Recognition: unknown
Mining Negative Sequential Patterns to Improve Viral Genomic Feature Representation and Classification
Pith reviewed 2026-05-07 14:04 UTC · model grok-4.3
The pith
Negative sequential patterns from viral genomes raise classification accuracy across classifiers.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
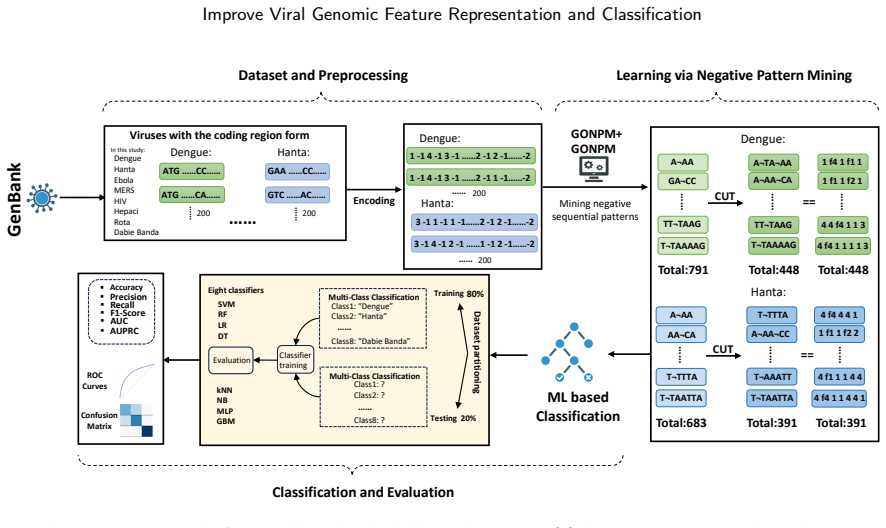
GeneNSPCla extracts negative sequential patterns as absence-based features from RNA viral genomes, converts them to numerical vectors, and feeds them into multiple classifiers; the adapted GONPM+ algorithm yields patterns that produce higher accuracy than standard negative mining or positive pattern methods alone.
What carries the argument
Negative sequential patterns (NSPs) as absence-based features from nucleotide sequences, discovered via the GONPM+ mining algorithm tailored for genomic data.
If this is right
- Absence signals from negative patterns complement presence signals and improve feature representation for viral sequences.
- GONPM+ produces longer, more biologically meaningful negative patterns than the original negative pattern mining algorithm.
- The resulting features raise average classifier accuracy by 10.03 percent over the baseline negative method and 24.75 percent over positive pattern mining.
- The framework supplies a complementary view to composition-based approaches for viral genome analysis.
Where Pith is reading between the lines
- If the gains survive controlled experiments with matched feature counts, the same absence-pattern approach could be tested on non-viral sequence classification tasks.
- Negative patterns may highlight evolutionary constraints or host-interaction motifs that positive patterns alone do not reveal.
- The method could be applied to metagenomic samples where viral sequences are mixed with host and bacterial DNA.
Load-bearing premise
The reported accuracy gains come from the negative sequential patterns themselves rather than from differences in total feature count, classifier hyperparameters, or preprocessing steps.
What would settle it
Re-running the eight classifiers on the same viral datasets while holding feature count, hyperparameters, and preprocessing fixed and using only positive patterns to check if accuracy gains disappear.
Figures






read the original abstract
Viruses represent the most abundant biological entities on Earth and play a pivotal role in microbial ecosystems, yet, as prominent human pathogens, they are closely linked to human morbidity and mortality. Accurate identification of viral sequences from viral genome sequences is therefore essential, but existing genome-based classification models that largely relying on composition- or frequency-based subsequence features often suffer from limited interpretability and reduced accuracy, particularly on complex or imbalanced datasets. To address these limitations, we propose GeneNSPCla (Genomic Negative Sequential Pattern-based Classification), a novel viral classification framework based on Negative Sequential Patterns (NSPs) that extracts discriminative absence-based features from nucleotide sequences of RNA viral genomes. By transforming these NSPs into numerical feature vectors and integrating them into multiple supervised classifiers, GeneNSPCla effectively captures both presence and absence signals in viral sequences. Furthermore, we propose a negative pattern mining algorithm adapted for processing genomic data: GONPM+, which can discover longer and more biologically meaningful negative sequential patterns. The experimental results demonstrate that the average accuracy of GONPM+ in 8 classifiers has improved by 10.03% compared to the original negative pattern mining algorithm and by 24.75% compared to the positive pattern mining algorithm. These findings highlight the effectiveness of incorporating absence-based sequential information, providing a new and complementary perspective for viral genome analysis and classification.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes GeneNSPCla, a viral genome classification framework that extracts negative sequential patterns (NSPs) from RNA viral nucleotide sequences using a new mining algorithm GONPM+ to generate absence-based discriminative features. These features are converted to numerical vectors and fed into supervised classifiers; the work claims that GONPM+ yields longer, more biologically meaningful patterns and produces average accuracy gains of 10.03% over the original negative pattern mining algorithm and 24.75% over positive pattern mining across eight classifiers.
