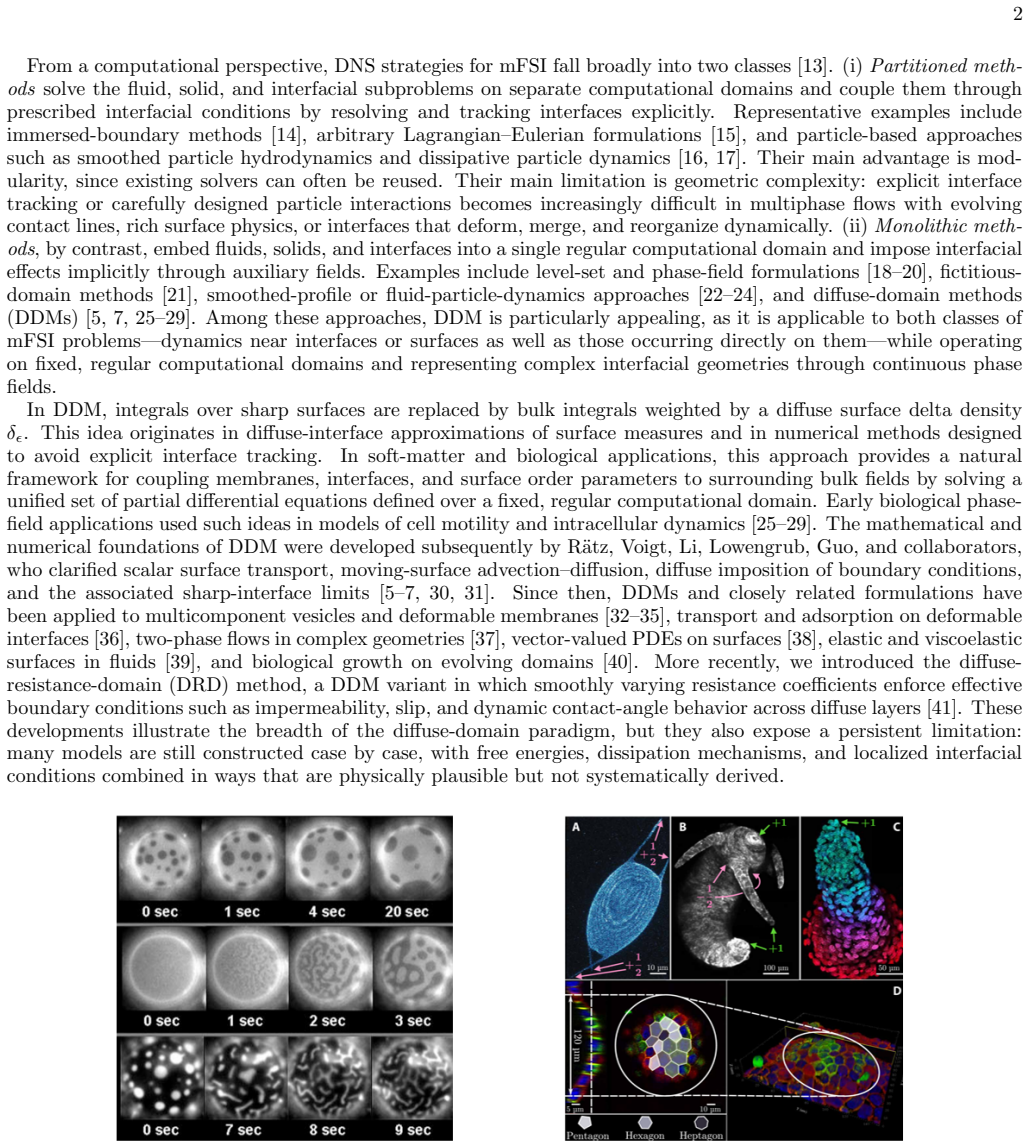
Onsager principle yields unified diffuse-domain models for interfaces
Sharp-surface energies are embedded via delta density to recover sharp limits and produce vesicle and active-shell models.
 full image
full image
Biological Physics
Molecular biophysics, cellular biophysics, neurological biophysics, membrane biophysics, single-molecule biophysics, ecological biophysics, quantum phenomena in biological systems (quantum biophysics), theoretical biophysics, molecular dynamics/modeling and simulation, game theory, biomechanics, bioinformatics, microorganisms, virology, evolution, biophysical methods.
Sharp-surface energies are embedded via delta density to recover sharp limits and produce vesicle and active-shell models.
 full image
full image
Kinetics of Mycoprotein Production from Alternative Carbon Substrates
Screening shows expired functional drink beats pure sugars in speed, biomass titre and reduced byproducts.
Simulations at pH 3.7 show low urea dehydrates the surface while high urea lets water re-enter via self-clustering, leaving secondary folds
 full image
full image
Trapped and flowing nuclei exchange diffusible messages to achieve rapid coordination over long distances, outpacing diffusion up to twenty,
 full image
full image
Growth Dynamics of S. aureus with Sugars and Sugar Alcohols in Weak Magnetic Fields
Weak magnetic fields near 2 mT add only small shifts beyond the pH effect of sweetener metabolism.
Chiral-Induced Spin Selectivity Regulates Triplet formation in Heliobacterial Photosynthesis
Simulations of radical-pair spin dynamics show that chirality-induced selectivity reduces unwanted triplets without needing magnetic fields.
 full image
full image
Cellular-scale mechanism of cell crawling responding to substrate stiffness
Model shows speed and diffusion peak at one stiffness value while persistence rises only on softer substrates.
