Recognition: unknown
Training cell stress patterns in 3D cellular packings
Pith reviewed 2026-05-07 13:52 UTC · model grok-4.3
The pith
A 3D vertex model shows cellular packings can learn prescribed stress patterns by updating hidden-cell shape indices with contrastive learning.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Using a three-dimensional vertex model, cellular packings can be trained to realize prescribed cell stress patterns through a contrastive learning algorithm to update hidden-cell shape indices. Learning is intrinsically collective, requiring coordinated system-wide parameter adjustments, with learnability governed by an interplay between mechanical state, capacity, and training protocol. The rigidity of the tissue controls an effective exploration-exploitation tradeoff: fluid-like regimes enhance exploration through cellular rearrangements, while rigid regimes constrain dynamics and favor exploitation of existing configurations. These rearrangements introduce discontinuous learning dynamics,
What carries the argument
The contrastive learning algorithm that iteratively updates the shape indices of hidden cells in the 3D vertex model to reduce the mismatch between realized and target stress patterns.
If this is right
- Learning slows and becomes more heterogeneous with increasing ratio of target cells or constraint load, relying on rare rearrangements to escape constrained states.
- Sequential training of cells yields more robust outcomes than parallel protocols but requires more time for the loads examined.
- Tissue rigidity sets an exploration-exploitation tradeoff that produces a phase diagram of learnability controlled by rigidity, load, and protocol.
- Rearrangements create discontinuous jumps that let the system move between distinct local minima in the cost landscape.
Where Pith is reading between the lines
- Engineered cell-based materials could be designed to self-organize toward desired internal mechanical states by applying similar update rules.
- Biological tissues may already exploit rigidity-dependent rearrangements to adapt stress patterns during growth or response to injury.
- Varying the order of cell updates could be tested as a practical way to improve learning speed in physical analogs of the model.
- The approach opens the possibility of using cell packings as physical reservoirs that store and process mechanical information.
Load-bearing premise
The 3D vertex model together with contrastive updates on cell shape indices accurately captures how real cellular tissues would adjust to achieve prescribed stress patterns.
What would settle it
An experiment on a real three-dimensional cell aggregate in which cells fail to produce the target stress pattern after shape-index adjustments predicted by the trained model, or succeed under conditions the model says should be impossible.
Figures







read the original abstract
The task of learning patterns is typically associated with systems that update parameters on fixed architectures, such as neural networks, where learning proceeds through continuous optimization. Here, we demonstrate that pattern learning can also emerge in reconfigurable cellular tissue, where both mechanical parameters and network topology evolve. Using a three-dimensional vertex model, we show that cellular packings can be trained to realize prescribed cell stress patterns through a contrastive learning algorithm to update hidden-cell shape indices. We find that learning is intrinsically collective, requiring coordinated, system-wide parameter adjustments, with learnability governed by an interplay between mechanical state, capacity, and training protocol. In particular, the rigidity of the tissue controls an effective exploration-exploitation tradeoff: fluid-like regimes enhance exploration through cellular rearrangements, while rigid regimes constrain dynamics and favor exploitation of existing configurations. These rearrangements introduce discontinuous learning dynamics, enabling the system to transition between distinct local minima in the cost function landscape. As the ratio of target cells to the total number cells in the packing or constraint load increases, learning becomes slower, more heterogeneous, and increasingly dependent on rare rearrangements that allow escape from geometrically constrained states. Finally, training cells in sequence, in contrast to parallel protocols, provides an alternative route that can be more robust but generally takes longer to train for the constraint loads studied. These results suggest a learning phase diagram governed by constraint load, cell packing rigidity, and training protocol. By enabling the training of localized internal states, this work positions tissues not only as adaptive materials, but as nonconventional AI platforms.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript uses a 3D vertex model of cellular packings to demonstrate that prescribed cell stress patterns can be realized by applying a contrastive learning algorithm that updates the shape indices of hidden cells. Learning is shown to be collective and governed by an interplay of mechanical rigidity (controlling exploration via rearrangements versus exploitation), constraint load (ratio of target to total cells), and training protocol (sequential versus parallel), with rearrangements producing discontinuous jumps between cost minima; the authors propose a learning phase diagram in the space of these parameters.