Significance. If the reported accuracy improvements are reproducible and causally attributable to the negative patterns rather than confounding experimental factors, the approach supplies a complementary signal (absence of subsequences) to existing composition- or frequency-based genomic features. This could enhance both predictive performance and interpretability on imbalanced viral datasets and suggests a generalizable adaptation of sequential pattern mining to biological sequence data.
major comments (1)
- [Experimental Results] Experimental Results (or equivalent section containing the accuracy claims): the stated 10.03% and 24.75% average accuracy lifts are presented without any description of dataset sizes, class balance, train/test split protocol, exact number of features retained by GONPM+ versus the two baselines, hyperparameter search procedure, or statistical significance testing. Because these controls are absent, it is impossible to determine whether the observed deltas arise from the negative-pattern representation itself or from uncontrolled differences in feature cardinality or tuning.
minor comments (1)
- [Abstract] Abstract: the clause 'existing genome-based classification models that largely relying on' contains a subject-verb agreement error and should read 'that largely rely on'.
Simulated Author's Rebuttal
We thank the referee for the detailed and constructive review. The concern regarding missing experimental controls is valid and we have revised the manuscript accordingly to improve reproducibility and clarify the source of the reported accuracy gains.
read point-by-point responses
-
Referee: [Experimental Results] Experimental Results (or equivalent section containing the accuracy claims): the stated 10.03% and 24.75% average accuracy lifts are presented without any description of dataset sizes, class balance, train/test split protocol, exact number of features retained by GONPM+ versus the two baselines, hyperparameter search procedure, or statistical significance testing. Because these controls are absent, it is impossible to determine whether the observed deltas arise from the negative-pattern representation itself or from uncontrolled differences in feature cardinality or tuning.
Authors: We agree that these methodological details are essential for evaluating whether the accuracy improvements are attributable to the negative sequential patterns. In the revised manuscript we have expanded the Experimental Results section to explicitly describe: the eight RNA viral genome datasets (including total sequence counts and class imbalance ratios), the train/test split protocol (70/30 stratified split with 5-fold cross-validation), the exact number of retained features for GONPM+ versus the original negative-pattern miner and the positive-pattern baseline, the grid-search hyperparameter procedure applied uniformly to all eight classifiers, and the statistical significance tests (paired t-tests) performed on the accuracy differences. These additions demonstrate that the gains arise from the NSP representation rather than differences in feature cardinality or tuning. revision: yes
Circularity Check
No circularity: empirical accuracy gains are measured outcomes, not tautological predictions
full rationale
The paper proposes the GONPM+ algorithm for mining negative sequential patterns from viral genomes and integrates the resulting absence-based features into standard classifiers. All reported improvements (10.03% and 24.75% average accuracy) are presented as direct experimental measurements on held-out data rather than as first-principles derivations or predictions. No equations, uniqueness theorems, or ansatzes are introduced that reduce to the input data or to prior self-citations by construction. The central claim therefore remains an empirical observation whose validity depends on experimental controls, not on definitional equivalence.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Negative sequential patterns extracted from RNA viral genomes provide complementary discriminative signals to positive frequency-based features.
