Recognition: unknown
3DPipe: A Pipelined GPU Framework for Scalable Generalized Spatial Join over Polyhedral Objects
Pith reviewed 2026-05-10 00:38 UTC · model grok-4.3
The pith
3DPipe is a pipelined GPU framework that performs scalable spatial joins over 3D polyhedral objects by overlapping CPU preparation, data transfer, and GPU computation while using multi-level pruning and chunked streaming.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
3DPipe exploits GPU parallelism across both filtering and refinement stages, incorporates a multi-level pruning strategy for efficient candidate reduction, and employs chunked streaming to handle datasets exceeding GPU memory; its pipelined execution overlaps CPU data preparation, host-device data transfer, and GPU computation to improve throughput, delivering up to 9.0× speedup over TDBase.
What carries the argument
The pipelined execution model that overlaps CPU data preparation, host-device data transfer, and GPU computation, supported by a multi-level pruning strategy and chunked streaming for polyhedral objects.
If this is right
- Spatial joins become practical for 3D datasets that exceed single-GPU memory limits.
- Both the filter and refinement phases gain from GPU parallelism without custom user-level out-of-core code.
- Overall query throughput rises because data movement latency is hidden behind ongoing computation.
- The approach scales to larger inputs while keeping the same pruning and streaming logic.
Where Pith is reading between the lines
- The same overlap pattern could be applied to other 3D spatial operations such as range queries or k-nearest-neighbor joins.
- Domains that already generate 3D polyhedral data, including object detection from LiDAR point clouds, could see reduced end-to-end processing times.
- If pruning thresholds are made adaptive to object complexity, further candidate reduction may be possible on heterogeneous datasets.
Load-bearing premise
The multi-level pruning and chunked streaming strategy will continue to deliver high candidate reduction and overlap efficiency on arbitrary real-world polyhedral datasets without hidden bottlenecks in data transfer or load imbalance.
What would settle it
Running the system on a large, irregular polyhedral dataset where candidate reduction stays low or transfer overhead dominates, producing either no speedup or out-of-memory failures despite chunking.
Figures















read the original abstract
Spatial join is a fundamental operation in spatial databases. With the rapid growth of 3D data in applications such as LiDAR-based object detection and 3D digital pathology, there is an increasing need to support spatial join over 3D datasets. However, existing techniques are largely designed for 2D data, leaving 3D spatial join underexplored and computationally expensive. We present 3DPipe, a pipelined GPU framework for scalable spatial join over polyhedral objects. 3DPipe exploits GPU parallelism across both filtering and refinement stages, incorporates a multi-level pruning strategy for efficient candidate reduction, and employs chunked streaming to handle datasets exceeding GPU memory. Its pipelined execution overlaps CPU data preparation, host-device data transfer, and GPU computation to improve throughput. Experiments show that 3DPipe achieves up to 9.0$\times$ speedup over the state-of-the-art GPU solution, TDBase, while maintaining excellent scalability. 3DPipe is open-sourced at https://github.com/lyuheng/3dpipe.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper presents 3DPipe, a pipelined GPU framework for scalable generalized spatial joins over 3D polyhedral objects. It exploits GPU parallelism for both filtering and refinement, incorporates multi-level pruning for candidate reduction, uses chunked streaming to process datasets larger than GPU memory, and overlaps CPU data preparation, host-device transfers, and GPU computation via pipelining. The central empirical claim is that 3DPipe delivers up to 9.0× speedup over the prior GPU baseline TDBase while exhibiting strong scalability; the implementation is released as open source.
Significance. If the reported speedups and scalability hold under rigorous evaluation, the work fills an important gap in spatial database systems by extending efficient GPU-accelerated joins to 3D polyhedra, which are relevant to growing applications such as LiDAR processing and 3D digital pathology. The open-source release and falsifiable performance claims constitute a clear strength for reproducibility and further research.
major comments (2)
- [§5] §5 (Experimental Evaluation): The abstract and results claim up to 9.0× speedup and 'excellent scalability,' yet the provided text contains no dataset descriptions, table of input sizes, error bars, number of runs, or ablation studies isolating the contribution of pipelining versus multi-level pruning. This absence prevents verification of the central performance claim and should be addressed with concrete experimental details.