Significance. If the simulation results are quantitatively validated, the work establishes that reconfigurable mechanical networks with evolving topology can perform pattern-learning tasks, extending machine-learning concepts to active matter and suggesting tissues as adaptive or computational materials. The explicit mapping of rigidity to an exploration-exploitation tradeoff and the role of topological rearrangements in escaping local minima are potentially novel contributions.
major comments (2)
- [Results (rigidity and rearrangements subsection)] The central claim that rigidity gates an exploration-exploitation tradeoff and produces discontinuous learning dynamics rests on the observed correlation between rearrangement events and cost-function jumps. The manuscript must supply quantitative support (e.g., histograms of rearrangement times versus learning epochs, or the fraction of successful escapes from local minima) in the results section that compares fluid and rigid regimes at fixed constraint load; without these metrics the tradeoff remains qualitative.
- [Training protocols comparison] The statement that sequential training is 'more robust but generally takes longer' for the studied constraint loads requires explicit comparison data (learning curves, success rates, and variance across multiple initial conditions) between sequential and parallel protocols at the same target-cell fractions; the current description leaves open whether the robustness advantage is statistically significant or merely anecdotal.
minor comments (2)
- [Abstract and Introduction] The term 'hidden-cell shape indices' is introduced without an immediate definition or reference to the vertex-model energy functional; a brief equation or parenthetical clarification on first use would improve readability.
- [Figures] Figure captions should explicitly state the number of independent realizations, the precise definition of 'constraint load,' and the color scale or normalization used for stress-pattern visualizations.
Simulated Author's Rebuttal
We thank the referee for their detailed and constructive report. We have addressed the major comments by planning to include additional quantitative data and analyses in the revised manuscript, as outlined in the point-by-point responses below.
read point-by-point responses
-
Referee: [Results (rigidity and rearrangements subsection)] The central claim that rigidity gates an exploration-exploitation tradeoff and produces discontinuous learning dynamics rests on the observed correlation between rearrangement events and cost-function jumps. The manuscript must supply quantitative support (e.g., histograms of rearrangement times versus learning epochs, or the fraction of successful escapes from local minima) in the results section that compares fluid and rigid regimes at fixed constraint load; without these metrics the tradeoff remains qualitative.
Authors: We agree with the referee that providing quantitative metrics will make the evidence for the rigidity-controlled exploration-exploitation tradeoff more rigorous. In the revised version, we will add a new figure or panel in the Results section showing histograms of rearrangement event timings relative to learning epochs for fluid and rigid regimes at the same constraint load. We will also compute and report the fraction of successful escapes from local minima facilitated by rearrangements in each regime, based on multiple simulation runs. This will quantify the discontinuous jumps and the difference in dynamics between the regimes. revision: yes
-
Referee: [Training protocols comparison] The statement that sequential training is 'more robust but generally takes longer' for the studied constraint loads requires explicit comparison data (learning curves, success rates, and variance across multiple initial conditions) between sequential and parallel protocols at the same target-cell fractions; the current description leaves open whether the robustness advantage is statistically significant or merely anecdotal.
Authors: We acknowledge that the current description of the training protocol comparison could be strengthened with statistical details. In the revision, we will include averaged learning curves with error bars representing variance across multiple independent initial conditions for both sequential and parallel protocols at identical target-cell fractions. We will also tabulate success rates (defined as achieving a cost below a certain threshold) and the average number of training steps required, allowing a direct statistical comparison of robustness and efficiency. revision: yes
Circularity Check
Simulation protocol is self-contained with no circular reductions
full rationale
The paper presents a computational demonstration in which a 3D vertex model is driven by an explicitly defined contrastive learning rule that updates hidden-cell shape indices to match target stress patterns. All reported outcomes (collective learning, rigidity-gated rearrangements, effects of sequential vs. parallel training, dependence on constraint load) follow directly from running the stated energy functional, update dynamics, and protocol on the model; none are obtained by fitting a parameter to a subset of the target data and then relabeling the fit as a prediction, nor by importing a uniqueness theorem or ansatz from self-citation that is itself unverified. The derivation chain is therefore a forward simulation whose results are independent of the target patterns once the model and algorithm are fixed.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption The 3D vertex model accurately captures cell mechanics and rearrangements in tissue packings.
- domain assumption Contrastive learning can be implemented by updating hidden-cell shape indices to minimize a cost function for target stress patterns.