Reference graph
Works this paper leans on
-
[1]
Miningassociation rules between sets of items in large databases
Agrawal,R.,Imieliński,T.,andSwami,A.(1993). Miningassociation rules between sets of items in large databases. InThe ACM SIGMOD International Conference on Management of Data, pages 207–216. Wenxi Zhu et al.:Preprint submitted to Elsevier Page 16 of 18 Improve Viral Genomic Feature Representation and Classification
1993
-
[2]
Miningsequentialpatterns
Agrawal,R.andSrikant,R.(1995). Miningsequentialpatterns. In The Eleventh International Conference on Data Engineering, pages 3–14. IEEE
1995
-
[3]
and Jeon, G
Ahmed, I. and Jeon, G. (2022). Enabling artificial intelligence for genome sequence analysis of COVID-19 and alike viruses.Interdis- ciplinary Sciences: Computational Life Sciences, 14(2):504–519
2022
-
[4]
Anew profilingapproachforDNAsequencesbasedonthenucleotides’physic- ochemicalfeaturesforaccurateanalysisofSARS-CoV-2genomes
AkbariRoknAbadi,S.,Mohammadi,A.,andKoohi,S.(2023). Anew profilingapproachforDNAsequencesbasedonthenucleotides’physic- ochemicalfeaturesforaccurateanalysisofSARS-CoV-2genomes. BMC Genomics, 24(1):266
2023
-
[5]
Almeida,J.S.,Carrico,J.A.,Maretzek,A.,Noble,P.A.,andFletcher, M. (2001). Analysis of genomic sequences by chaos game representa- tion. Bioinformatics, 17(5):429–437
2001
-
[6]
H., Sindhu, S
Alshayeji, M. H., Sindhu, S. C., and Abed, S. (2023). Viral genome prediction from raw human DNA sequence samples by combining natural language processing and machine learning techniques.Expert Systems with Applications, 218:119641
2023
-
[7]
F., Gish, W., Miller, W., Myers, E
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman, D. J. (1990).Basiclocalalignmentsearchtool. JournalofMolecularBiology , 215(3):403–410
1990
-
[8]
S., de Souza, L
Azevedo, K. S., de Souza, L. C., Coutinho, M. G., de M. Barbosa, R., and Fernandes, M. A. (2024). DeepVirusClassifier: a deep learning tool for classifying SARS-CoV-2 based on viral subtypes within the coronaviridae family.BMC Bioinformatics, 25(1):231
2024
-
[9]
L., Pimentel, H., Melsted, P., and Pachter, L
Bray, N. L., Pimentel, H., Melsted, P., and Pachter, L. (2016). Near- optimal probabilistic RNA-seq quantification. Nature Biotechnology, 34(5):525–527
2016
-
[10]
Burki, T. (2023). First shared SARS-CoV-2 genome: GISAID vs virological. org.The Lancet Microbe, 4(6):e395
2023
-
[11]
XGBoost:Ascalabletreeboosting system
Chen,T.andGuestrin,C.(2016). XGBoost:Ascalabletreeboosting system. In The 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, pages 785–794
2016
-
[12]
e-RNSP:Anefficientmethod forminingrepetitionnegativesequentialpatterns
Dong,X.,Gong,Y.,andCao,L.(2018). e-RNSP:Anefficientmethod forminingrepetitionnegativesequentialpatterns. IEEETransactionson Cybernetics, 50(5):2084–2096
2018
-
[13]
Fournier-Viger, P., Gan, W., Wu, Y., Nouioua, M., Song, W., Truong, T., and Duong, H. (2022). Pattern mining: Current challenges and opportunities. In International Conference on Database Systems for Advanced Applications, pages 34–49. Springer
2022
-
[14]
Fast vertical mining of sequential patterns using co-occurrence infor- mation
Fournier-Viger,P.,Gomariz,A.,Campos,M.,andThomas,R.(2014). Fast vertical mining of sequential patterns using co-occurrence infor- mation. InPacific-Asia Conference on Knowledge Discovery and Data Mining, pages 40–52. Springer
2014
-
[15]
Fournier-Viger,P.,Lin,J.C.-W.,Kiran,R.U.,Koh,Y.S.,andThomas, R. (2017). A survey of sequential pattern mining.Data Science and Pattern Recognition, 1(1):54–77
