kyle Swanson, Wesley Wu, Nash L Bulaong, John E Pak, and James Zou. The virtual lab of ai agents designs new sars-cov-2 nanobodies.Nature, 646(8085):716–723, 2025
2025
-
[42]
The virtual biotech: A multi-agent ai framework for therapeutic discovery and development.bioRxiv, pages 2026–02, 2026
Harrison G Zhang, Peter Eckmann, Jiacheng Miao, Andrew B Mahon, and James Zou. The virtual biotech: A multi-agent ai framework for therapeutic discovery and development.bioRxiv, pages 2026–02, 2026
2026
-
[43]
Agent laboratory: Using llm agents as research assistants
Samuel Schmidgall, Yusheng Su, Ze Wang, Ximeng Sun, Jialian Wu, Xiaodong Yu, Jiang Liu, Michael Moor, Zicheng Liu, and Emad Barsoum. Agent laboratory: Using llm agents as research assistants. Findings of the Association for Computational Linguistics: EMNLP 2025, pages 5977–6043, 2025. 14 VERITASA PREPRINT
2025
-
[44]
Many heads are better than one: Improved scientific idea generation by a llm-based multi-agent system
Haoyang Su, Renqi Chen, Shixiang Tang, Zhenfei Yin, Xinzhe Zheng, Jinzhe Li, Biqing Qi, Qi Wu, Hui Li, Wanli Ouyang, et al. Many heads are better than one: Improved scientific idea generation by a llm-based multi-agent system. InProceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pages 28201–28240, 2025
2025
-
[45]
arXiv preprint arXiv:2404.07738
Jinheon Baek, Sujay Kumar Jauhar, Silviu Cucerzan, and Sung Ju Hwang. Researchagent: Itera- tive research idea generation over scientific literature with large language models.arXiv preprint arXiv:2404.07738, 2024
-
[46]
Sciagents: automating scientific discovery through bioin- spired multi-agent intelligent graph reasoning.Advanced Materials, 37(22):2413523, 2025
Alireza Ghafarollahi and Markus J Buehler. Sciagents: automating scientific discovery through bioin- spired multi-agent intelligent graph reasoning.Advanced Materials, 37(22):2413523, 2025
2025
-
[47]
Autogen: Enabling next-gen llm applications via multi-agent conversa- tions
Qingyun Wu, Gagan Bansal, Jieyu Zhang, Yiran Wu, Beibin Li, Erkang Zhu, Li Jiang, Xiaoyun Zhang, Shaokun Zhang, Jiale Liu, et al. Autogen: Enabling next-gen llm applications via multi-agent conversa- tions. InFirst conference on language modeling, 2024
2024
-
[48]
Metagpt: Meta programming for a multi-agent collaborative framework
Sirui Hong, Mingchen Zhuge, Jonathan Chen, Xiawu Zheng, Yuheng Cheng, Jinlin Wang, Ceyao Zhang, Zili Wang, Steven Ka Shing Yau, Zijuan Lin, et al. Metagpt: Meta programming for a multi-agent collaborative framework. InThe twelfth international conference on learning representations, 2023
2023
-
[49]
Zonglin Yang, Wanhao Liu, Ben Gao, Yujie Liu, Wei Li, Tong Xie, Lidong Bing, Wanli Ouyang, Erik Cambria, and Dongzhan Zhou. Moose-chem2: Exploring llm limits in fine-grained scientific hypothesis discovery via hierarchical search.arXiv preprint arXiv:2505.19209, 2025
-
[50]
Scimon: Scientific inspiration machines optimized for novelty
Qingyun Wang, Doug Downey, Heng Ji, and Tom Hope. Scimon: Scientific inspiration machines optimized for novelty. InProceedings of the 62nd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pages 279–299, 2024
2024
-
[51]
Can ai conduct autonomous scientific research? case studies on two real-world tasks.bioRxiv, pages 2026–01, 2026
Shreyansh Agrawal, Harsh B Anadkat, Kiran K Athimoolam, Harsh Bhardwaj, Trishul Chowdhury, Shengtao Gao, Purva K Kamat, Vishwadeepsinh Makwana, Mohammed H Shariff, Amitesh Badkul, et al. Can ai conduct autonomous scientific research? case studies on two real-world tasks.bioRxiv, pages 2026–01, 2026
