Recognition: unknown
Deciphering the chemical grammar of protein-RNA condensates
Pith reviewed 2026-05-10 06:53 UTC · model grok-4.3
The pith
Even ultrashort dipeptides encode the instructions for spontaneous protein-RNA condensation through base-specific interactions.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
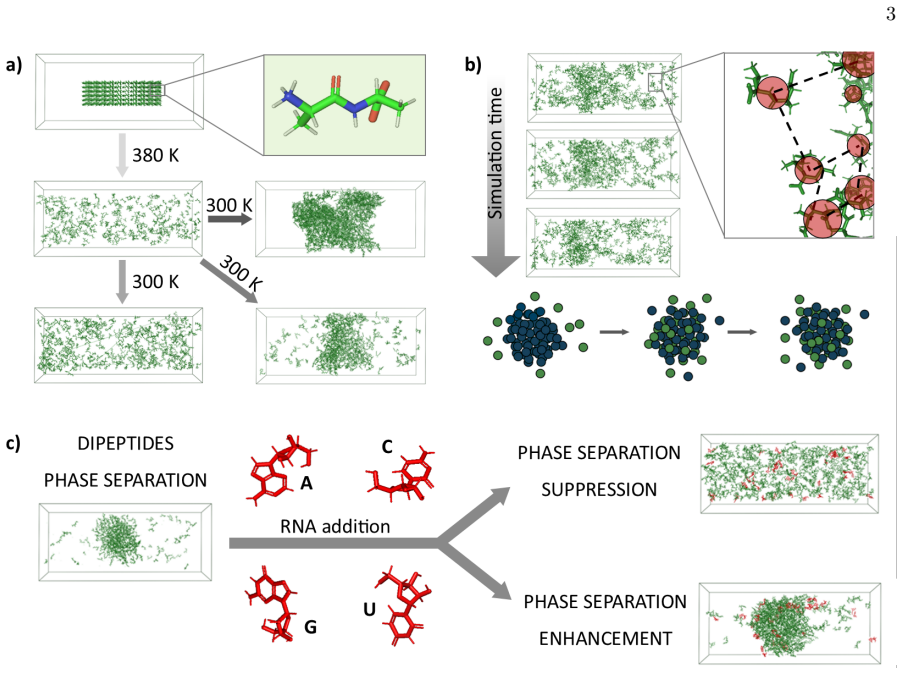
Screening the phase behavior of the complete dipeptide library with and without nucleic acids via full-atomistic molecular dynamics simulations demonstrates that ultrashort peptide units encode instructions for spontaneous condensation. This proves phase separation is rooted at a sub-polymeric level. Nucleic acids exert a base-specific regulatory logic rather than serving as generic anionic glue. Individual nucleobases act as chemical tuners that can dissolve, stabilize, or fluidize the condensates depending on their identity. While polymer length enhances assembly, the core properties are governed by this fine-tuned chemical alphabet of peptides and nucleobases.
What carries the argument
Full-atomistic molecular dynamics simulations screening the complete dipeptide library in the presence and absence of nucleic acids to isolate specific chemical interactions at the sub-polymeric scale.
Load-bearing premise
That full-atomistic molecular dynamics simulations of isolated dipeptides and nucleobases accurately capture the driving forces and phase behavior that occur in longer chains and crowded cellular environments.
What would settle it
Laboratory experiments mixing purified dipeptides with specific nucleobases that fail to produce the predicted condensation behavior or base-dependent tuning effects.
Figures








read the original abstract
Biomolecular phase separation is typically attributed to the polymer physics of long, disordered chains. However, the underlying chemical grammar, i.e. the specific interactions between protein and RNA building blocks, remains poorly understood. We decouple those effects by screening the phase behavior of the complete dipeptide library in presence and absence of nucleic acids using full-atomistic molecular dynamics simulations. We demonstrate that (i) even these ultrashort units encode the instructions for spontaneous condensation, proving that phase separation is fundamentally rooted at a sub-polymeric level. (ii) Nucleic acids do not act as generic anionic glue but exert instead a base-specific regulatory logic. (iii) Individual nucleobases function as chemical tuners that dissolve, stabilize, or fluidize condensates based on their molecular identity. Overall, our minimal framework reveals that while polymer length enhances assembly, the core properties and regulatory control of condensates may be also governed by a fine-tuned chemical alphabet of peptides and nucleobases.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript uses full-atomistic molecular dynamics simulations to screen the complete dipeptide library in the presence and absence of nucleic acids. It claims that ultrashort dipeptides spontaneously condense, establishing that biomolecular phase separation is rooted at a sub-polymeric level; that nucleic acids exert base-specific regulatory logic rather than acting as generic anionic glue; and that individual nucleobases function as chemical tuners that can dissolve, stabilize, or fluidize condensates. The work positions this minimal chemical alphabet as governing core condensate properties even when polymer length enhances assembly.