 full image
full image
Indirect Detection of Lactate Through Voltammetry Using Glassy Carbon Microelectrodes
Lactate oxidase in chitosan on glassy carbon turns the molecule into measurable hydrogen peroxide on flexible neural arrays
 full image
full image
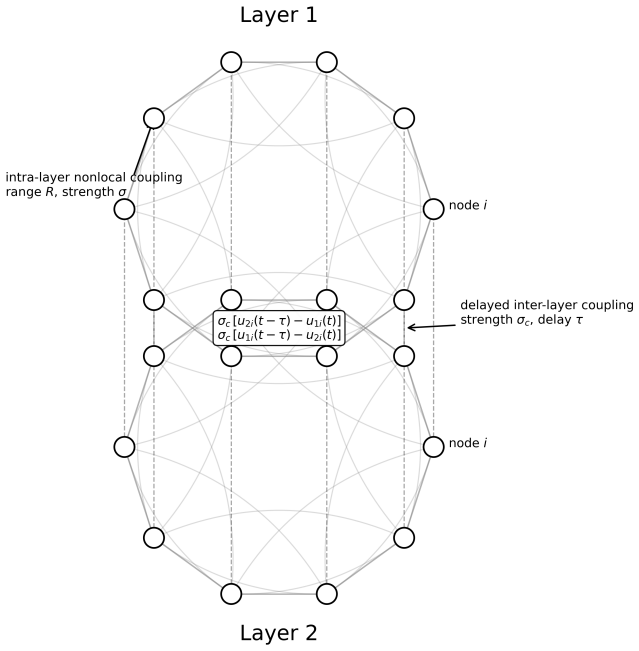
Delay-induced chimera transitions via mode selection in a multiplex FitzHugh Nagumo network
In a two-layer multiplex system, increasing inter-layer delay selectively stabilizes and destabilizes spatial modes, producing partial then全
 full image
full image
Predicting and controlling nonlinear neuro-mechanical locomotion dynamics
Framework turns neural activity recordings into real-time predictions and control signals for C. elegans movement using spectral and decom
 full image
full image
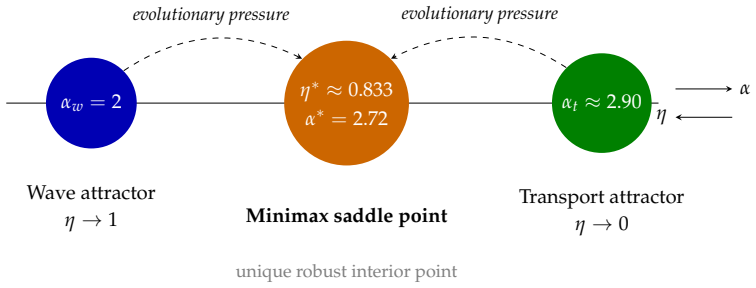
The Incommensurability Principle in Biological Transport
The minimax duty cycle becomes an exact invariant, explaining why the same pattern appears across species and growth stages.
 full image
full image
Optimal information transmission in a sequential model for cell division
A branching process model shows that too few steps make populations unpredictable while too many dilute each step's impact.
 full image
full image
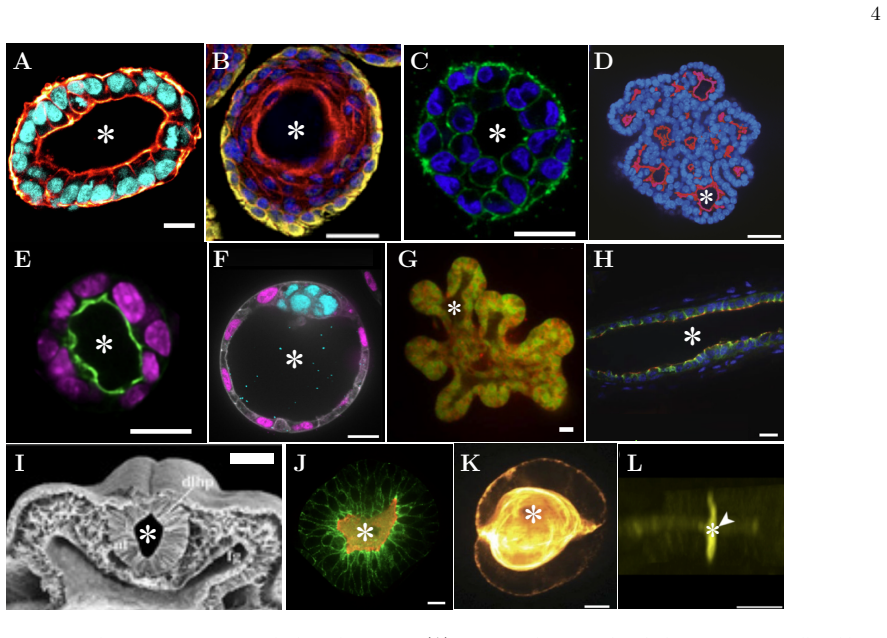
Lumens as active balloons: a biological physics review
Review unifies cavity development across tissues via out-of-equilibrium physics and coupled active processes.
 full image
full image
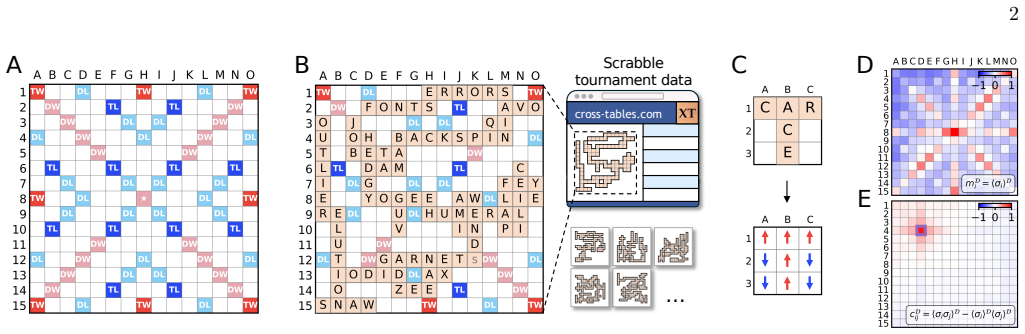
Statistical mechanics for Scrabble predicts strategy, entropy and language
Tile placement statistics alone predict word lengths, reveal strategies, and assign games to languages without seeing the letters.
