Recognition: unknown
Bayesian Sparsity Modeling of Shared Neural Response in Functional Magnetic Resonance Imaging Data
Pith reviewed 2026-05-08 13:17 UTC · model grok-4.3
The pith
A Bayesian model using sparse Gaussian processes and horseshoe priors detects shared neural responses in fMRI while quantifying uncertainty.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The authors develop a Bayesian model that simultaneously detects spatial regions of shared activity and estimates the shared neural response function by combining sparse Gaussian process estimation with a horseshoe-inspired Bayesian sparsity prior. A spatially structured extension encourages similar activation patterns in neighboring voxels. When applied to task-based and naturalistic fMRI data, the model provides principled uncertainty quantification for the shared response and demonstrates improved or comparable activation detection and response estimation relative to intersubject correlation methods.
What carries the argument
Sparse Gaussian process estimation of the shared response function combined with a horseshoe-inspired Bayesian sparsity prior that selects active voxels while enabling uncertainty quantification.
If this is right
- Direct estimation of the shared neural response function instead of relying solely on correlation summaries.
- Improved activation detection without performing thousands of separate voxelwise tests.
- Principled uncertainty quantification for both voxel activation and the estimated response function.
- Comparable or superior performance to intersubject correlation on both task-based and naturalistic fMRI datasets.
- Framework that supports potential clinical biomarker identification through modeled synchronous responses.
Where Pith is reading between the lines
- The uncertainty estimates could help evaluate reliability when applying the model to patient groups for biomarker discovery.
- The spatial prior might extend naturally to connectivity analyses in other modalities such as EEG.
- The model could serve as a building block for stimulus decoding tasks that require interpretable shared signals.
- Direct comparisons with flexible machine learning alternatives would clarify trade-offs between uncertainty calibration and predictive accuracy.
Load-bearing premise
That the combination of sparse Gaussian process estimation and the horseshoe-inspired Bayesian sparsity prior will correctly separate shared signal from noise and individual variability without introducing bias or missing true activations in real fMRI data.
What would settle it
A controlled simulation with known shared response signals, added noise, and individual variability where the model either misses true activations, produces intervals that fail to cover the true function, or shows worse activation detection metrics than intersubject correlation.
Figures







read the original abstract
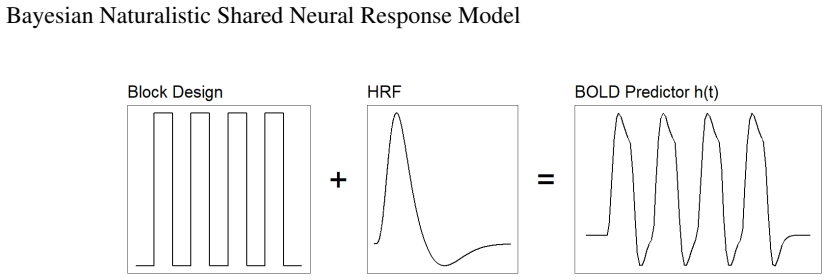
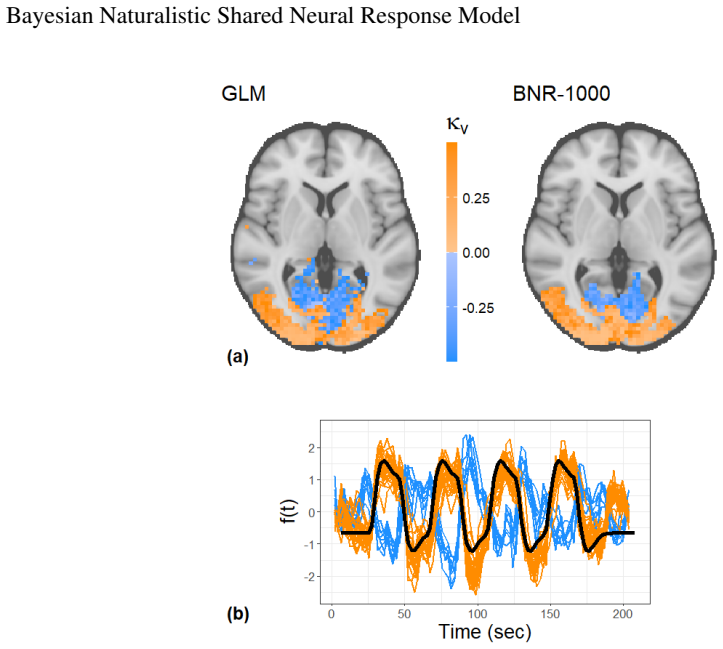
Detecting shared neural activity from functional magnetic resonance imaging (fMRI) across individuals exposed to the same stimulus can reveal synchronous brain responses, functional roles of regions, and potential clinical biomarkers. Intersubject correlation (ISC) is the main method for identifying voxelwise shared responses and per-subject variability, but it relies on heavy data summarization and thousands of regional tests, leading to poor uncertainty quantification and multiple testing issues. ISC also does not directly estimate a shared neural response (SNR) function. We propose a model-based alternative applicable to both task-based and naturalistic fMRI that simultaneously identifies spatial regions of shared activity and estimates the SNR function. The model combines sparse Gaussian process estimation of the response function with a Bayesian sparsity prior inspired by the horseshoe prior to detect voxel activation. A spatially structured extension encourages neighboring voxels to exhibit similar activation patterns. We examine the model's properties, evaluate performance via simulations, and analyze two real-world fMRI datasets, including one task-based and one naturalistic dataset. The Bayesian framework provides principled uncertainty quantification for the shared response function and shows improved activation detection and response estimation compared to standard approaches. Model fits demonstrate comparable or superior performance relative to ISC, while the framework opens avenues for clinical applications.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proposes a Bayesian sparsity model for detecting and estimating shared neural responses (SNR) in fMRI data across subjects exposed to the same stimulus. It integrates sparse Gaussian process estimation of the response function with a horseshoe-inspired prior for voxel activation detection, plus a spatially structured extension, and evaluates the approach via model properties, simulations with ground truth, and applications to one task-based and one naturalistic real fMRI dataset. The framework is positioned as providing principled posterior uncertainty on the shared response while matching or exceeding ISC in activation detection and response estimation.