Significance. If the central claims hold after addressing validation concerns, the work would be significant for reframing condensate research around sequence-specific chemical interactions at the building-block scale rather than solely polymer physics. The systematic dipeptide screen offers a potentially useful reference framework for dissecting regulatory logic, and the atomistic resolution provides molecular detail on hydrogen bonding, stacking, and hydrophobic contacts. Strengths include the exhaustive library coverage and the attempt to decouple peptide and nucleic-acid contributions.
major comments (3)
- [Abstract and Results] Abstract and Results: The central claim that dipeptides undergo 'spontaneous condensation' and thereby prove a sub-polymeric root for LLPS requires explicit demonstration that the observed clustering constitutes true phase separation (e.g., via density histograms, order parameters, or finite-size scaling) rather than local association; standard dipeptide MD typically reports radial distribution functions or small-cluster statistics, and the manuscript must show how these extrapolate to macroscopic behavior.
- [Methods] Methods: Force-field choice, system sizes, simulation lengths, and convergence diagnostics for the reported phase behavior are not detailed; without these, the base-specific regulatory effects and nucleobase-tuning claims cannot be assessed for robustness against known limitations of current force fields in describing hydrogen bonding and π-stacking in crowded environments.
- [Discussion] Discussion: The assertion that dipeptide-level interactions govern the same regulatory logic as in longer polymeric condensates lacks direct transferability tests (e.g., comparison simulations with oligopeptides or experimental cross-validation); this assumption is load-bearing for the claim that the chemical grammar is fundamentally sub-polymeric.
minor comments (2)
- [Figures] Figure captions should specify the exact simulation conditions (temperature, concentration, box size) used to generate each panel so that readers can evaluate the reported condensation behavior.
- Notation for nucleobases and dipeptides should be standardized throughout (e.g., consistent use of one-letter codes or full names) to improve readability.
Simulated Author's Rebuttal
We thank the referee for the thoughtful and constructive report. The comments highlight important aspects of validating phase separation in atomistic simulations and the transferability of our findings. We address each major comment below and outline revisions that strengthen the manuscript without altering its core claims.
read point-by-point responses
-
Referee: [Abstract and Results] The central claim that dipeptides undergo 'spontaneous condensation' and thereby prove a sub-polymeric root for LLPS requires explicit demonstration that the observed clustering constitutes true phase separation (e.g., via density histograms, order parameters, or finite-size scaling) rather than local association; standard dipeptide MD typically reports radial distribution functions or small-cluster statistics, and the manuscript must show how these extrapolate to macroscopic behavior.
Authors: We agree that distinguishing true phase separation from transient local associations is critical. Our simulations show persistent, large-scale clusters (hundreds of dipeptides) that remain stable over multi-microsecond trajectories, with clear separation from the dilute phase, which we interpret as condensation. To strengthen this, we will add density histograms across the simulation box and cluster-size distributions as a function of time in the revised Results section. Finite-size scaling is computationally prohibitive at atomistic resolution for the full dipeptide library, but we will include a discussion of system-size effects and cite supporting coarse-grained studies that confirm the same trends extrapolate to larger scales. revision: partial
-
Referee: [Methods] Force-field choice, system sizes, simulation lengths, and convergence diagnostics for the reported phase behavior are not detailed; without these, the base-specific regulatory effects and nucleobase-tuning claims cannot be assessed for robustness against known limitations of current force fields in describing hydrogen bonding and π-stacking in crowded environments.
Authors: We acknowledge that the Methods section in the current version is concise and should be expanded for reproducibility. The simulations employed the CHARMM36 force field with TIP3P water, systems of 200-500 dipeptides plus 50-100 nucleotides in periodic boxes of ~10-15 nm, and production runs of 1-5 µs per condition after equilibration. Convergence was assessed via block averaging of cluster sizes and interaction energies. We will add a dedicated subsection with these parameters, plus additional diagnostics (e.g., time series of radius of gyration and hydrogen-bond counts) to the revised Methods and Supplementary Information. revision: yes
-
Referee: [Discussion] The assertion that dipeptide-level interactions govern the same regulatory logic as in longer polymeric condensates lacks direct transferability tests (e.g., comparison simulations with oligopeptides or experimental cross-validation); this assumption is load-bearing for the claim that the chemical grammar is fundamentally sub-polymeric.