 full image
full image
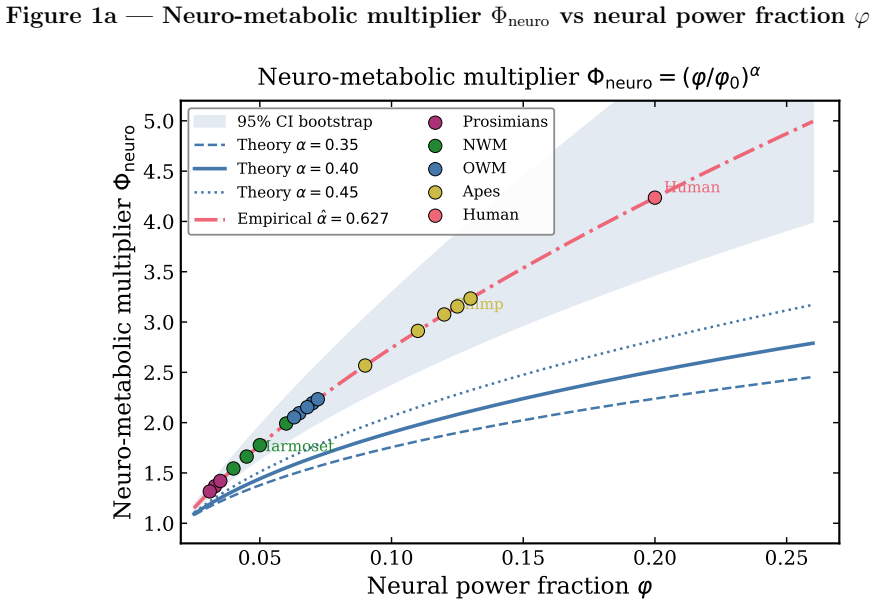
By increasing the share of metabolism going to the brain, primates complete more physiological cycles within the same total entropy budget,
 full image
full image
Dataset spanning mammals, birds and corrected ectotherms shows total cardiac cycles stay near 10^9 after phylogenetic and physiological adj
 full image
full image
Delayed control driven oscillations in plant roots
Minimal model finds period equals four times the delay and matches arc lengths measured in Arabidopsis images.
Statistical tomography shows alternative spatial strategies likely key to transduction success.
 full image
full image
GSM2-based Markov chains for individual cells generate observed survival trends across dose rates and ion types from explicit damage-repair-
 full image
full image
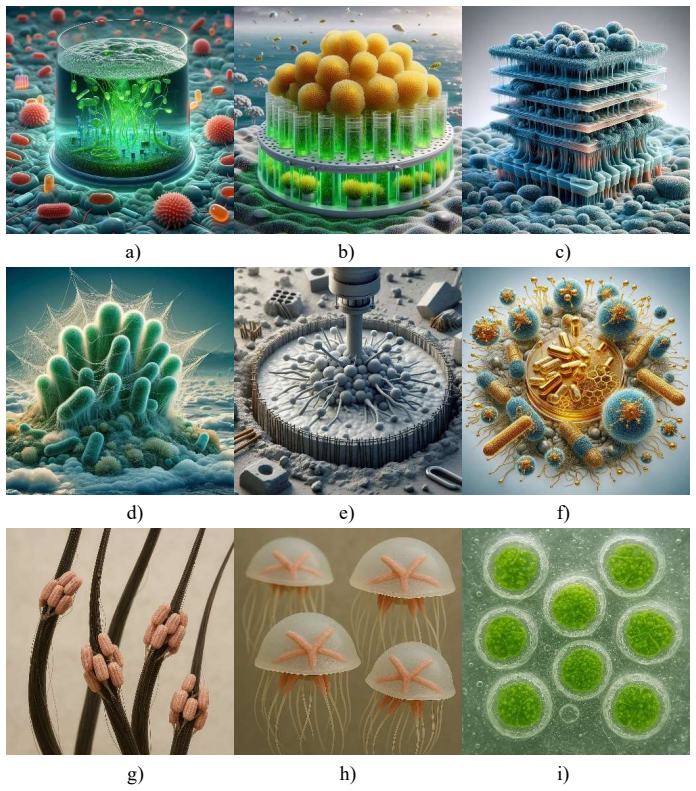
Can we teach generative artificial intelligence the design language of engineered living materials?
A new ontology codifies ELM families, applications and methods so generative AI can describe examples and invent novel bio-materials.
 full image
full image
Orientation-Dependent Protein Binding at Nanoparticle Interfaces
United-atom and docking calculations produce similar angular distributions to experiments for several allergens
 full image
full image
Bayesian Rate Inference for Sequence Motif Dynamics in Systems of Reactive Nucleic Acids
The approach calibrates simplified models to detailed simulations while quantifying uncertainties for RNA world studies.
 full image
full image
Analysis of DNA thermal stability across a broad range of thionine concentrations
Intercalation below 1.5 mg/L gives way to groove and electrostatic binding above it, yet melting temperature rises in both regimes.
The material assembles in place and adapts to brain conditions for stronger adhesion and clearer signals without added conductive particles.
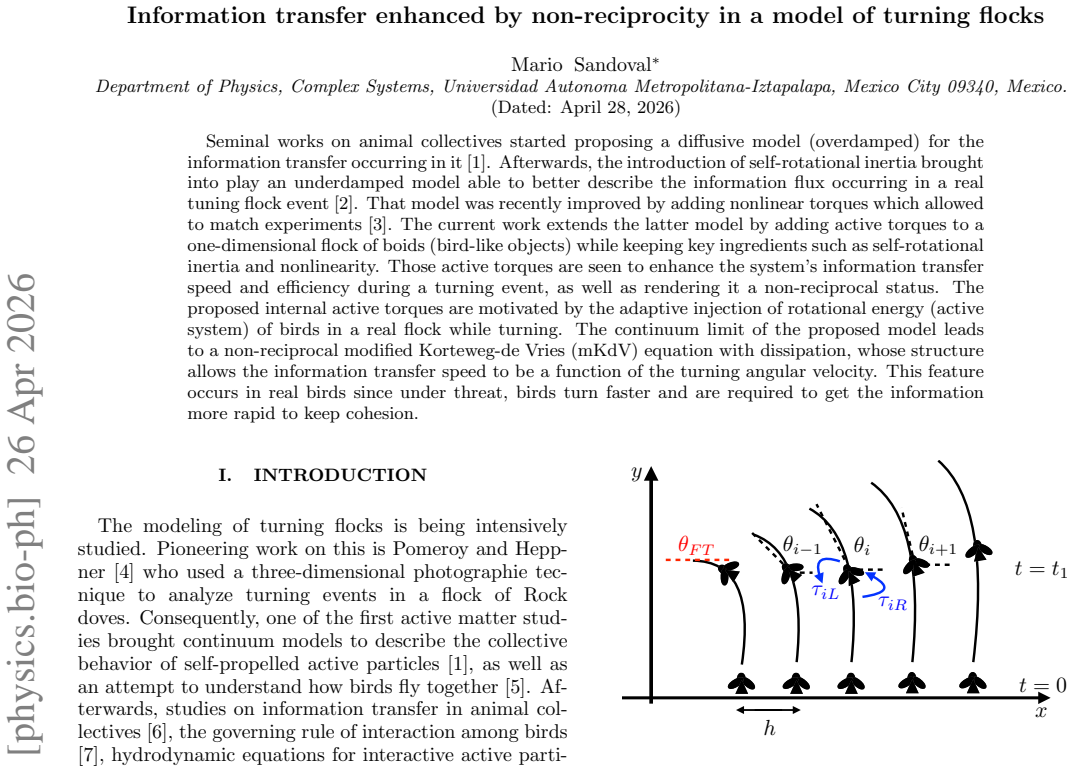
Information transfer enhanced by non-reciprocity in a model of turning flocks
Non-reciprocal effects make propagation speed rise with turning rate, matching how real groups exchange signals faster during sharp turns.