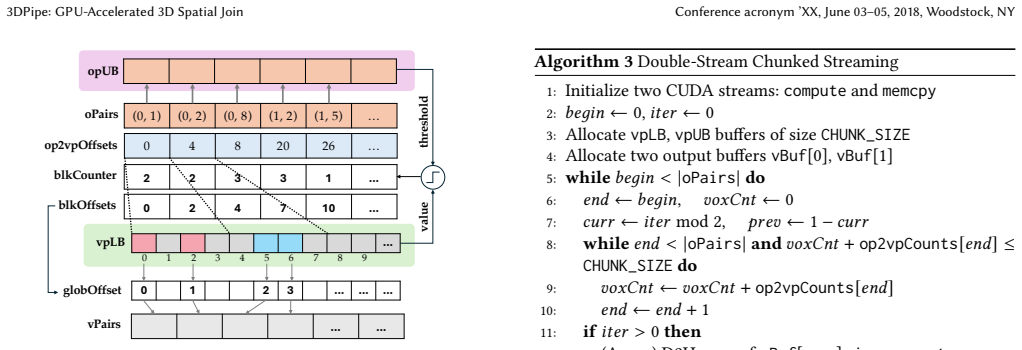
- [§3.3] §3.3 (Chunked Streaming): The description of chunked streaming for out-of-core datasets does not quantify potential host-device transfer overhead or load imbalance across polyhedral objects of varying complexity; without such analysis or measurements, it is unclear whether the strategy remains efficient on arbitrary real-world distributions as assumed in the weakest point of the evaluation.
minor comments (2)
- [Abstract] The abstract and introduction would benefit from a brief comparison table or sentence contrasting 3DPipe's design choices with TDBase to clarify the source of the reported gains.
- [§3] Notation for polyhedral representation (e.g., how faces and edges are encoded for GPU kernels) is introduced without a dedicated figure or pseudocode example, which would aid clarity for readers unfamiliar with 3D spatial data structures.
Simulated Author's Rebuttal
We thank the referee for the constructive feedback and the recommendation of minor revision. We appreciate the emphasis on strengthening the experimental evaluation and the analysis of chunked streaming. We address each major comment below and will revise the manuscript accordingly.
read point-by-point responses
-
Referee: [§5] §5 (Experimental Evaluation): The abstract and results claim up to 9.0× speedup and 'excellent scalability,' yet the provided text contains no dataset descriptions, table of input sizes, error bars, number of runs, or ablation studies isolating the contribution of pipelining versus multi-level pruning. This absence prevents verification of the central performance claim and should be addressed with concrete experimental details.
Authors: We agree that the current experimental section lacks sufficient detail for independent verification of the reported speedups and scalability. In the revised manuscript, we will add a table summarizing all datasets with their sizes, object counts, and complexity metrics. We will explicitly state that all timing results are averaged over 5 runs and include error bars in the figures. We will also add an ablation study that isolates the performance contributions of pipelining, multi-level pruning, and chunked streaming. revision: yes
-
Referee: [§3.3] §3.3 (Chunked Streaming): The description of chunked streaming for out-of-core datasets does not quantify potential host-device transfer overhead or load imbalance across polyhedral objects of varying complexity; without such analysis or measurements, it is unclear whether the strategy remains efficient on arbitrary real-world distributions as assumed in the weakest point of the evaluation.