Authors: This is a fair point; our work deliberately focuses on the minimal dipeptide units to isolate the chemical grammar, and we do not claim identical behavior in polymers. We will revise the Discussion to explicitly state this scope limitation, add a paragraph comparing our dipeptide trends to published experimental and simulation data on short peptides and IDRs (e.g., similar base-specific modulation), and note that polymer length amplifies but does not qualitatively alter the underlying interactions. Direct oligopeptide simulations or new experiments are beyond the present computational scope but represent valuable future work. revision: yes
Circularity Check
No circularity: claims rest on direct MD simulation outputs
full rationale
The paper performs a computational screen of dipeptide-nucleic acid mixtures via full-atomistic MD and reports observed clustering or phase behaviors as its primary evidence. No mathematical derivation, parameter fitting, or model is constructed whose outputs are then re-used as inputs. Claims about sub-polymeric encoding of condensation are presented as simulation results rather than as predictions derived from a self-referential equation or ansatz. No load-bearing self-citations or uniqueness theorems are invoked that reduce the central argument to prior work by the same authors. The study is therefore self-contained as an empirical simulation report.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Full-atomistic force fields and simulation protocols are sufficient to capture the driving forces of dipeptide-RNA condensation
Reference graph
Works this paper leans on
-
[1]
Regulation of cellular metabolism through phase separation of enzymes.Biomolecules, 8(4):160, 2018
Mano¨ el Prouteau and Robbie Loewith. Regulation of cellular metabolism through phase separation of enzymes.Biomolecules, 8(4):160, 2018
2018
-
[2]
Phase separation in transcription fac- tor dynamics and chromatin organization.Current Opinion in Structural Biology, 71:148–155, 2021
Kaustubh Wagh, David A Garcia, and Arpita Upadhyaya. Phase separation in transcription fac- tor dynamics and chromatin organization.Current Opinion in Structural Biology, 71:148–155, 2021
2021
-
[3]
Biomolecular con- densates: organizers of cellular biochemistry.Na- ture reviews Molecular cell biology, 18(5):285–298, 2017
Salman F Banani, Hyun O Lee, Anthony A Hy- man, and Michael K Rosen. Biomolecular con- densates: organizers of cellular biochemistry.Na- ture reviews Molecular cell biology, 18(5):285–298, 2017
2017
-
[4]
Liquid phase condensation in cell physiology and disease
Yongdae Shin and Clifford P Brangwynne. Liquid phase condensation in cell physiology and disease. Science, 357(6357):eaaf4382, 2017
2017
-
[5]
Multi-condensate state as a functional strategy to optimize the cell signaling output.Nature Com- munications, 15(1):6268, 2024
Aniruddha Chattaraj and Eugene I Shakhnovich. Multi-condensate state as a functional strategy to optimize the cell signaling output.Nature Com- munications, 15(1):6268, 2024
2024
-
[6]
Transcription regulation by biomolecular condensates.Nature Reviews Molec- ular Cell Biology, 26(3):213–236, 2025
Gaofeng Pei, Heankel Lyons, Pilong Li, and Ben- jamin R Sabari. Transcription regulation by biomolecular condensates.Nature Reviews Molec- ular Cell Biology, 26(3):213–236, 2025
2025
-
[7]
Rna granules: a view from the rna per- spective.Molecules, 25(14):3130, 2020
Siran Tian, Harrison A Curnutte, and Tatjana Trcek. Rna granules: a view from the rna per- spective.Molecules, 25(14):3130, 2020
2020
-
[8]
Paraspeckle nuclear condensates: Global sensors of cell stress? Bioessays, 43(5):2000245, 2021
Finn McCluggage and Archa H Fox. Paraspeckle nuclear condensates: Global sensors of cell stress? Bioessays, 43(5):2000245, 2021
2021
-
[9]
Sequestration within biomolecular con- densates inhibits aβ-42 amyloid formation.Chem- ical Science, 12(12):4373–4382, 2021
Andreas M K¨ uffner, Miriam Linsenmeier, Fulvio Grigolato, Marc Prodan, Remo Zuccarini, Um- berto Capasso Palmiero, Lenka Faltova, and Paolo Arosio. Sequestration within biomolecular con- densates inhibits aβ-42 amyloid formation.Chem- ical Science, 12(12):4373–4382, 2021
2021
-
[10]
Noise buffering by biomolecular condensates in glucose sensing.Current Opinion in Cell Biology, 69:1–6, 2021
Kobi Simpson-Lavy and Martin Kupiec. Noise buffering by biomolecular condensates in glucose sensing.Current Opinion in Cell Biology, 69:1–6, 2021
2021
-
[11]
Germline p granules are liq- uid droplets that localize by controlled dissolu- tion/condensation.Science, 324(5935):1729–1732, 2009