2017
-
[16]
C.-W., Fournier-Viger, P., Chao, H.-C., Tseng, V
Gan, W., Lin, J. C.-W., Fournier-Viger, P., Chao, H.-C., Tseng, V. S., and Yu, P. S. (2021). A survey of utility-oriented pattern mining.IEEE Transactions on Knowledge and Data Engineering, 33(4):1306–1327
2021
-
[17]
Georgakopoulos-Soares, I., Yizhar-Barnea, O., Mouratidis, I., Hem- berg, M., and Ahituv, N. (2021). Absent from DNA and protein: ge- nomic characterization of nullomers and nullpeptides across functional categories and evolution.Genome Biology, 22(1):245
2021
-
[18]
R., Clooney, A
Guerin, E., Shkoporov, A., Stockdale, S. R., Clooney, A. G., Ryan, F.J.,Sutton,T.D.,Draper,L.A.,Gonzalez-Tortuero,E.,Ross,R.P.,and Hill,C.(2018). Biologyandtaxonomyofcrass-likebacteriophages,the mostabundantvirusinthehumangut. CellHost&Microbe ,24(5):653– 664
2018
-
[19]
Gunasekaran, H., Ramalakshmi, K., Rex Macedo Arokiaraj, A., Deepa Kanmani, S., Venkatesan, C., and Suresh Gnana Dhas, C. (2021). Analysis of DNA sequence classification using CNN and hybridmodels. ComputationalandMathematicalMethodsinMedicine , 2021(1):1835056
2021
-
[20]
Z., Verma, H., Kumar, R., Sood, U., Hira, P., et al
Gupta,V.,Haider,S.,Verma,M.,Singhvi,N.,Ponnusamy,K.,Malik, M. Z., Verma, H., Kumar, R., Sood, U., Hira, P., et al. (2021). Com- parativegenomicsandintegratednetworkapproachunveiledundirected phylogeny patterns, co-mutational hot spots, functional cross talk, and regulatory interactions in SARS-CoV-2.MSystems, 6(1):10–1128
2021
-
[21]
and Quiniou, R
Guyet, T. and Quiniou, R. (2020). NegPSpan: efficient extraction of negative sequential patterns with embedding constraints.Data Mining and Knowledge Discovery, 34:563–609
2020
-
[22]
Hossain,M.M.,Wu,Y.,Fournier-Viger,P.,Li,Z.,Guo,L.,andLi,Y. (2021). HSNP-Miner:Highutilityself-adaptivenonoverlappingpattern mining. In IEEE International Conference on Big Knowledge, pages 70–77. IEEE
2021
-
[23]
A., and Sun, F
Hou, S., Tang, T., Cheng, S., Liu, Y., Xia, T., Chen, T., Fuhrman, J. A., and Sun, F. (2024). DeepMicroClass sorts metagenomic contigs into prokaryotes, eukaryotes and viruses.NAR Genomics and Bioinfor- matics, 6(2):lqae044
2024
-
[24]
Huang, J.-W., Wu, Y.-B., and Jaysawal, B. P. (2020). On mining progressive positive and negative sequential patterns simultaneously. Journal of Information Science & Engineering, 36(1)
2020
-
[25]
Islam,M.S.,AlFarid,F.,Shamrat,F.J.M.,Islam,M.N.,Rashid,M., Bari,B.S.,Abdullah,J.,Islam,M.N.,Akhtaruzzaman,M.,Kabir,M.N., et al. (2024). Challenges issues and future recommendations of deep learning techniques for SARS-CoV-2 detection utilising X-ray and CT images: a comprehensive review.PeerJ Computer Science, 10:e2517
2024
-
[26]
and Burge, C
Kariin, S. and Burge, C. (1995). Dinucleotide relative abundance extremes: a genomic signature.Trends in Genetics, 11(7):283–290
1995
-
[27]
Kim,J.,Lee,K.,Rupasinghe,R.,Rezaei,S.,Martínez-López,B.,and Liu, X. (2021). Applications of machine learning for the classification of porcine reproductive and respiratory syndrome virus sublineages using amino acid scores of orf5 gene.Frontiers in Veterinary Science, 8:683134
2021
-
[28]
and Frith, M
Koulouras, G. and Frith, M. C. (2021). Significant non-existence of sequences in genomes and proteomes. Nucleic Acids Research, 49(6):3139–3155
2021
-
[29]
J., Dempsey, D
Lefkowitz, E. J., Dempsey, D. M., Hendrickson, R. C., Orton, R. J., Siddell, S. G., and Smith, D. B. (2018). Virus taxonomy: the database oftheinternationalcommitteeontaxonomyofviruses(ICTV). Nucleic Acids Research, 46(D1):D708–D717