Authors: We acknowledge the need for quantitative analysis of transfer overhead and load imbalance. In the revision, we will include new measurements breaking down the time spent in host-device transfers versus GPU computation for different chunk sizes. We will also add a discussion and supporting experiments on load imbalance for datasets containing polyhedra of heterogeneous complexity, along with any mitigation strategies employed. revision: yes
Circularity Check
No significant circularity in empirical systems paper
full rationale
The paper describes a pipelined GPU framework for 3D spatial joins over polyhedral objects, with claims resting entirely on measured experimental speedups (up to 9.0× over TDBase) and scalability observations. No mathematical derivation chain, fitted parameters, predictions, or equations are present that could reduce to inputs by construction. The approach relies on implementation techniques (multi-level pruning, chunked streaming, pipelined execution) whose performance is externally falsifiable via runtime benchmarks on real datasets. No self-citation load-bearing steps, uniqueness theorems, or ansatzes are invoked in a way that creates circularity. This is a standard empirical systems contribution whose validity depends on reproducible experiments rather than internal definitional reductions.
Axiom & Free-Parameter Ledger
Reference graph
Works this paper leans on
-
[1]
[n. d.]. Cesium. https://cesium.com
-
[2]
[n. d.]. ERSI 3D GIS. https://www.esri.com/en-us/capabilities/3d-gis
-
[3]
[n. d.]. ModelNet40 - Princeton 3D Object Dataset. https://www.kaggle.com/dat asets/balraj98/modelnet40-princeton-3d-object-dataset
-
[4]
[n. d.]. OFF (Object File Format). https://en.wikipedia.org/wiki/OFF_(file_format )
-
[5]
[n. d.]. Polyhedron Model in PolarDB. https://www.alibabacloud.com/help/en/p olardb/polardb-for-postgresql/models-pg
-
[6]
[n. d.]. PostGIS. https://postgis.net/
-
[7]
[n. d.]. TDBase Source Code. https://github.com/tengdj/tdbase
-
[8]
Ablimit Aji, Fusheng Wang, Hoang Vo, Rubao Lee, Qiaoling Liu, Xiaodong Zhang, and Joel Saltz. 2013. Hadoop-GIS: A high performance spatial data warehousing system over MapReduce. InProceedings of the VLDB endowment international conference on very large data bases, Vol. 6. p1009
2013
-
[9]
Lars Arge, Octavian Procopiuc, Sridhar Ramaswamy, Torsten Suel, and Jef- frey Scott Vitter. 1998. Scalable sweeping-based spatial join. InVLDB, Vol. 98. 570–581
1998
-
[10]
Furqan Baig, Hoang Vo, Tahsin Kurc, Joel Saltz, and Fusheng Wang. 2017. Sparkgis: Resource aware efficient in-memory spatial query processing. InPro- ceedings of the 25th ACM SIGSPATIAL international conference on advances in geographic information systems. 1–10
2017
-
[11]
Rushmeier, Cláudio T
Fausto Bernardini, Joshua Mittleman, Holly E. Rushmeier, Cláudio T. Silva, and Gabriel Taubin. 1999. The Ball-Pivoting Algorithm for Surface Reconstruction. IEEE Trans. Vis. Comput. Graph.5, 4 (1999), 349–359
1999
-
[12]
Thomas Brinkhoff, H-P Kriegel, and Bernhard Seeger. 1996. Parallel processing of spatial joins using R-trees. InProceedings of the Twelfth International Conference on Data Engineering. IEEE, 258–265
1996
-
[13]
Lang, Sourabh Vora, Venice Erin Liong, Qiang Xu, Anush Krishnan, Yu Pan, Giancarlo Baldan, and Oscar Beijbom
Holger Caesar, Varun Bankiti, Alex H. Lang, Sourabh Vora, Venice Erin Liong, Qiang Xu, Anush Krishnan, Yu Pan, Giancarlo Baldan, and Oscar Beijbom. 2020. nuScenes: A Multimodal Dataset for Autonomous Driving. InCVPR. Computer Vision Foundation / IEEE, 11618–11628
2020
-
[14]
Xingyu Chen, Fu-Jen Chu, Pierre Gleize, Kevin J. Liang, Alexander Sax, Hao Tang, Weiyao Wang, Michelle Guo, Thibaut Hardin, Xiang Li, Aohan Lin, Jiawei Liu, Ziqi Ma, Anushka Sagar, Bowen Song, Xiaodong Wang, Jianing Yang, Bowen Zhang, Piotr Dollár, Georgia Gkioxari, Matt Feiszli, and Jitendra Malik. 2025. SAM 3D: 3Dfy Anything in Images.arXiv preprint arX...