Clifford P Brangwynne, Christian R Eckmann, David S Courson, Agata Rybarska, Carsten Hoege, J¨ obin Gharakhani, Frank J¨ ulicher, and Anthony A Hyman. Germline p granules are liq- uid droplets that localize by controlled dissolu- tion/condensation.Science, 324(5935):1729–1732, 2009
2009
-
[12]
Liquid-liquid phase separation in biology.Annual review of cell and developmental biology, 30(1):39–58, 2014
Anthony A Hyman, Christoph A Weber, and Frank J¨ ulicher. Liquid-liquid phase separation in biology.Annual review of cell and developmental biology, 30(1):39–58, 2014
2014
-
[13]
A new era for understanding amyloid structures and disease.Nature reviews Molecular cell biology, 19(12):755–773, 2018
Matthew G Iadanza, Matthew P Jackson, Eric W Hewitt, Neil A Ranson, and Sheena E Radford. A new era for understanding amyloid structures and disease.Nature reviews Molecular cell biology, 19(12):755–773, 2018
2018
-
[14]
Mechanism underly- ing liquid-to-solid phase transition in fused in sar- coma liquid droplets.Physical Chemistry Chemi- cal Physics, 24(32):19346–19353, 2022
Shujie Li, Takuya Yoshizawa, Yutaro Shiramasa, Mako Kanamaru, Fumika Ide, Keiji Kitamura, Norika Kashiwagi, Naoya Sasahara, Soichiro Ki- tazawa, and Ryo Kitahara. Mechanism underly- ing liquid-to-solid phase transition in fused in sar- coma liquid droplets.Physical Chemistry Chemi- cal Physics, 24(32):19346–19353, 2022
2022
-
[15]
Biomolecular condensates undergo a generic shear-mediated liquid-to-solid transition.Nature nanotechnology, 15(10):841–847, 2020
Yi Shen, Francesco Simone Ruggeri, Daniele Vigolo, Ayaka Kamada, Seema Qamar, Aviad Levin, Christiane Iserman, Simon Alberti, Pe- ter St George-Hyslop, and Tuomas PJ Knowles. Biomolecular condensates undergo a generic shear-mediated liquid-to-solid transition.Nature nanotechnology, 15(10):841–847, 2020
2020
-
[16]
The physics of liquid-to-solid transitions in multi- domain protein condensates.Biophysical Journal, 121(14):2751–2766, 2022
Srivastav Ranganathan and Eugene Shakhnovich. The physics of liquid-to-solid transitions in multi- domain protein condensates.Biophysical Journal, 121(14):2751–2766, 2022
2022
-
[17]
Challenges in study- ing the liquid-to-solid phase transitions of proteins using computer simulations.Current opinion in chemical biology, 75:102333, 2023
Beata Sza la-Mendyk, Tien Minh Phan, Priyesh Mohanty, and Jeetain Mittal. Challenges in study- ing the liquid-to-solid phase transitions of proteins using computer simulations.Current opinion in chemical biology, 75:102333, 2023
2023
-
[18]
A computational approach to investigate tdp-43 rna-recognition motif 2 c-terminal frag- ments aggregation in amyotrophic lateral sclerosis
Greta Grassmann, Mattia Miotto, Lorenzo Di Rienzo, Federico Salaris, Beatrice Silvestri, Elsa Zacco, Alessandro Rosa, Gian Gaetano Tartaglia, Giancarlo Ruocco, and Edoardo Mi- lanetti. A computational approach to investigate tdp-43 rna-recognition motif 2 c-terminal frag- ments aggregation in amyotrophic lateral sclerosis. Biomolecules, 11(12):1905, 2021
1905
-
[19]
Biomolec- ular condensates at the nexus of cellular stress, protein aggregation disease and ageing.Nature re- views Molecular cell biology, 22(3):196–213, 2021
Simon Alberti and Anthony A Hyman. Biomolec- ular condensates at the nexus of cellular stress, protein aggregation disease and ageing.Nature re- views Molecular cell biology, 22(3):196–213, 2021
2021
-
[20]
Atomic structures of low-complexity protein segments reveal kinked βsheets that assemble networks.Science, 359(6376):698–701, 2018
Michael P Hughes, Michael R Sawaya, David R Boyer, Lukasz Goldschmidt, Jose A Rodriguez, Duilio Cascio, Lisa Chong, Tamir Gonen, and David S Eisenberg. Atomic structures of low-complexity protein segments reveal kinked βsheets that assemble networks.Science, 359(6376):698–701, 2018
2018
-
[21]
Cell-free formation of rna granules: low complexity sequence domains form dynamic fibers within hydrogels.Cell, 149(4):753–767, 2012
Masato Kato, Tina W Han, Shanhai Xie, Kevin Shi, Xinlin Du, Leeju C Wu, Hamid Mirzaei, Eliz- abeth J Goldsmith, Jamie Longgood, Jimin Pei, et al. Cell-free formation of rna granules: low complexity sequence domains form dynamic fibers within hydrogels.Cell, 149(4):753–767, 2012
2012
-
[22]
Atomic structures of fus lc domain segments reveal bases for re- versible amyloid fibril formation.Nature structural & molecular biology, 25(4):341–346, 2018
Feng Luo, Xinrui Gui, Heng Zhou, Jinge Gu, Yichen Li, Xiangyu Liu, Minglei Zhao, Dan Li, Xueming Li, and Cong Liu. Atomic structures of fus lc domain segments reveal bases for re- versible amyloid fibril formation.Nature structural & molecular biology, 25(4):341–346, 2018
2018
-
[23]