 full image
full image
Planar vector replaces scalar in base-pair model to match entropy, force and rate data.
 full image
full image
Exact Resistance of an Orifice in a 2D Membrane Blocked by a Cylindrical Obstruction
Curvilinear coordinates map all boundaries and yield a closed-form expression usable as access resistance in molecular-sensing channels
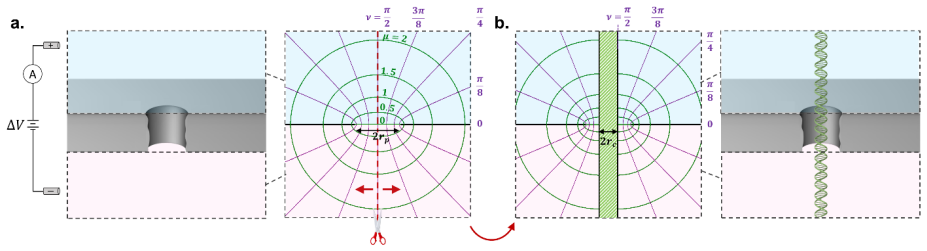 full image
full image
Mitochondrial mechanics nucleates axonal jamming and swelling
Simulations show flexible mitochondria pile up, stressing the axon membrane until it deforms, while rigid ones pass freely.
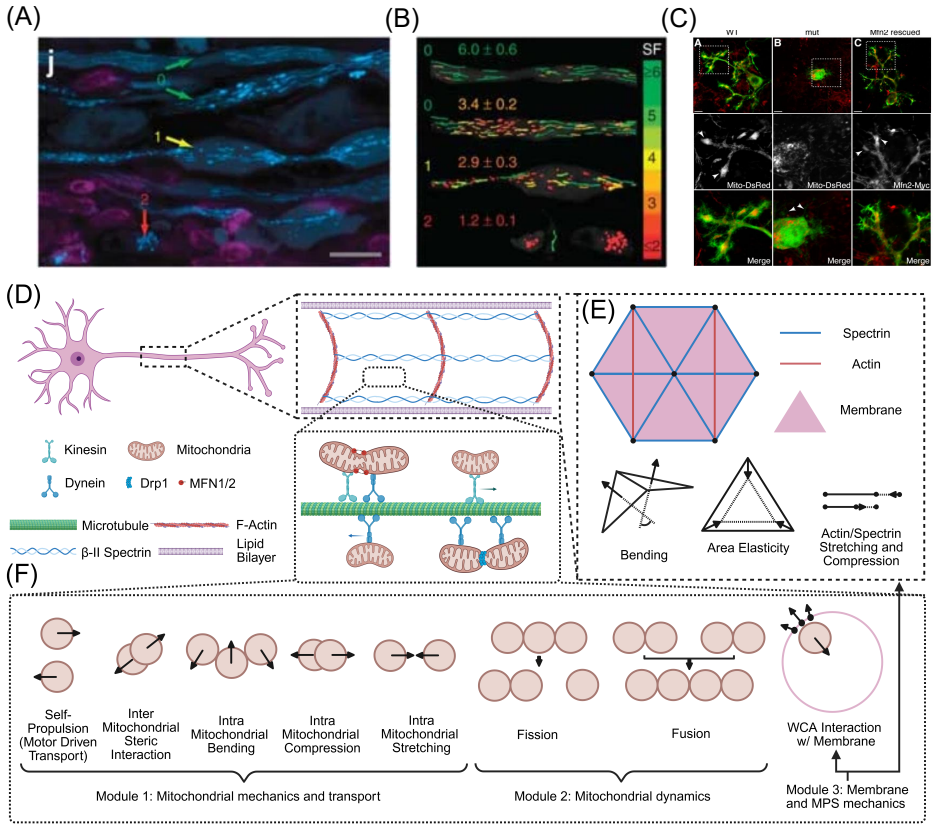 full image
full image
Wave physics as a choreographic notation for partner dance
Dance sequences show interference and emerging harmonics that map to musical dyads, yielding an analytical motion notation.
 full image
full image
Shaping nematic order in bacterial films with single-cell resolution patterning
Parallel spore orientations produce millimetre-scale nematic alignment, synchronous buckling, and light-polarising properties in growing B.