Reference graph
Works this paper leans on
-
[1]
Maintenance of this assignment of tar- get shape indices shall be referred to as thefree stateof the system
Target Cells: For a training task we assignN T target cells with initial target shape index assign- ments{s T 0 }. Maintenance of this assignment of tar- get shape indices shall be referred to as thefree stateof the system. In contrast, aclamped stateis achieved by changing{s T 0 }. The training task seeks to achieve target maximum shear stresses{σ T }on ...
-
[2]
Training involves chang- ing these assignments using a learning rule
Hidden Cells: These cells have initial target shape index assignments{s H 0 }. Training involves chang- ing these assignments using a learning rule. In other words,{s H 0 }constitute the learning, or mod- ifiable, parameters of the system
-
[3]
clamp- ing
Passive Cells: These cells have initial target shape index assignments{s P 0 }, which will remain un- changed in all stages of the training algorithm. That is, while these cells appear in the total energy of the system, they are not part of the training al- gorithm. We note that the set of passive cells may be empty. As we will choose some fraction of cel...
-
[4]
Boussard, A
A. Boussard, A. Fessel, C. Oettmeier, L. Briard, H.- G. D¨ obereiner, and A. Dussutour, Adaptive behaviour and learning in slime moulds: the role of oscillations, Philosophical Transactions of the Royal Society B376, 20190757 (2021)
2021
-
[5]
Schick, E
L. Schick, E. Eichenlaub, F. Drexel, A. Mayer, S. Chen, M. Roper, and K. Alim, Decision-making in light-trapped slime molds involves active mechanical processes, PRX Life , (2026)
2026
-
[6]
Rajan, T
D. Rajan, T. Makushok, A. Kalish, L. Acuna, A. Bonville, K. C. Almanza, B. Garibay, E. Tang, M. Voss, A. Lin,et al., Single-cell analysis of habituation in stentor coeruleus, Current Biology33, 241 (2023)
2023
-
[7]
S. J. Gershman, P. E. Balbi, C. R. Gallistel, and J. Gu- nawardena, Reconsidering the evidence for learning in single cells, Elife10, e61907 (2021)
2021
-
[8]
D. H. Rajan and W. F. Marshall, A receptor-inactivation model for single-celled habituation in stentor coeruleus, Current Biology35, 3327 (2025)
2025
-
[9]
A. Zentner, E. V. Halingstad, C. Chalk, M. P. Bren- ner, A. Murugan, E. Winfree, and K. Shrinivas, Informa- tion processing driven by multicomponent surface con- densates, arXiv preprint arXiv:2509.08100 (2025)
-
[10]
C. G. Evans, J. O’Brien, E. Winfree, and A. Murugan, Pattern recognition in the nucleation kinetics of non- equilibrium self-assembly, Nature625, 500 (2024)
2024
-
[11]
Bahri, J
Y. Bahri, J. Kadmon, J. Pennington, S. S. Schoenholz, J. Sohl-Dickstein, and S. Ganguli, Statistical mechanics of deep learning, Annual Review of Condensed Matter Physics11, 501 (2020)
2020
- [12]
-
[13]
L. G. Wright, T. Onodera, M. M. Stein, T. Wang, D. T. Schachter, Z. Hu, and P. L. McMahon, Deep physical neural networks trained with backpropagation, Nature 601, 549 (2022)