2018
-
[30]
and Sun, F
Li, H. and Sun, F. (2018). Comparative studies of alignment, alignment-free and SVM based approaches for predicting the hosts of viruses based on viral sequences.Scientific Reports, 8(1):10032
2018
-
[31]
Li, Y., Wang, Z., Liu, J., Guo, L., Fournier-Viger, P., Wu, Y., and Wu, X. (2025). Mining repetitive negative sequential patterns with gap constraints. ACM Transactions on Knowledge Discovery from Data, 19(4):1–29
2025
-
[32]
Liao, V. C.-C. and Chen, M.-S. (2013). Efficient mining gapped sequential patterns for motifs in biological sequences.BMC Systems Biology, 7(Suppl 4):S7
2013
-
[33]
Liu, G., Chen, X., Luan, Y., and Li, D. (2024). VirusPredictor: Xgboost-based software to predict virus-related sequences in human data. Bioinformatics, 40(4):btae192
2024
-
[34]
A., Chan, C
Moeckel, C., Mareboina, M., Konnaris, M. A., Chan, C. S., Moura- tidis, I., Montgomery, A., Chantzi, N., Pavlopoulos, G. A., and Georgakopoulos-Soares, I. (2024). A survey of k-mer methods and applications in bioinformatics.Computational and Structural Biotech- nology Journal, 23:2289–2303
2024
-
[35]
Sequentialpatternmining– approaches and algorithms.ACM Computing Surveys, 45(2):1–39
Mooney,C.H.andRoddick,J.F.(2013). Sequentialpatternmining– approaches and algorithms.ACM Computing Surveys, 45(2):1–39
2013
-
[36]
Mordvanyuk, N., Bifet, A., and López, B. (2022). VEPRECO: Vertical databases with pre-pruning strategies and common candidate selection policies to fasten sequential pattern mining.Expert Systems with Applications, 204:117517
2022
-
[37]
S-PDB:Analysis and classification of SARS-CoV-2 spike protein structures
Nawaz,M.S.,Fournier-Viger,P.,andHe,Y.(2022). S-PDB:Analysis and classification of SARS-CoV-2 spike protein structures. InIEEE International Conference on Bioinformatics and Biomedicine, pages 2259–2265. IEEE
2022
-
[38]
S., Fournier-Viger, P., He, Y., and Zhang, Q
Nawaz, M. S., Fournier-Viger, P., He, Y., and Zhang, Q. (2023). PSAC-PDB: Analysis and classification of protein structures.Comput- ers in Biology and Medicine, 158:106814
2023
-
[39]
S., Fournier-Viger, P., Nawaz, S., Gan, W., and He, Y
Nawaz, M. S., Fournier-Viger, P., Nawaz, S., Gan, W., and He, Y. (2024a). FSP4HSP: Frequent sequential patterns for the improved classification of heat shock proteins, their families, and sub-types.In- ternational Journal of Biological Macromolecules, 277:134147. Wenxi Zhu et al.:Preprint submitted to Elsevier Page 17 of 18 Improve Viral Genomic Feature R...
-
[40]
S., Fournier-Viger, P., Nawaz, S., Zhu, H., and Yun, U
Nawaz, M. S., Fournier-Viger, P., Nawaz, S., Zhu, H., and Yun, U. (2024b). SPM4GAC: SPM-based approach for genome analysis and classification of macromolecules.International Journal of Biological Macromolecules, 266:130984
-
[41]
S., Nawaz, M
Nawaz, M. S., Nawaz, M. Z., Junyi, Z., Fournier-Viger, P., and Qu, J.-F. (2024c). Exploiting the sequential nature of genomic data for improved analysis and identification. Computers in Biology and Medicine, 183:109307
-
[42]
Novakovsky,G.,Fornes,O.,Saraswat,M.,Mostafavi,S.,andWasser- man, W. W. (2023). ExplaiNN: interpretable and transparent neural networks for genomics.Genome Biology, 24(1):154
2023
-
[43]
RCOVID19:Recurrence-basedSARS-CoV-2featuresusingchaosgame representation
Olyaee,M.H.,Pirgazi,J.,Khalifeh,K.,andKhanteymoori,A.(2020). RCOVID19:Recurrence-basedSARS-CoV-2featuresusingchaosgame representation. Data in Brief, 32:106144