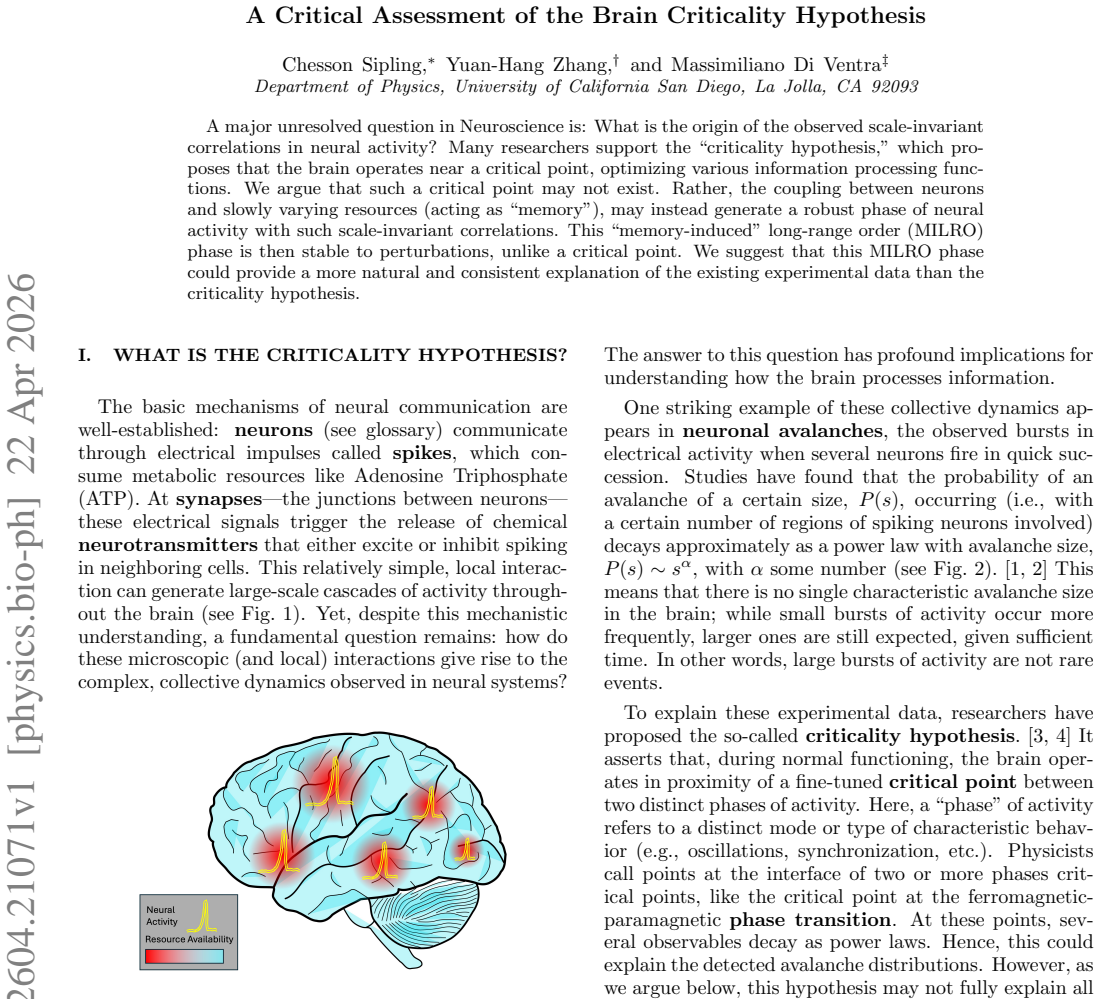
A Critical Assessment of the Brain Criticality Hypothesis
Neuron coupling to slow resources produces stable scale-invariant activity that persists without fine-tuning to a critical point.
 full image
full image
A Critical Assessment of the Brain Criticality Hypothesis
Memory effects from varying resources generate long-range neural correlations more naturally than criticality.
 full image
full image
The transient state loosens the PBL and PL in a pattern unlike the rigid fully reduced form, connecting spin chemistry to navigation signals
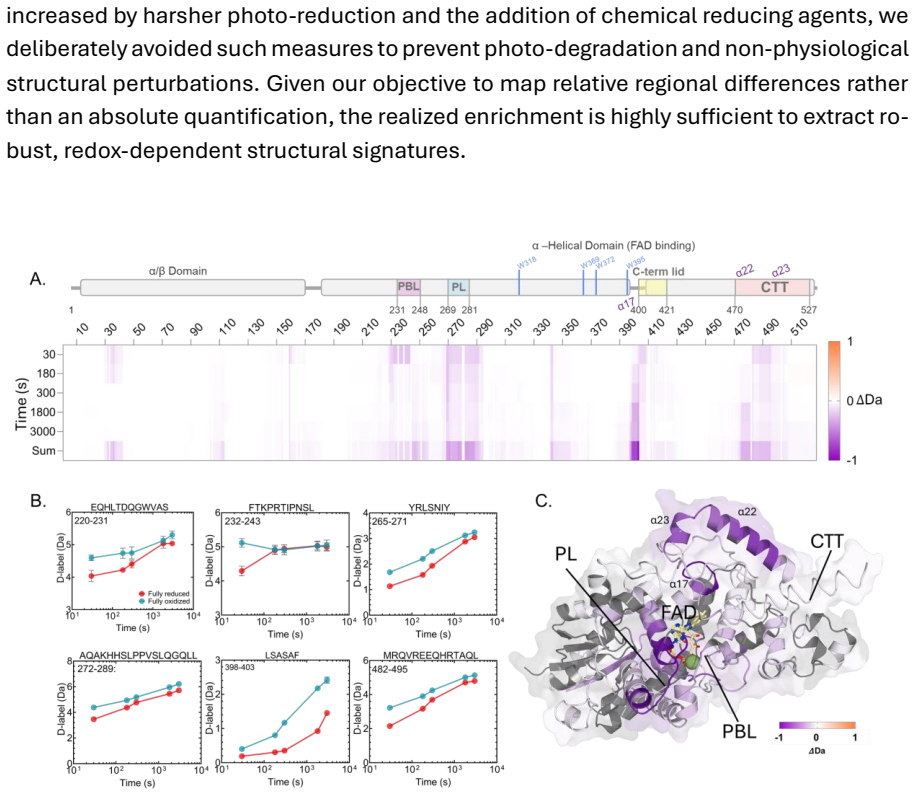 full image
full image
Advancing optical imaging systems with digital fabrication
Open microscopy examples show how 3D-printed parts speed adaptation and allow shared refinements while preserving performance.