Polymer physics of intracellular phase transitions.Nature Physics, 11(11):899– 904, 2015
Clifford P Brangwynne, Peter Tompa, and Ro- hit V Pappu. Polymer physics of intracellular phase transitions.Nature Physics, 11(11):899– 904, 2015
2015
-
[24]
Classifi- cation of intrinsically disordered regions and pro- teins.Chemical reviews, 114(13):6589–6631, 2014
Robin Van Der Lee, Marija Buljan, Benjamin Lang, Robert J Weatheritt, Gary W Daughdrill, A Keith Dunker, Monika Fuxreiter, Julian Gough, Joerg Gsponer, David T Jones, et al. Classifi- cation of intrinsically disordered regions and pro- teins.Chemical reviews, 114(13):6589–6631, 2014
2014
-
[25]
Intrinsically 13 disordered proteins in cellular signalling and reg- ulation.Nature reviews Molecular cell biology, 16(1):18–29, 2015
Peter E Wright and H Jane Dyson. Intrinsically 13 disordered proteins in cellular signalling and reg- ulation.Nature reviews Molecular cell biology, 16(1):18–29, 2015
2015
-
[26]
Condensates in rna repeat sequences are heteroge- neously organized and exhibit reptation dynamics
Hung T Nguyen, Naoto Hori, and D Thirumalai. Condensates in rna repeat sequences are heteroge- neously organized and exhibit reptation dynamics. Nature chemistry, 14(7):775–785, 2022
2022
-
[27]
Electrostatic complementarity at the interface drives transient protein-protein interactions.Scientific Reports, 13(1):10207, 2023
Greta Grassmann, Lorenzo Di Rienzo, Giorgio Gosti, Marco Leonetti, Giancarlo Ruocco, Mat- tia Miotto, and Edoardo Milanetti. Electrostatic complementarity at the interface drives transient protein-protein interactions.Scientific Reports, 13(1):10207, 2023
2023
-
[28]
Rna con- tributions to the form and function of biomolec- ular condensates.Nature Reviews Molecular Cell Biology, 22(3):183–195, 2021
Christine Roden and Amy S Gladfelter. Rna con- tributions to the form and function of biomolec- ular condensates.Nature Reviews Molecular Cell Biology, 22(3):183–195, 2021
2021
-
[29]
Sequence determinants of intra- cellular phase separation by complex coacervation of a disordered protein.Molecular cell, 63(1):72– 85, 2016
Chi W Pak, Martyna Kosno, Alex S Holehouse, Shae B Padrick, Anuradha Mittal, Rustam Ali, Ali A Yunus, David R Liu, Rohit V Pappu, and Michael K Rosen. Sequence determinants of intra- cellular phase separation by complex coacervation of a disordered protein.Molecular cell, 63(1):72– 85, 2016
2016
-
[30]
mrna structure determines specificity of a polyq-driven phase separation.Science, 360(6391):922–927, 2018
Erin M Langdon, Yupeng Qiu, Amirhossein Ghan- bari Niaki, Grace A McLaughlin, Chase A Weid- mann, Therese M Gerbich, Jean A Smith, John M Crutchley, Christina M Termini, Kevin M Weeks, et al. mrna structure determines specificity of a polyq-driven phase separation.Science, 360(6391):922–927, 2018
2018
-
[31]
m6a enhances the phase sep- aration potential of mrna.Nature, 571(7765):424– 428, 2019
Ryan J Ries, Sara Zaccara, Pierre Klein, Anthony Olarerin-George, Sim Namkoong, Brian F Picker- ing, Deepak P Patil, Hojoong Kwak, Jun Hee Lee, and Samie R Jaffrey. m6a enhances the phase sep- aration potential of mrna.Nature, 571(7765):424– 428, 2019
2019
-
[32]
The disordered p gran- ule protein laf-1 drives phase separation into droplets with tunable viscosity and dynamics
Shana Elbaum-Garfinkle, Younghoon Kim, Krzysztof Szczepaniak, Carlos Chih-Hsiung Chen, Christian R Eckmann, Sua Myong, and Clifford P Brangwynne. The disordered p gran- ule protein laf-1 drives phase separation into droplets with tunable viscosity and dynamics. Proceedings of the National Academy of Sciences, 112(23):7189–7194, 2015
2015
-
[33]
Rna controls polyq protein phase transitions.Molecular cell, 60(2):220–230, 2015
Huaiying Zhang, Shana Elbaum-Garfinkle, Erin M Langdon, Nicole Taylor, Patricia Occhipinti, An- drew A Bridges, Clifford P Brangwynne, and Amy S Gladfelter. Rna controls polyq protein phase transitions.Molecular cell, 60(2):220–230, 2015
2015
-
[34]
Rna buffers the phase separation behav- ior of prion-like rna binding proteins.Science, 360(6391):918–921, 2018
Shovamayee Maharana, Jie Wang, Dimitrios K Papadopoulos, Doris Richter, Andrey Pozni- akovsky, Ina Poser, Marc Bickle, Sandra Rizk, Jordina Guill´ en-Boixet, Titus M Franzmann, et al. Rna buffers the phase separation behav- ior of prion-like rna binding proteins.Science, 360(6391):918–921, 2018
2018
-
[35]
Inhibition of sars-cov-2 replication by a ssdna aptamer target- ing the nucleocapsid protein.Microbiology Spec- trum, 12(4):e03410–23, 2024
Yanping Huang, Congcong Huang, Junkai Chen, Siwei Chen, Bei Li, Jian Li, Zhixiong Jin, Qiwei Zhang, Pan Pan, Weixing Du, et al. Inhibition of sars-cov-2 replication by a ssdna aptamer target- ing the nucleocapsid protein.Microbiology Spec- trum, 12(4):e03410–23, 2024