2026
-
[52]
Multi-agent reasoning for cardiovascular imaging phenotype analysis
Weitong Zhang, Mengyun Qiao, Chengqi Zang, Steven Niederer, Paul M Matthews, Wenjia Bai, and Bernhard Kainz. Multi-agent reasoning for cardiovascular imaging phenotype analysis. InInternational Conference on Medical Image Computing and Computer-Assisted Intervention, pages 429–439. Springer, 2025
2025
-
[53]
Moving to a world beyond “p< 0.05”, 2019
Ronald L Wasserstein, Allen L Schirm, and Nicole A Lazar. Moving to a world beyond “p< 0.05”, 2019
2019
-
[54]
Redefine statistical significance.Nature human behaviour, 2(1):6–10, 2018
Daniel J Benjamin, James O Berger, Magnus Johannesson, Brian A Nosek, E-J Wagenmakers, Richard Berk, Kenneth A Bollen, Björn Brembs, Lawrence Brown, Colin Camerer, et al. Redefine statistical significance.Nature human behaviour, 2(1):6–10, 2018
2018
-
[55]
The earth is round (p<
Jacob Cohen. The earth is round (p<. 05).American psychologist, 49(12):997, 1994
1994
-
[56]
Why most published research findings are false.PLoS medicine, 2(8):e124, 2005
John PA Ioannidis. Why most published research findings are false.PLoS medicine, 2(8):e124, 2005
2005
-
[57]
Power failure: why small sample size undermines the reliability of neuroscience.Nature reviews neuroscience, 14(5):365–376, 2013
Katherine S Button, John PA Ioannidis, Claire Mokrysz, Brian A Nosek, Jonathan Flint, Emma SJ Robinson, and Marcus R Munafò. Power failure: why small sample size undermines the reliability of neuroscience.Nature reviews neuroscience, 14(5):365–376, 2013
2013
-
[58]
Equivalence tests: A practical primer for t tests, correlations, and meta-analyses.Social psychological and personality science, 8(4):355–362, 2017
Daniël Lakens. Equivalence tests: A practical primer for t tests, correlations, and meta-analyses.Social psychological and personality science, 8(4):355–362, 2017
2017
-
[59]
Equivalence testing for psychological research: A tutorial.Advances in methods and practices in psychological science, 1(2):259–269, 2018
Daniël Lakens, Anne M Scheel, and Peder M Isager. Equivalence testing for psychological research: A tutorial.Advances in methods and practices in psychological science, 1(2):259–269, 2018
2018
-
[60]
The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping.Radiology, 295(2):328–338, 2020
Alex Zwanenburg, Martin Vallières, Mahmoud A Abdalah, Hugo JWL Aerts, Vincent Andrearczyk, Aditya Apte, Saeed Ashrafinia, Spyridon Bakas, Roelof J Beukinga, Ronald Boellaard, et al. The image biomarker standardization initiative: standardized quantitative radiomics for high-throughput image-based phenotyping.Radiology, 295(2):328–338, 2020
2020
-
[61]
Computational radiomics system to decode the radiographic phenotype.Cancer research, 77(21):e104– e107, 2017
Joost JM Van Griethuysen, Andriy Fedorov, Chintan Parmar, Ahmed Hosny, Nicole Aucoin, Vivek Narayan, Regina GH Beets-Tan, Jean-Christophe Fillion-Robin, Steve Pieper, and Hugo JWL Aerts. Computational radiomics system to decode the radiographic phenotype.Cancer research, 77(21):e104– e107, 2017
2017
-
[62]
Introducing gpt-oss, 8 2025
OpenAI. Introducing gpt-oss, 8 2025. URL https://openai.com/index/ introducing-gpt-oss/. Accessed: 2026-03-03
2025
-
[63]
An Yang, Anfeng Li, Baosong Yang, Beichen Zhang, Binyuan Hui, Bo Zheng, Bowen Yu, Chang Gao, Chengen Huang, Chenxu Lv, et al. Qwen3 technical report.arXiv preprint arXiv:2505.09388, 2025. 15 VERITASA PREPRINT