Noise-Driven Differentiation via Gene Frustration and Epigenetic Fixation
Logarithmic dependence on noise strength and input-biased fate selection emerge from switching followed by epigenetic locking.
 full image
full image
Deciphering the chemical grammar of protein-RNA condensates
Simulations reveal that base-specific interactions govern phase separation at the smallest chemical scale rather than polymer length alone.
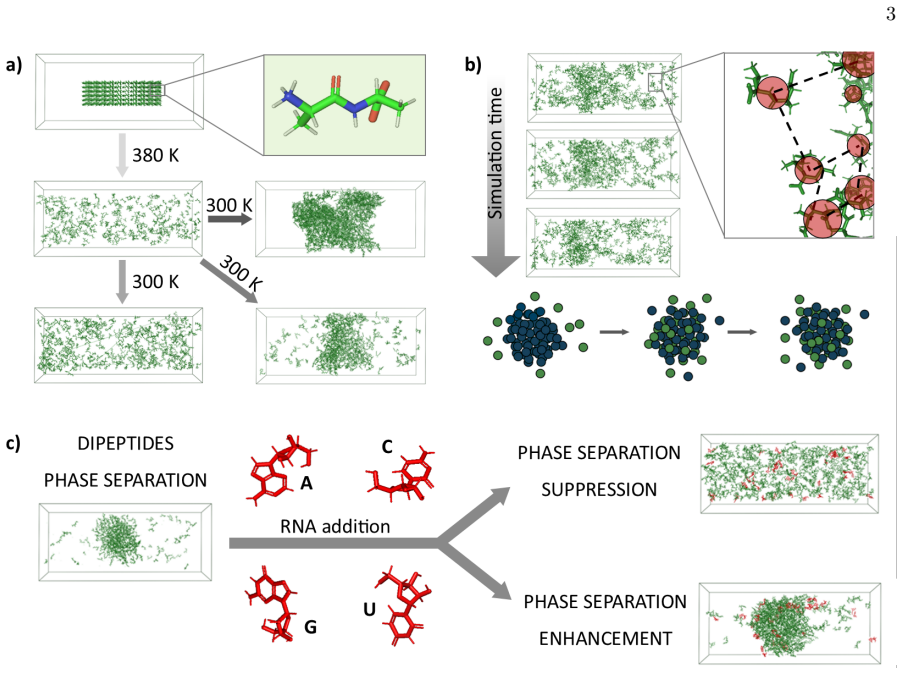 full image
full image
Casimir-Lifshitz forces allow clusters to regulate tasks like approach and stabilization through reproducible attractors and gradients.
 full image
full image
Deformation of Bacterial Cell Membranes by Action of Metal Surface under Plasmon Resonance Condition
Modelling shows surface plasmons expand the van der Waals interaction region with S. aureus cell walls, pointing to mechanical antibacterial
Self-propelled particles driven by light
Experiments and simulations show shape breaking enables net momentum transfer from light, offering a chemical-free drive for microscale act
Spectral map analysis of simulations shows protonation state controls which parts of the SARS-CoV-2 pseudoknot are affected.
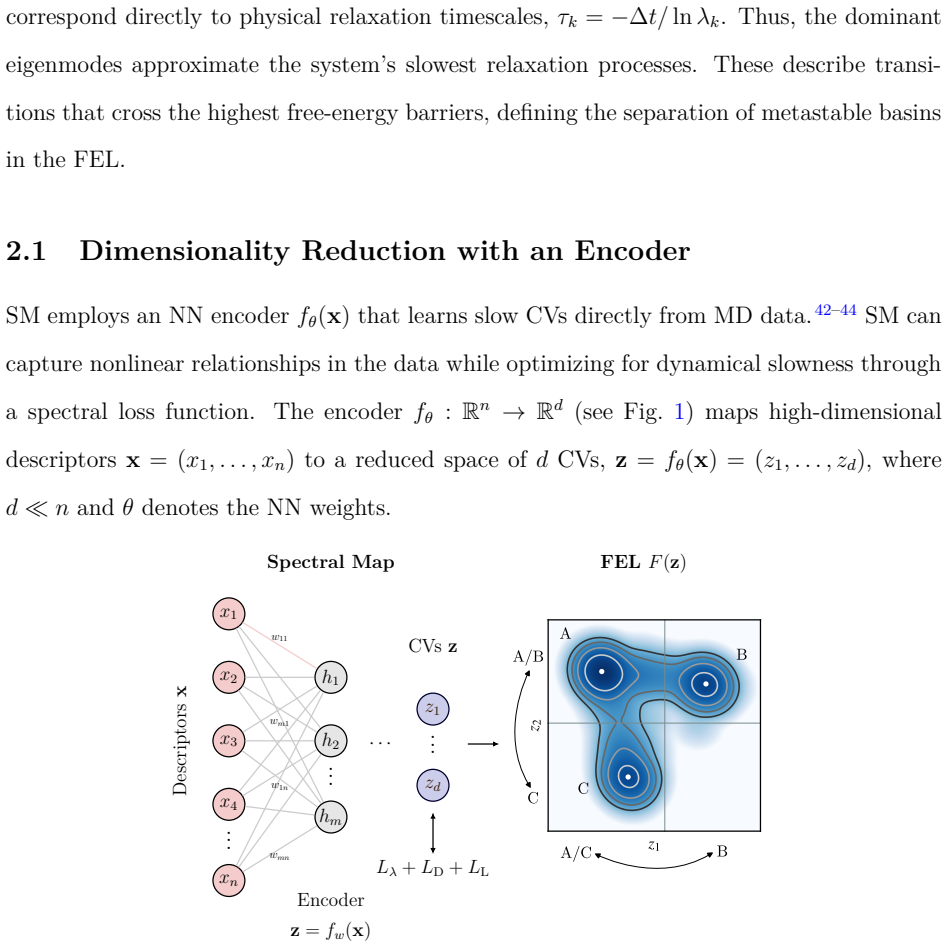 full image
full image
Seabird trajectories map onto a reduced optimal-control bound for dynamic soaring
Normalized data from three seabird species place albatrosses nearest the theoretical minimum derived from a reduced optimal-control model.
 full image
full image
Membrane Tension Governs Particle Wrapping-Unwrapping Transitions and Stalling
Energy calculation reveals when wrapping stalls or reverses based on adhesion, tension, and size.