work page internal anchor Pith review arXiv 2025
-
[15]
HuBMAP Consortium. 2019. The human body at cellular resolution: the NIH Human Biomolecular Atlas Program.Nature574, 7777 (2019), 187–192
2019
-
[16]
Ahmed Eldawy and Mohamed F Mokbel. 2015. Spatialhadoop: A mapreduce framework for spatial data. In2015 IEEE 31st international conference on Data Engineering. IEEE, 1352–1363
2015
-
[17]
Navid Farahani, Alex Braun, Dylan Jutt, Todd Huffman, Nick Reder, Zheng Liu, Yukako Yagi, and Liron Pantanowitz. 2017. Three-dimensional imaging and scanning: current and future applications for pathology.Journal of pathology informatics8, 1 (2017), 36
2017
-
[18]
Andreas Geiger, Philip Lenz, and Raquel Urtasun. 2012. Are we ready for au- tonomous driving? The KITTI vision benchmark suite. InCVPR. IEEE Computer Society, 3354–3361
2012
-
[19]
S Gnanakaran, Hugh Nymeyer, John Portman, Kevin Y Sanbonmatsu, and Angel E Garcıa. 2003. Peptide folding simulations.Current opinion in structural biology 13, 2 (2003), 168–174
2003
- [20]
-
[21]
W. Daniel Hillis and Guy L. Steele Jr. 1986. Data Parallel Algorithms.Commun. ACM29, 12 (1986), 1170–1183. doi:10.1145/7902.7903
-
[22]
Wenqi Jiang, Oleh-Yevhen Khavrona, Martin Parvanov, and Gustavo Alonso
-
[23]
Swiftspatial: Spatial joins on modern hardware.Proceedings of the ACM on Management of Data3, 3 (2025), 1–27
2025
-
[24]
Kazhdan, Matthew Bolitho, and Hugues Hoppe
Michael M. Kazhdan, Matthew Bolitho, and Hugues Hoppe. 2006. Poisson surface reconstruction. InProceedings of the Fourth Eurographics Symposium on Geometry Processing (ACM International Conference Proceeding Series), Alla Sheffer and Konrad Polthier (Eds.). Eurographics Association, 61–70
2006
-
[25]
Andrei Khodakovsky, Peter Schröder, and Wim Sweldens. 2000. Progressive geometry compression. InProceedings of the 27th annual conference on Computer graphics and interactive techniques. 271–278
2000
-
[26]
Yanhui Liang, Hoang Vo, Jun Kong, and Fusheng Wang. 2017. iSPEED: an Efficient In-Memory Based Spatial Query System for Large-Scale 3D Data with Complex Structures. InProceedings of the 25th ACM SIGSPATIAL International Conference on Advances in Geographic Information Systems, GIS 2017, Redondo Beach, CA, USA, November 7-10, 2017. ACM, 17:1–17:10
2017
-
[27]
Yanhui Liang, Fusheng Wang, Pengyue Zhang, Joel H Saltz, Daniel J Brat, and Jun Kong. 2017. Development of a framework for large scale three-dimensional pathology and biomarker imaging and spatial analytics.AMIA Summits on Translational Science Proceedings2017 (2017), 75
2017
-
[28]
Ming-Ling Lo and Chinya V Ravishankar. 1996. Spatial hash-joins. InProceedings of the 1996 ACM SIGMOD international conference on Management of data. 247– 258
1996
-
[29]
Lorensen and Harvey E
William E. Lorensen and Harvey E. Cline. 1987. Marching cubes: A high resolution 3D surface construction algorithm. InSIGGRAPH, Maureen C. Stone (Ed.). ACM, 163–169
1987
-
[30]