2022
-
[14]
Stern and A
M. Stern and A. Murugan, Learning without neurons in physical systems, Annual Review of Condensed Matter Physics14, 417 (2023)
2023
- [15]
-
[16]
Momeni, B
A. Momeni, B. Rahmani, B. Scellier, L. G. Wright, P. L. McMahon, C. C. Wanjura, Y. Li, A. Skalli, N. G. Berloff, T. Onodera,et al., Training of physical neural networks, Nature645, 53 (2025)
2025
-
[17]
Scellier and Y
B. Scellier and Y. Bengio, Equilibrium propagation: Bridging the gap between energy-based models and back- propagation, Frontiers in computational neuroscience11, 24 (2017). 16
2017
-
[18]
V. R. Anisetti, B. Scellier, and J. M. Schwarz, Learning by non-interfering feedback chemical signaling in physical networks, Physical Review Research5, 023024 (2023)
2023
-
[19]
V. R. Anisetti, A. Kandala, B. Scellier, and J. M. Schwarz, Frequency propagation: Multimechanism learn- ing in nonlinear physical networks, Neural Computation 36, 596 (2024)
2024
-
[20]
Li and X
S. Li and X. Mao, Training all-mechanical neural net- works for task learning through in situ backpropagation, Nature Communications15, 10528 (2024)
2024
-
[21]
M. J. Falk, A. T. Strupp, B. Scellier, and A. Muru- gan, Temporal contrastive learning through implicit non- equilibrium memory, Nature Communications16, 2163 (2025)
2025
-
[22]
Hexner, Training precise stress patterns, Soft Matter 19, 2120 (2023)
D. Hexner, Training precise stress patterns, Soft Matter 19, 2120 (2023)
2023
-
[23]
D. O. Hebb, The organization of behavior: A neuropsy- chological theory, Wiley (1949)
1949
-
[24]
Gopinathan, K.-C
A. Gopinathan, K.-C. Lee, J. M. Schwarz, and A. J. Liu, Branching, capping, and severing in dynamic actin struc- tures, Physical review letters99, 058103 (2007)
2007
-
[25]
D. A. Fletcher and R. D. Mullins, Cell mechanics and the cytoskeleton, Nature463, 485 (2010)
2010
- [26]
-
[27]
Berger-Tal, R
O. Berger-Tal, R. Nathan, E. Meron, and D. Saltz, The exploration–exploitation dilemma: A multidisciplinary framework, PLoS ONE9, e95693 (2014)
2014
-
[28]
Zhang, S
T. Zhang, S. Gupta, M. Lancaster, and J. M. Schwarz, How human-derived brain organoids are built differently from brain organoids derived from genetically-close rela- tives: A multi-scale hypothesis, Soft Matter (2026)
2026
-
[29]
Nasrollahi, C
S. Nasrollahi, C. Walter, A. J. Loza, G. V. Schimizzi, G. D. Longmore, and A. Pathak, Past matrix stiffness primes epithelial cells and regulates their future collective migration through a mechanical memory, Biomaterials 146, 146 (2017)
2017
-
[30]
Honda, M
H. Honda, M. Tanemura, and T. Nagai, A three- dimensional vertex dynamics cell model of space-filling polyhedra simulating cell behavior in a cell aggregate, Journal of Theoretical Biology226, 439 (2004)
2004
-
[31]
Okuda, Y
S. Okuda, Y. Inoue, M. Eiraku, Y. Sasai, and T. Adachi, Reversible network reconnection model for simulating large deformation in dynamic tissue morphogenesis, Biomechanics and Modeling in Mechanobiology12, 627 (2013)