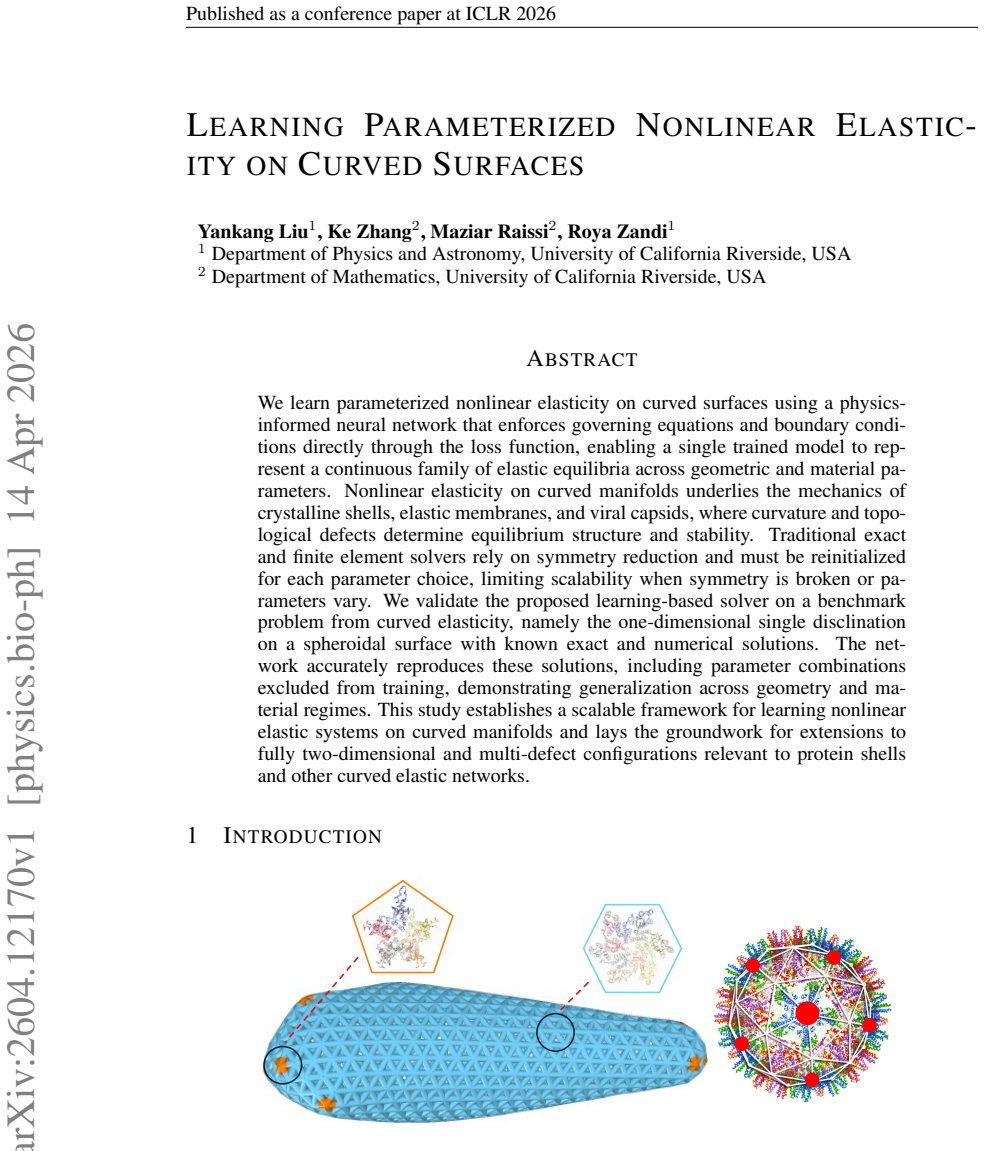
Learning Parameterized Nonlinear Elasticity on Curved Surfaces
One model captures solutions for any geometry or material stiffness by enforcing the nonlinear equations directly in the loss, matching held
 full image
full image
Dynamic Functional Connectivity Resolves Brain Integration-Segregation Trade-off Under Costly Links
Empirical dFC outperforms static networks in reach and speed while preserving local clusters and rapid recirculation.
A toy Hamiltonian yields closed-form singlet populations and reframes zero field as phase locking rather than simple mixing.
The Dynamic Origin of Kleiber's Law
Dynamic impedance matching in proximal vessels sets the exponent and predicts scaling shifts in small animals without free parameters.
 full image
full image
Simulations and experiments demonstrate that altering the tryptophan network modulates emission efficiency, pointing to rules for biological
Active Transport as a Mechanism of Microphase Selection in Biomolecular Condensates
Low-fraction stochastic transport along filaments selects finite sizes tunable from nanometers to micrometers.
 full image
full image
Models with lipid and fibrous plaque show reduced artery opening, higher wall stress and altered blood flow compared to plaque-free cases.
Biogenic bubbles enable microbial escape from physical confinement
Fermentation produces CO2 bubbles that yield the matrix and carry immotile colonies far upward.