2024
-
[36]
Liquid-liquid phase separation and fibrillation of the prion protein modulated by a high-affinity dna aptamer.The FASEB Journal, 34(1):365–385, 2020
Carolina O Matos, Yulli M Passos, Mariana J do Amaral, Bruno Macedo, Matheus H Tempone, Ohanna CL Bezerra, Milton O Moraes, Marcius S Almeida, Gerald Weber, Sotiris Missailidis, et al. Liquid-liquid phase separation and fibrillation of the prion protein modulated by a high-affinity dna aptamer.The FASEB Journal, 34(1):365–385, 2020
2020
-
[37]
Toward predictive coarse-grained simulations of biomolec- ular condensates.Biochemistry, 64(8):1750–1761, 2025
Shuming Liu, Cong Wang, and Bin Zhang. Toward predictive coarse-grained simulations of biomolec- ular condensates.Biochemistry, 64(8):1750–1761, 2025
2025
-
[38]
Tuning formation of protein–dna coacervates by sequence and environment.The Journal of Physical Chem- istry B, 126(12):2407–2419, 2022
Kathryn M Lebold and Robert B Best. Tuning formation of protein–dna coacervates by sequence and environment.The Journal of Physical Chem- istry B, 126(12):2407–2419, 2022
2022
-
[39]
Coarse-grained model of disor- dered rna for simulations of biomolecular conden- sates.Journal of chemical theory and computa- tion, 21(5):2766–2779, 2025
Ikki Yasuda, Sooren von Buulow, Giulio Te- sei, Eiji Yamamoto, Kenji Yasuoka, and Kresten Lindorff-Larsen. Coarse-grained model of disor- dered rna for simulations of biomolecular conden- sates.Journal of chemical theory and computa- tion, 21(5):2766–2779, 2025
2025
-
[40]
Phase-separated coacer- vates of low-molecular-weight compounds for cy- tosolic delivery and disease treatment.JACS Au, 5(11):5267–5285, 2025
Yishu Bao and Jiang Xia. Phase-separated coacer- vates of low-molecular-weight compounds for cy- tosolic delivery and disease treatment.JACS Au, 5(11):5267–5285, 2025
2025
-
[41]
A short peptide synthon for liquid–liquid phase separation.Nature Chemistry, 13(11):1046–1054, 2021
Manzar Abbas, Wojciech P Lipi´ nski, Karina K Nakashima, Wilhelm TS Huck, and Evan Spruijt. A short peptide synthon for liquid–liquid phase separation.Nature Chemistry, 13(11):1046–1054, 2021
2021
-
[42]
Virtual screening for dipeptide aggregation: toward predictive tools for peptide self-assembly.The journal of physical chemistry letters, 2(19):2380–2384, 2011
Pim WJM Frederix, Rein V Ulijn, Neil T Hunt, and Tell Tuttle. Virtual screening for dipeptide aggregation: toward predictive tools for peptide self-assembly.The journal of physical chemistry letters, 2(19):2380–2384, 2011
2011
-
[43]
Untersuchungen ¨ uber die guanyls¨ aure
I Bang. Untersuchungen ¨ uber die guanyls¨ aure. Biochem. Z, 26:293–311, 1910
1910
-
[44]
Phase transitions in the assembly of multivalent signalling proteins.Na- ture, 483(7389):336–340, 2012
Pilong Li, Sudeep Banjade, Hui-Chun Cheng, Soyeon Kim, Baoyu Chen, Liang Guo, Marc Lla- guno, Javoris V Hollingsworth, David S King, Salman F Banani, et al. Phase transitions in the assembly of multivalent signalling proteins.Na- ture, 483(7389):336–340, 2012
2012
-
[45]
Theory of simple liquids: with applications to soft matter
Jean-Pierre Hansen and Ian Ranald McDonald. Theory of simple liquids: with applications to soft matter. Academic press, 2013
2013
-
[46]
Nucleation: theory and appli- cations to protein solutions and colloidal suspen- sions.Journal of Physics: Condensed Matter, 19(3):033101, 2007
Richard P Sear. Nucleation: theory and appli- cations to protein solutions and colloidal suspen- sions.Journal of Physics: Condensed Matter, 19(3):033101, 2007
2007
-
[47]
Casting metal nanowires within discrete self-assembled peptide nanotubes.Science, 300(5619):625–627, 2003
Meital Reches and Ehud Gazit. Casting metal nanowires within discrete self-assembled peptide nanotubes.Science, 300(5619):625–627, 2003
2003
-
[48]
Self- assembly and application of diphenylalanine- based nanostructures.Chemical Society Reviews, 39(6):1877–1890, 2010
Xuehai Yan, Pengli Zhu, and Junbai Li. Self- assembly and application of diphenylalanine- based nanostructures.Chemical Society Reviews, 39(6):1877–1890, 2010
2010
-
[49]
Aggregation propen- sity of amyloidogenic and elastomeric dipep- tides constituents.Tetrahedron, 72(35):5369–5376, 2016
Vikas Kumar, K Vijaya Krishna, Shruti Khanna, and Khashti Ballabh Joshi. Aggregation propen- sity of amyloidogenic and elastomeric dipep- tides constituents.Tetrahedron, 72(35):5369–5376, 2016
2016
-
[50]
Ile-phe dipeptide self-assembly: clues to amyloid formation.Biophysical journal, 92(5):1732–1741, 2007