2013
-
[32]
Okuda, Y
S. Okuda, Y. Inoue, and T. Adachi, Three-dimensional vertex model for simulating multicellular morphogenesis, Biophysics and Physicobiology12, 13 (2015)
2015
-
[33]
Zhang and J
T. Zhang and J. M. Schwarz, Topologically-protected in- terior for three-dimensional confluent cellular collectives, Phys. Rev. Res.4, 043148 (2022)
2022
- [34]
-
[35]
Bitzek, P
E. Bitzek, P. Koskinen, F. G¨ ahler, M. Moseler, and P. Gumbsch, Structural relaxation made simple, Phys. Rev. Lett.97, 170201 (2006)
2006
-
[36]
Bitzek and P
E. Bitzek and P. Gumbsch, An efficient algorithm for structural relaxation of atomic systems, Modelling and Simulation in Materials Science and Engineering15, S159 (2007)
2007
-
[37]
Geman, E
S. Geman, E. Bienenstock, and R. Doursat, Neural net- works and the bias/variance dilemma, Neural Computa- tion4, 1 (1992)
1992
-
[38]
Hastie, R
T. Hastie, R. Tibshirani, and J. Friedman,The Elements of Statistical Learning, 2nd ed. (Springer, 2009)
2009
-
[39]
M. S. Advani, A. M. Saxe, and H. Sompolinsky, High- dimensional dynamics of generalization error in neural networks, Neural Networks132, 428 (2020)
2020
-
[40]
Parker, M
A. Parker, M. C. Marchetti, M. L. Manning, and J. Schwarz, How does the extracellular matrix affect the rigidity of an embedded spheroid?, Physical Review E 111, 044410 (2025)
2025
- [41]
-
[42]
Arzash, I
S. Arzash, I. Tah, A. J. Liu, and M. L. Manning, Rigidity of epithelial tissues as a double optimization problem, Physical Review Research7, 013157 (2025)
2025
-
[43]
Arzash and S
S. Arzash and S. Banerjee, Learning epithelial elasticity via local tension remodeling, bioRxiv , 2025 (2025)
2025
-
[44]
Arzash, A
S. Arzash, A. J. Liu, and M. L. Manning, Epithelial con- vergent extension as a tuning process, bioRxiv , 2025 (2025)
2025
-
[45]
VertAX: a differentiable vertex model for learning epithelial tissue mechanics
A. Pasqui, J. M. C. Ocana, A. Sinha, M. Perez, F. Del- bary, G. Gosti, M. Miotto, D. Caudo, M. Ernoult, and H. Turlier, Vertax: a differentiable vertex model for learning epithelial tissue mechanics, arXiv preprint arXiv:2604.06896 (2026)
work page internal anchor Pith review Pith/arXiv arXiv 2026
-
[46]
D. S. Banerjee, M. J. Falk, M. L. Gardel, A. M. Wal- czak, T. Mora, and S. Vaikuntanathan, Learning via mechanosensitivity and activity in cytoskeletal networks, PRX Life4, 013001 (2026)
2026
-
[47]
Shomar, O
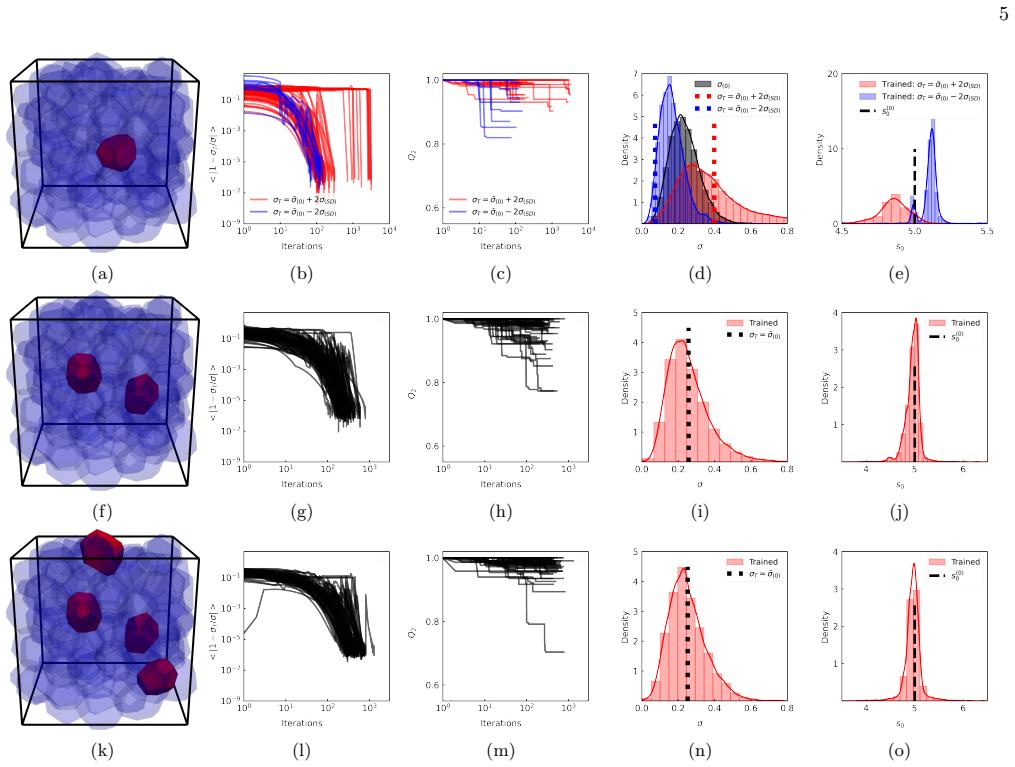
A. Shomar, O. Barak, and N. Brenner, Cancer progres- sion as a learning process, Iscience25(2022). 17 SUPPLEMENT AR Y FIGURES (a) (b) (c) (d) (e) (f) (g) (h) (i) (j) (k) (l) FIG. S1:Reducing geometric frustration enhances learning.The same parameters are used as in Figure 1, with the exception ofk V = 1. The row and column layout logic, as well as color s...
2022
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.