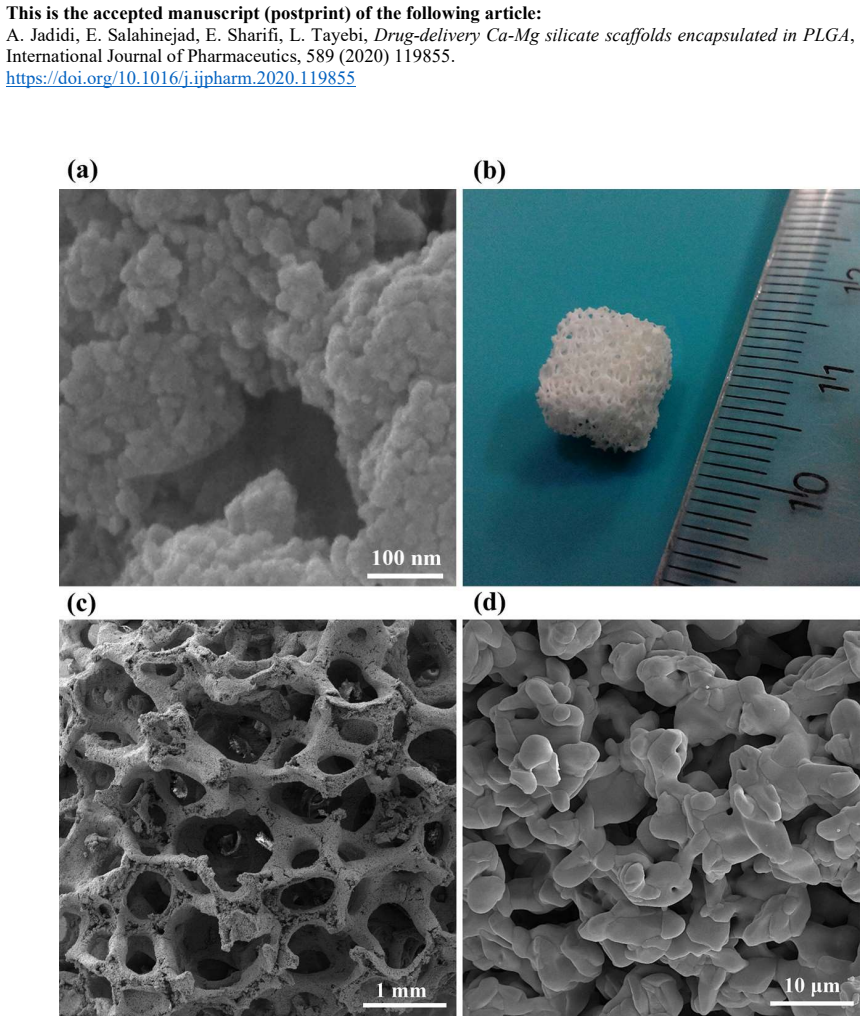
Drug-delivery Ca-Mg silicate scaffolds encapsulated in PLGA
The polymer layer keeps pores open for tissue growth while extending vancomycin delivery and lifting stem-cell survival in lab tests.
 full image
full image
Stem cell viability ranks diopside highest, then akermanite, then bredigite for vancomycin-loaded carriers, indicating carrier breakdown as
 full image
full image
Slovakia's Mass Testing: A Critical Look at the Negative Effects
Re-evaluation finds testing rounds mismatched with case declines and coincided with rising deaths through maintained mobility.
On phenomenology of physical effects in axons
Hybrid models use observables for coupled effects to describe signal propagation in nerve fibres more realistically.
Thermal fluctuations set fundamental limits on ion channel function
Thermal fluctuations from discrete ions set a hard bound on 10 microsecond gating timescales close to observed performance.
 full image
full image
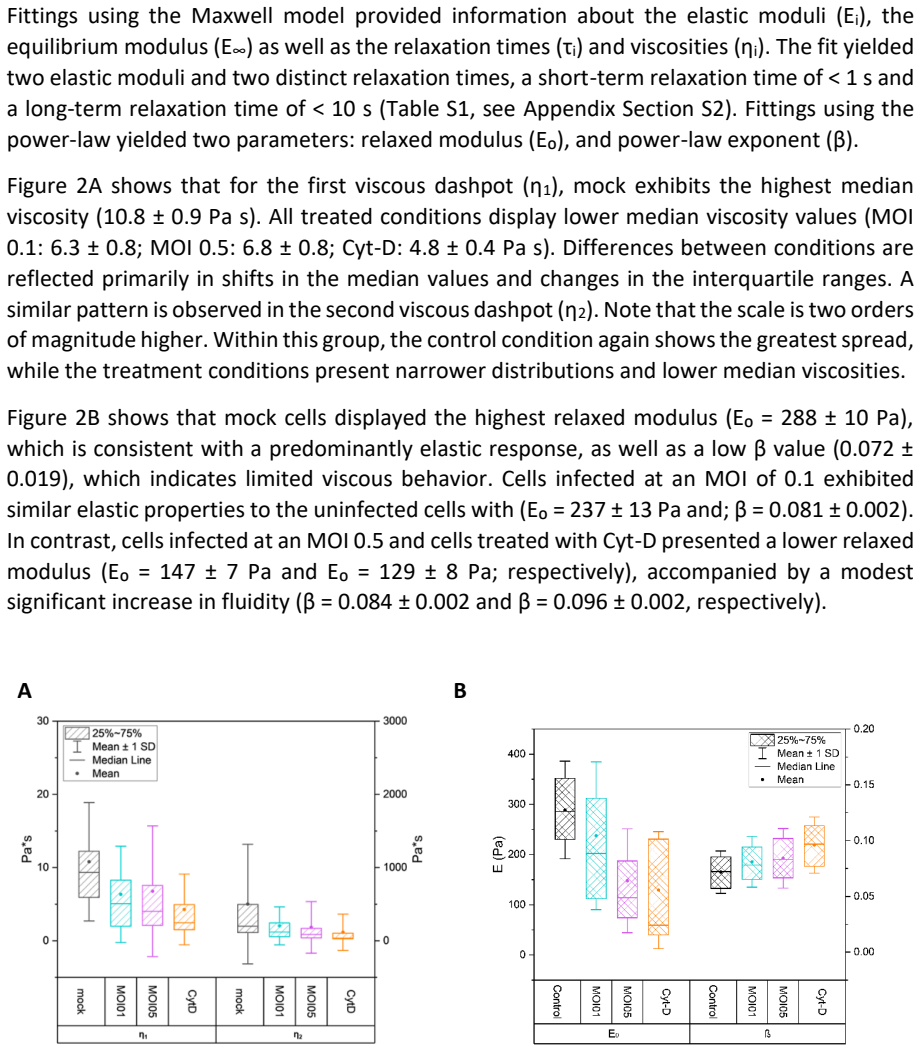
Mechanical Softening of Vero Cells Induced by an Attenuated Measles Vaccine Virus
AFM shows reduced elastic modulus and actin reorganization in infected cells, matching effects of chemical actin depolymerization
 full image
full image