Natalia S´ anchez De Groot, Teodor Parella, Francesc X Aviles, Josep Vendrell, and Sal- vador Ventura. Ile-phe dipeptide self-assembly: clues to amyloid formation.Biophysical journal, 92(5):1732–1741, 2007. 14
2007
-
[51]
Prediction and characterization of liquid-liquid phase separation of minimalistic peptides.Cell Reports Physical Science, 2(9), 2021
Yiming Tang, Santu Bera, Yifei Yao, Jiyuan Zeng, Zenghui Lao, Xuewei Dong, Ehud Gazit, and Guanghong Wei. Prediction and characterization of liquid-liquid phase separation of minimalistic peptides.Cell Reports Physical Science, 2(9), 2021
2021
-
[52]
Confor- mational landscape of substituted prolines.Bio- physical reviews, 12(1):25–39, 2020
Himal Kanti Ganguly and Gautam Basu. Confor- mational landscape of substituted prolines.Bio- physical reviews, 12(1):25–39, 2020
2020
-
[53]
An analytic structure factor for macroion solutions.Molecu- lar Physics, 42(1):109–118, 1981
John B Hayter and Jeff Penfold. An analytic structure factor for macroion solutions.Molecu- lar Physics, 42(1):109–118, 1981
1981
-
[54]
Structure factor scaling in colloidal phase separation.Physical Re- view E—Statistical, Nonlinear, and Soft Matter Physics, 70(5):051405, 2004
Juan J Cerd` a, Tom´ as Sintes, CM Sorensen, and A Chakrabarti. Structure factor scaling in colloidal phase separation.Physical Re- view E—Statistical, Nonlinear, and Soft Matter Physics, 70(5):051405, 2004
2004
-
[55]
Non-equilibrium phase transitions in hy- brid voronoi models of cell colonies.arXiv, 2024
Mattia Miotto, Giancarlo Ruocco, and Matteo Paoluzzi. Non-equilibrium phase transitions in hy- brid voronoi models of cell colonies.arXiv, 2024
2024
-
[56]
Valence and patterning of aromatic residues determine the phase behavior of prion-like domains.Science, 367(6478):694–699, 2020
Erik W Martin, Alex S Holehouse, Ivan Peran, Mina Farag, J Jeremias Incicco, Anne Bremer, Christy R Grace, Andrea Soranno, Rohit V Pappu, and Tanja Mittag. Valence and patterning of aromatic residues determine the phase behavior of prion-like domains.Science, 367(6478):694–699, 2020
2020
-
[57]
Pi-pi contacts are an overlooked protein fea- ture relevant to phase separation.elife, 7:e31486, 2018
Robert McCoy Vernon, Paul Andrew Chong, Brian Tsang, Tae Hun Kim, Alaji Bah, Patrick Farber, Hong Lin, and Julie Deborah Forman- Kay. Pi-pi contacts are an overlooked protein fea- ture relevant to phase separation.elife, 7:e31486, 2018
2018
-
[58]
Spatial organization of hydrophobic and charged residues affects protein thermal stability and binding affinity.Scientific Reports, 12(1):12087, 2022
Fausta Desantis, Mattia Miotto, Lorenzo Di Rienzo, Edoardo Milanetti, and Giancarlo Ruocco. Spatial organization of hydrophobic and charged residues affects protein thermal stability and binding affinity.Scientific Reports, 12(1):12087, 2022
2022
-
[59]
Sequence determinants of protein phase behavior from a coarse-grained model.PLoS computational biol- ogy, 14(1):e1005941, 2018
Gregory L Dignon, Wenwei Zheng, Young C Kim, Robert B Best, and Jeetain Mittal. Sequence determinants of protein phase behavior from a coarse-grained model.PLoS computational biol- ogy, 14(1):e1005941, 2018
2018
-
[60]
Fluores- cence recovery after photobleaching: a versatile tool for mobility and interaction measurements in pharmaceutical research.Pharmaceutical re- search, 16(8):1153–1162, 1999
Tom KL Meyvis, Stefaan C De Smedt, Patrick Van Oostveldt, and Joseph Demeester. Fluores- cence recovery after photobleaching: a versatile tool for mobility and interaction measurements in pharmaceutical research.Pharmaceutical re- search, 16(8):1153–1162, 1999
1999
-
[61]
Computational ap- proaches to predict protein–protein interactions in crowded cellular environments.Chemical Reviews, 124(7):3932–3977, 2024
Greta Grassmann, Mattia Miotto, Fausta Desan- tis, Lorenzo Di Rienzo, Gian Gaetano Tartaglia, Annalisa Pastore, Giancarlo Ruocco, Michele Monti, and Edoardo Milanetti. Computational ap- proaches to predict protein–protein interactions in crowded cellular environments.Chemical Reviews, 124(7):3932–3977, 2024
2024
-
[62]
Jorge R Espinosa, Jerelle A Joseph, Ignacio Sanchez-Burgos, Adiran Garaizar, Daan Frenkel, and Rosana Collepardo-Guevara. Liquid network connectivity regulates the stability and composi- tion of biomolecular condensates with many com- ponents.Proceedings of the National Academy of Sciences, 117(24):13238–13247, 2020
2020
-
[63]
Reentrant liquid con- densate phase of proteins is stabilized by hy- drophobic and non-ionic interactions.Biophysical Journal, 120(3):28a, 2021