2020
-
[44]
Pearson, W. R. (1994). Using the FASTA program to search protein andDNAsequencedatabases. In ComputerAnalysisofSequenceData: Part I, pages 307–331. Springer
1994
-
[45]
Pearson, W. R. (2013). BLAST and FASTA similarity searching for multiplesequencealignment. In MultipleSequenceAlignmentMethods , pages 75–101. Springer
2013
-
[46]
Pei, J., Han, J., and Wang, W. (2007). Constraint-based sequential pattern mining: the pattern-growth methods. Journal of Intelligent Information Systems, 28(2):133–160
2007
-
[47]
ViraLM: empowering virus discovery through the genome foundation model
Peng,C.,Shang,J.,Guan,J.,Wang,D.,andSun,Y.(2024). ViraLM: empowering virus discovery through the genome foundation model. Bioinformatics, 40(12):btae704
2024
-
[48]
Deeplearninginmicrobiomeanalysis:acom- prehensivereviewofneuralnetworkmodels
Przymus, P., Rykaczewski, K., Martín-Segura, A., Truu, J., Carrillo De Santa Pau, E., Kolev, M., Naskinova, I., Gruca, A., Sampri, A., Frohme,M.,etal.(2025). Deeplearninginmicrobiomeanalysis:acom- prehensivereviewofneuralnetworkmodels. FrontiersinMicrobiology , 15:1516667
2025
-
[49]
Pu,L.andShamir,R.(2024).4CAC:4-classclassifierofmetagenome contigs using machine learning and assembly graphs.Nucleic Acids Research, 52(19):e94–e94
2024
-
[50]
Qiang, X.-L., Xu, P., Fang, G., Liu, W.-B., and Kou, Z. (2020). Using the spike protein feature to predict infection risk and monitor the evolutionary dynamic of coronavirus.Infectious Diseases of Poverty, 9(1):33
2020
-
[51]
S., Soltysiak, M
Randhawa, G. S., Soltysiak, M. P., El Roz, H., de Souza, C. P., Hill, K. A., and Kari, L. (2020). Machine learning using intrinsic genomic signatures for rapid classification of novel pathogens: COVID-19 case study. Plos One, 15(4):e0232391
2020
-
[52]
Remita,M.A.,Halioui,A.,MalickDiouara,A.A.,Daigle,B.,Kiani, G., and Diallo, A. B. (2017). A machine learning approach for viral genome classification.BMC Bioinformatics, 18(1):208
2017
-
[53]
VirFinder:anovelk-merbasedtoolforidentifyingviralsequencesfrom assembled metagenomic data.Microbiome, 5(1):69
Ren,J.,Ahlgren,N.A.,Lu,Y.Y.,Fuhrman,J.A.,andSun,F.(2017). VirFinder:anovelk-merbasedtoolforidentifyingviralsequencesfrom assembled metagenomic data.Microbiome, 5(1):69
2017
-
[54]
Salmi, M., Atif, D., Oliva, D., Abraham, A., and Ventura, S. (2024). Handling imbalanced medical datasets: review of a decade of research. Artificial Intelligence Review, 57(10):273
2024
-
[55]
W., Cavanaugh, M., Clark, K., Ostell, J., Pruitt, K
Sayers, E. W., Cavanaugh, M., Clark, K., Ostell, J., Pruitt, K. D., and Karsch-Mizrachi, I. (2019). GenBank. Nucleic Acids Research, 47(D1):D94–D99
2019
-
[56]
Shi, C. H. and Yip, K. Y. (2020). A general near-exact k-mer counting method with low memory consumption enables de novo as- sembly of 106× human sequence data in 2.7 hours. Bioinformatics, 36(Supplement_2):i625–i633
2020
-
[57]
ClassificationofSARS-CoV-2andnon- SARS-CoV-2usingmachinelearningalgorithms
Singh,O.P.,Vallejo,M.,El-Badawy,I.M.,Aysha,A.,Madhanagopal, J.,andFaudzi,A.A.M.(2021). ClassificationofSARS-CoV-2andnon- SARS-CoV-2usingmachinelearningalgorithms. ComputersinBiology and Medicine, 136:104650
2021
-
[58]
and Lapalme, G
Sokolova, M. and Lapalme, G. (2009). A systematic analysis of performance measures for classification tasks.Information Processing & Management, 45(4):427–437
2009
-
[59]
Sun, C., Gong, Y., Guo, Y., Zhao, L., Guan, H., Liu, X., and Dong, X. (2024). SN-RNSP: Mining self-adaptive nonoverlapping repetitive negativesequentialpatternsintransactionsequences. Knowledge-Based Systems, 287:111449