Significance. If the simulation and real-data results hold under scrutiny, the work supplies a coherent model-based alternative to ISC that directly estimates the shared response function and supplies posterior uncertainty quantification, addressing ISC's reliance on heavy summarization and multiple-testing burdens. The combination of established components (sparse GPs, horseshoe prior) with explicit checks against known ground truth and two distinct fMRI paradigms constitutes a substantive contribution to statistical neuroimaging methodology.
major comments (1)
- §4 (Simulations): the reported improvements in activation detection and response estimation are described as 'comparable or superior' to ISC, yet the section does not appear to include formal statistical comparisons (e.g., paired tests or confidence intervals on AUC/MSE differences across Monte Carlo replicates); without these, the strength of the superiority claim remains difficult to assess.
minor comments (3)
- Abstract and §2: the abbreviation SNR is introduced for 'shared neural response' but is already standard for signal-to-noise ratio; a distinct acronym (e.g., SRF) would avoid confusion.
- §3.1 (Model): the exact parameterization of the horseshoe-inspired prior (e.g., local/global shrinkage scales and their hyperpriors) should be stated explicitly with equations to ensure reproducibility.
- Figure captions and §5 (Real data): axis labels, color scales, and voxel-count thresholds are not uniformly described; adding these details would improve interpretability of the activation maps.
Simulated Author's Rebuttal
We thank the referee for the positive assessment of our work and the recommendation for minor revision. We address the single major comment below.
read point-by-point responses
-
Referee: [—] §4 (Simulations): the reported improvements in activation detection and response estimation are described as 'comparable or superior' to ISC, yet the section does not appear to include formal statistical comparisons (e.g., paired tests or confidence intervals on AUC/MSE differences across Monte Carlo replicates); without these, the strength of the superiority claim remains difficult to assess.
Authors: We agree that formal statistical comparisons would strengthen the claims in the simulation section. In the revised manuscript we will add paired t-tests (or Wilcoxon signed-rank tests if normality is violated) on the per-replicate differences in AUC and MSE, together with 95% confidence intervals for the mean performance metrics across the Monte Carlo runs. These additions will allow readers to assess the statistical significance and precision of the reported improvements. revision: yes
Circularity Check
No significant circularity: model combines established components with independent empirical validation
full rationale
The paper constructs a Bayesian model for shared neural response in fMRI by combining sparse Gaussian process estimation with a horseshoe-inspired sparsity prior, plus a spatial extension. These are standard tools (GP regression, horseshoe prior from external literature) applied to the problem of detecting shared activation and estimating the SNR function. The derivation chain consists of specifying the hierarchical model, deriving posterior inference (likely via MCMC or variational methods), and then validating through simulation studies with known ground truth plus application to task-based and naturalistic datasets. No equation reduces a prediction to a fitted parameter by construction, no uniqueness theorem is imported from self-citation, and no ansatz is smuggled via prior work by the same authors. Performance claims are benchmarked against ISC on explicit metrics, providing falsifiable external checks. This is the normal case of a self-contained modeling paper whose central results do not collapse to tautology.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption fMRI signals contain a shared neural response function across subjects that can be represented by a sparse Gaussian process
- domain assumption A horseshoe-inspired Bayesian sparsity prior can reliably identify voxels with shared activation while controlling for individual variability
Reference graph
Works this paper leans on
-
[1]
URLhttps://doi.org/10.5281/zenodo.3937849
doi: 10.5281/zenodo.3937849. URLhttps://doi.org/10.5281/zenodo.3937849. Gang Chen, Paul-Christian Bürkner, Paul A Taylor, Zhihao Li, Lijun Yin, Daniel R Glen, Joshua Kinnison, Robert W Cox, and Luiz Pessoa. An integrative Bayesian approach to matrix-based analysis in neuroimaging.Human Brain Mapping, 40(14):4072–4090,
-
[2]
Prasenjit Ghosh and Arijit Chakrabarti
R package version 0.7.1, https://discourse.mc-stan.org. Prasenjit Ghosh and Arijit Chakrabarti. Posterior concentration properties of a general class of shrinkage estimators around nearly black vectors.arXiv preprint arXiv:1412.8161,
-
[3]
Accessed: 2026-02-26. 15 Bayesian Naturalistic Shared Neural Response Model Qawi K Telesford, Eduardo Gonzalez-Moreira, Ting Xu, Yiwen Tian, Stanley J Colcombe, Jessica Cloud, Brian E Russ, Arnaud Falchier, Maximilian Nentwich, Jens Madsen, et al. An open-access dataset of naturalistic viewing using simultaneous eeg-fmri.Scientific Data, 10(1):554,
2026
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.