Adrien Maglo, Clement Courbet, Pierre Alliez, and Céline Hudelot. 2012. Pro- gressive compression of manifold polygon meshes.Comput. Graph.36, 5 (2012), 349–359
2012
-
[31]
Adrien Maglo, Guillaume Lavoué, Florent Dupont, and Céline Hudelot. 2015. 3D Mesh Compression: Survey, Comparisons, and Emerging Trends.ACM Comput. Surv.47, 3 (2015), 44:1–44:41
2015
-
[32]
Srinivasan, Matthew Tancik, Jonathan T
Ben Mildenhall, Pratul P. Srinivasan, Matthew Tancik, Jonathan T. Barron, Ravi Ramamoorthi, and Ren Ng. 2020. NeRF: Representing Scenes as Neural Radiance Fields for View Synthesis. InECCV (Lecture Notes in Computer Science), Andrea Vedaldi, Horst Bischof, Thomas Brox, and Jan-Michael Frahm (Eds.). Springer, 405–421
2020
-
[33]
Tomas Möller. 1997. A Fast Triangle-Triangle Intersection Test.J. Graphics, GPU, & Game Tools2, 2 (1997), 25–30. 3DPipe: GPU-Accelerated 3D Spatial Join Conference acronym ’XX, June 03–05, 2018, Woodstock, NY
1997
-
[34]
Sadegh Nobari, Farhan Tauheed, Thomas Heinis, Panagiotis Karras, Stéphane Bressan, and Anastasia Ailamaki. 2013. TOUCH: in-memory spatial join by hierarchical data-oriented partitioning. InProceedings of the ACM SIGMOD Inter- national Conference on Management of Data, SIGMOD 2013, New York, NY, USA, June 22-27, 2013. ACM, 701–712
2013
-
[35]
Florence, Julian Straub, Richard A
Jeong Joon Park, Peter R. Florence, Julian Straub, Richard A. Newcombe, and Steven Lovegrove. 2019. DeepSDF: Learning Continuous Signed Distance Func- tions for Shape Representation. InCVPR. Computer Vision Foundation / IEEE, 165–174
2019
-
[36]
Patel and David J
Jignesh M. Patel and David J. DeWitt. 1996. Partition Based Spatial-Merge Join. InProceedings of the 1996 ACM SIGMOD International Conference on Management of Data, Montreal, Quebec, Canada, June 4-6, 1996. ACM Press, 259–270
1996
-
[37]
Lucas C Villa Real, Bruno Silva, Dikran S Meliksetian, and Kaique Sacchi. 2019. Large-scale 3D geospatial processing made possible. InProceedings of the 27th ACM SIGSPATIAL International Conference on Advances in Geographic Information Systems. 199–208
2019
-
[38]
Nick Roussopoulos, Stephen Kelley, and Frédéic Vincent. 1995. Nearest Neighbor Queries. InSIGMOD, Michael J. Carey and Donovan A. Schneider (Eds.). ACM Press, 71–79
1995
-
[39]
Dejun Teng, Furqan Baig, Zhaohui Peng, Jun Kong, and Fusheng Wang. 2024. Efficient spatial queries over complex polygons with hybrid representations. GeoInformatica28, 3 (2024), 459–497
2024
-
[40]
Dejun Teng, Furqan Baig, Hoang Vo, Yanhui Liang, Jun Kong, and Fusheng Wang
-
[41]
InProceedings of the 25th International Conference on Extending Database Technology, EDBT 2022, Edinburgh, UK, March 29 - April 1,
3DPro: Querying Complex Three-Dimensional Data with Progressive Compression and Refinement. InProceedings of the 25th International Conference on Extending Database Technology, EDBT 2022, Edinburgh, UK, March 29 - April 1,
2022
-
[42]
Dejun Teng, Zhaochuan Li, Zhaohui Peng, Shuai Ma, and Fusheng Wang. 2025. Efficient and Accurate Spatial Queries Using Lossy Compressed 3D Geometry Data.IEEE Trans. Knowl. Data Eng.37, 5 (2025), 2472–2487