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[64]
Qwen3-coder-next technical report.arXiv preprint arXiv:2603.00729, 2026
Ruisheng Cao, Mouxiang Chen, Jiawei Chen, Zeyu Cui, Yunlong Feng, Binyuan Hui, Yuheng Jing, Kaixin Li, Mingze Li, Junyang Lin, et al. Qwen3-coder-next technical report.arXiv preprint arXiv:2603.00729, 2026
-
[65]
Gpt-5 mini, 1 2025
OpenAI. Gpt-5 mini, 1 2025. URL https://openai.com/index/gpt-5-mini/. Accessed: 2026- 03-03
2025
-
[66]
The table 2 fallacy: presenting and interpreting confounder and modifier coefficients.American journal of epidemiology, 177(4):292–298, 2013
Daniel Westreich and Sander Greenland. The table 2 fallacy: presenting and interpreting confounder and modifier coefficients.American journal of epidemiology, 177(4):292–298, 2013
2013
-
[67]
Cambridge University Press, 2009
Judea Pearl.Causality: Models, Reasoning, and Inference. Cambridge University Press, 2009
2009
-
[68]
Ollama: Get up and running with large language models
Ollama Team. Ollama: Get up and running with large language models. https://ollama.com/,
-
[69]
Accessed: 2026-03-03
2026
-
[70]
Openrouter: A unified interface for llms
OpenRouter Team. Openrouter: A unified interface for llms. https://openrouter.ai/, 2024. Accessed: 2026-03-03
2024
-
[71]
Cameron Davidson-Pilon. lifelines: survival analysis in python.Journal of Open Source Software, 4(40): 1317, 2019. doi: 10.21105/joss.01317. URLhttps://doi.org/10.21105/joss.01317
-
[72]
Langgraph: Building stateful, multi-actor applications with llms
LangChain AI. Langgraph: Building stateful, multi-actor applications with llms. https://github. com/langchain-ai/langgraph, 2024. Accessed: 2026-03-03
2024
-
[73]
The use of predicted confidence intervals when planning experiments and the misuse of power when interpreting results.Annals of internal medicine, 121(3): 200–206, 1994
Steven N Goodman and Jesse A Berlin. The use of predicted confidence intervals when planning experiments and the misuse of power when interpreting results.Annals of internal medicine, 121(3): 200–206, 1994
1994
-
[74]
DCM patients show significantly lower LVEF than normal controls
John M Hoenig and Dennis M Heisey. The abuse of power: the pervasive fallacy of power calculations for data analysis.The American Statistician, 55(1):19–24, 2001. 16 VERITASA PREPRINT A Example Workflow Walkthrough This section illustrates the end-to-end VERITASpipeline on two representative hypotheses: one imaging- based group comparison (ACDC) and one m...
2001
-
[75]
Iterates over all patients, loading LV masks at ED and ES via the Imaging Analysis API
-
[76]
Computes voxel-level volumes using sat.calculate_volume(mask, spacing)
-
[77]
Derives per-patient LVEF from the exact formula
-
[78]
Applies five prespecified QC checks (fi- nite volumes, EDV >0 , ESV ≥0 , ESV ≤ EDV)
-
[79]
group") not in [
Performs a Welch t-test and saves complete statistics to statistical_results.json Key excerpt from the generated code: 1for pid in patients: 2md = sat.get_patient_metadata(pid) 3if md.get("group") not in ["DCM", "NOR"]: 4continue 5obs_map = sat.get_observation_identifiers(pid) 6for obs in ["ED", "ES"]: 7mask = sat.load_structure_mask( 8results_db_path, pi...
2000
-
[80]
Test-family correctness: the statistical test matches the hypothesis type (group difference → Mann- Whitney U; correlation→Spearman; survival→log-rank/Cox PH)
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.