2024
-
[60]
Suttle, C. A. (2007). Marine viruses—major players in the global ecosystem. Nature Reviews Microbiology, 5(10):801–812
2007
-
[61]
Tandan, M., Acharya, Y., Pokharel, S., and Timilsina, M. (2021). Discovering symptom patterns of COVID-19 patients using association rule mining.Computers in Biology and Medicine, 131:104249
2021
-
[62]
Detection of a SARS-CoV-2 variant of concern in south africa.Nature, 592(7854):438–443
Tegally,H.,Wilkinson,E.,Giovanetti,M.,Iranzadeh,A.,Fonseca,V., Giandhari,J.,Doolabh,D.,Pillay,S.,San,E.J.,Msomi,N.,etal.(2021). Detection of a SARS-CoV-2 variant of concern in south africa.Nature, 592(7854):438–443
2021
-
[63]
E., Chen, L., Deng, C., Zhou, G., and Hu, P
Wade, K. E., Chen, L., Deng, C., Zhou, G., and Hu, P. (2024). In- vestigatingalignment-freemachinelearningmethodsforHIV-1subtype classification. Bioinformatics Advances, 4(1):vbae108
2024
-
[64]
E., Lu, J., and Langmead, B
Wood, D. E., Lu, J., and Langmead, B. (2019). Improved metage- nomic analysis with kraken 2.Genome Biology, 20(1):257
2019
-
[65]
Multimodal large language models: A survey
Wu,J.,Gan,W.,Chen,Z.,Wan,S.,andYu,P.S.(2023a). Multimodal large language models: A survey. InIEEE International Conference on Big Data, pages 2247–2256. IEEE
-
[66]
Wu, X., Zhang, C., and Zhang, S. (2004). Efficient mining of both positive and negative association rules. ACM Transactions on Information Systems, 22(3):381–405
2004
-
[67]
Wu, Y., Chen, M., Li, Y., Liu, J., Li, Z., Li, J., and Wu, X. (2023b). ONP-Miner: One-off negative sequential pattern mining.ACM Trans- actions on Knowledge Discovery from Data, 17(3):1–24
-
[68]
HANP-Miner:Highaverageutilitynonoverlapping sequential pattern mining.Knowledge-Based Systems, 229:107361
Wu, Y.,Geng, M.,Li, Y.,Guo, L.,Li, Z.,Fournier-Viger, P.,Zhu, X., andWu,X.(2021a). HANP-Miner:Highaverageutilitynonoverlapping sequential pattern mining.Knowledge-Based Systems, 229:107361
-
[69]
Wu, Y., Wang, Y., Li, Y., Zhu, X., and Wu, X. (2021b). Top-k self-adaptive contrast sequential pattern mining.IEEE Transactions on Cybernetics, 52(11):11819–11833
-
[70]
Xu, T., Dong, X., Xu, J., and Gong, Y. (2017). E-msNSP: Effi- cient negative sequential patterns mining based on multiple minimum supports. International Journal of Pattern Recognition and Artificial Intelligence, 31(02):1750003
2017
-
[71]
Yang, M., Wang, Z., Yan, Z., Wang, W., Zhu, Q., and Jin, C. (2024). DNASimCLR: a contrastive learning-based deep learning approach for gene sequence data classification.BMC Bioinformatics, 25(1):328
2024
-
[72]
Zheng, Y., Gan, W., Chen, Z., Qi, Z., Liang, Q., and Yu, P. S. (2025). Largelanguagemodelsformedicine:asurvey. InternationalJournalof Machine Learning and Cybernetics, 16(2):1015–1040
2025
-
[73]
(2021).Machine learning
Zhou, Z.-H. (2021).Machine learning. Springer nature
2021
-
[74]
Zhu, W., Gan, W., and Qi, Z. (2025). Leveraging negative sequential patterns for advanced genomic analysis and classification. InIEEE International Conference on Bioinformatics and Biomedicine, pages 6913–6920. IEEE
2025
-
[75]
Alignment-free sequence comparison: benefits, applications, and tools
Zielezinski,A.,Vinga,S.,Almeida,J.,andKarlowski,W.M.(2017). Alignment-free sequence comparison: benefits, applications, and tools. Genome Biology, 18(1):186. Wenxi Zhu et al.:Preprint submitted to Elsevier Page 18 of 18
2017
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.