2025
-
[43]
Dejun Teng, Yanhui Liang, Hoang Vo, Jun Kong, and Fusheng Wang. 2022. Effi- cient 3D Spatial Queries for Complex Objects.ACM Trans. Spatial Algorithms Syst.8, 2 (2022), 1–26
2022
-
[44]
Haithem Turki, Deva Ramanan, and Mahadev Satyanarayanan. 2022. Mega-NeRF: Scalable Construction of Large-Scale NeRFs for Virtual Fly-Throughs. InCVPR. IEEE, 12912–12921
2022
-
[45]
Sébastien Valette, Raphaëlle Chaine, and Rémy Prost. 2009. Progressive lossless mesh compression via incremental parametric refinement. InComputer Graphics Forum, Vol. 28. 1301–1310
2009
-
[46]
Fusheng Wang, Jun Kong, Lee Cooper, Tony Pan, Tahsin Kurc, Wenjin Chen, Ashish Sharma, Cristobal Niedermayr, Tae W Oh, Daniel Brat, et al. 2011. A data model and database for high-resolution pathology analytical image informatics. Journal of pathology informatics2, 1 (2011), 32
2011
-
[47]
World Labs. [n. d.]. Marble Labs: Blueprints for Building with World Models. https://www.worldlabs.ai/labs
-
[48]
Zhirong Wu, Shuran Song, Aditya Khosla, Fisher Yu, Linguang Zhang, Xiaoou Tang, and Jianxiong Xiao. 2015. 3d shapenets: A deep representation for volumet- ric shapes. InProceedings of the IEEE conference on computer vision and pattern recognition. 1912–1920
2015
-
[49]
Jia Yu, Jinxuan Wu, and Mohamed Sarwat. 2015. GeoSpark: a cluster computing framework for processing large-scale spatial data. InProceedings of the 23rd SIGSPATIAL International Conference on Advances in Geographic Information Systems, Bellevue, W A, USA, November 3-6, 2015. ACM, 70:1–70:4
2015
-
[50]
Lyuheng Yuan, Da Yan, Akhlaque Ahmad, Jiao Han, Saugat Adhikari, and Yang Zhou. 2025. Out-of-Core Parallel Spatial Join Outperforming In-Memory Systems: A BFS-DFS Hybrid Approach. InProceedings of the 34th International Symposium on High-Performance Parallel and Distributed Computing, HPDC 2025, Notre Dame, IN, USA, July 20-23, 2025. ACM, 23:1–23:14
2025
-
[51]
Andi Zang, Shiyu Luo, Xin Chen, and Goce Trajcevski. 2019. Real-Time Appli- cations Using High Resolution 3D Objects in High Definition Maps (Systems Paper). InSIGSPATIAL. ACM, 229–238
2019
-
[52]
Jianting Zhang, Simin You, and Le Gruenwald. 2017. Parallel selectivity estimation for optimizing multidimensional spatial join processing on gpus. In2017 IEEE 33rd International Conference on Data Engineering (ICDE). IEEE, 1591–1598
2017
-
[53]
Rui Zhang, Jianzhong Qi, Dan Lin, Wei Wang, and Raymond Chi-Wing Wong
-
[54]
A highly optimized algorithm for continuous intersection join queries over moving objects.VLDB J.21, 4 (2012), 561–586
2012
-
[55]
Xiaofang Zhou, David J Abel, and David Truffet. 1998. Data partitioning for parallel spatial join processing.Geoinformatica2, 2 (1998), 175–204
1998
-
[56]
Yuchen Zhou, Jiayuan Gu, Tung Yen Chiang, Fanbo Xiang, and Hao Su. 2025. Point-SAM: Promptable 3D Segmentation Model for Point Clouds. InICLR. Open- Review.net
2025
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.