Georg Krainer, Timothy J Welsh, Jerelle A Joseph, Peter St George-Hyslop, Anthony A Hy- man, Rosana Collepardo-Guevara, Simon Alberti, and Tuomas PJ Knowles. Reentrant liquid con- densate phase of proteins is stabilized by hy- drophobic and non-ionic interactions.Biophysical Journal, 120(3):28a, 2021
2021
-
[64]
Biomolecular condensates: new opportunities for drug discovery and rna therapeutics.Trends in pharmacological sciences, 43(10):820–837, 2022
Brooke A Conti and Mariano Oppikofer. Biomolecular condensates: new opportunities for drug discovery and rna therapeutics.Trends in pharmacological sciences, 43(10):820–837, 2022
2022
-
[65]
On the aromatic character of the heterocyclic bases of dna and rna.The Journal of Organic Chemistry, 68(22):8607–8613, 2003
Micha l K Cyra´ nski, Miros law Gilski, Mariusz Jask´ olski, and Tadeusz Marek Krygowski. On the aromatic character of the heterocyclic bases of dna and rna.The Journal of Organic Chemistry, 68(22):8607–8613, 2003
2003
-
[66]
Coarse-grained simulation study of the association of selected dipeptides.The Journal of Physical Chemistry B, 128(50):12403–12415, 2024
Mateusz Lesniewski, Emilia I lowska, Justyna Saw- icka, Zihan Li, Chun Tang, and Adam Liwo. Coarse-grained simulation study of the association of selected dipeptides.The Journal of Physical Chemistry B, 128(50):12403–12415, 2024
2024
-
[67]
Self-assembly of heterochiral, aliphatic dipeptides with leu.Journal of Peptide Science, 30(5):e3559, 2024
Erica Scarel, Marco De Corti, Maurizio Polen- tarutti, Giovanni Pierri, Consiglia Tedesco, and Silvia Marchesan. Self-assembly of heterochiral, aliphatic dipeptides with leu.Journal of Peptide Science, 30(5):e3559, 2024
2024
-
[68]
Sequence-encoded and composition-dependent protein-rna interac- tions control multiphasic condensate morpholo- gies.Nature communications, 12(1):872, 2021
Taranpreet Kaur, Muralikrishna Raju, Ibraheem Alshareedah, Richoo B Davis, Davit A Po- toyan, and Priya R Banerjee. Sequence-encoded and composition-dependent protein-rna interac- tions control multiphasic condensate morpholo- gies.Nature communications, 12(1):872, 2021
2021
-
[69]
Ibraheem Alshareedah, Taranpreet Kaur, Ja- son Ngo, Hannah Seppala, Liz-Audrey Djom- nang Kounatse, Wei Wang, Mahdi Muhammad Moosa, and Priya R Banerjee. Interplay be- tween short-range attraction and long-range re- pulsion controls reentrant liquid condensation of ribonucleoprotein–rna complexes.Journal of the American Chemical Society, 141(37):14593–14602, 2019
2019
-
[70]
Phase transition of rna- protein complexes into ordered hollow conden- sates.Proceedings of the National Academy of Sciences, 117(27):15650–15658, 2020
Ibraheem Alshareedah, Mahdi Muhammad Moosa, Muralikrishna Raju, Davit A Potoyan, and Priya R Banerjee. Phase transition of rna- protein complexes into ordered hollow conden- sates.Proceedings of the National Academy of Sciences, 117(27):15650–15658, 2020
2020
-
[71]
Quantifying viscosity and surface tension of multicomponent protein- nucleic acid condensates.Biophysical journal, 120(7):1161–1169, 2021
Ibraheem Alshareedah, George M Thurston, and Priya R Banerjee. Quantifying viscosity and surface tension of multicomponent protein- nucleic acid condensates.Biophysical journal, 120(7):1161–1169, 2021
2021
-
[72]
Phase behaviour of disordered proteins underlying low density and high permeability of liquid organelles
Ming-Tzo Wei, Shana Elbaum-Garfinkle, Alex S Holehouse, Carlos Chih-Hsiung Chen, Marina Feric, Craig B Arnold, Rodney D Priestley, Ro- hit V Pappu, and Clifford P Brangwynne. Phase behaviour of disordered proteins underlying low density and high permeability of liquid organelles. Nature chemistry, 9(11):1118–1125, 2017
2017
-
[73]
Van Der Spoel, E
D. Van Der Spoel, E. Lindahl, B. Hess, G. Groen- hof, A. E. Mark, and H. J. C. Berendsen. GRO- MACS: Fast, flexible, and free.Journal of Com- putational Chemistry, 26(16):1701–1718, 2005
2005
-
[74]
MacKerell, Donald Bashford, Michael Bellott, Roland L
Alexander D. MacKerell, Donald Bashford, Michael Bellott, Roland L. Dunbrack, J. David Evanseck, Martin J. Field, Siegfried Fischer, Jun Gao, Haibo Guo, Suyong Ha, et al. All-atom em- pirical potential for molecular modeling and dy- 15 namics studies of proteins.The Journal of Physi- cal Chemistry B, 102(18):3586–3616, 1998
1998
-
[75]
Jorgensen, Jayaraman Chandrasekhar, Jeffry D
William L. Jorgensen, Jayaraman Chandrasekhar, Jeffry D. Madura, Roger W. Impey, and Michael L. Klein. Comparison of simple potential functions for simulating liquid water.The Journal of Chemical Physics, 79(2):926–935, July 1983. 1 FIG. 1:Persistent Clusters (PCs) as spheres.For eachP C, Radius of gyration (R g ) versus Asphericity on the left and Fracta...
1983
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.