Recognition: unknown
How is gene-regulatory evolution affected by cell-to-cell variability?
Pith reviewed 2026-05-07 13:41 UTC · model grok-4.3
The pith
Increased cell-to-cell variability selects for aligned and robust gene-regulatory networks enriched in specific motifs.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
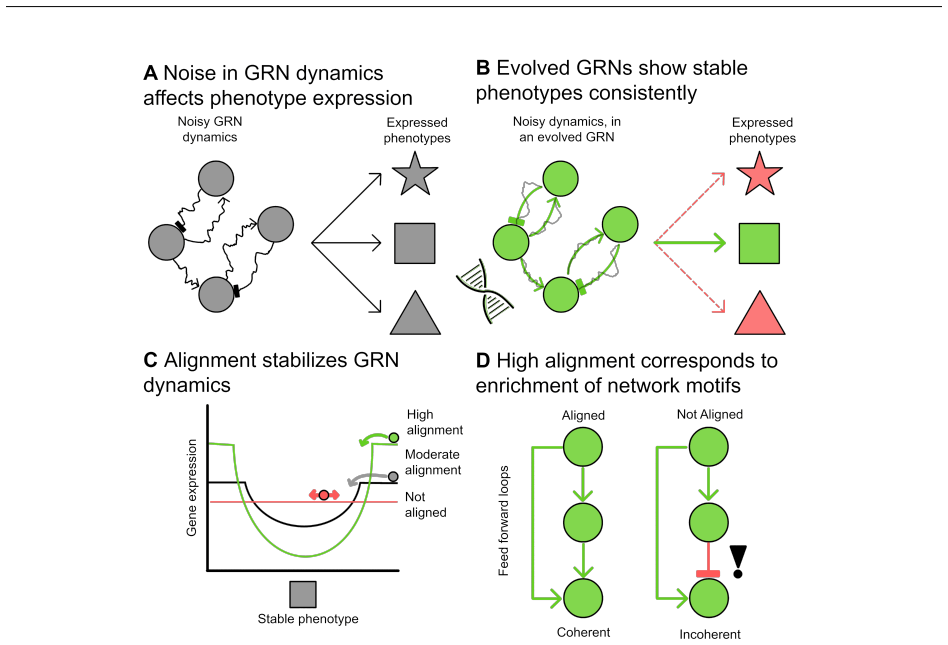
When developmental dynamics include noise, selection favors networks in which a larger fraction of gene-gene interactions carry the correct sign to support the target phenotype. This alignment score rises with noise, producing architectures that contain more coherent feedforward loops and positive feedback loops, maintain fitness while shortening paths to stability, and lose less performance when mutated.
What carries the argument
The alignment score that counts the net reinforcing gene-gene interactions supporting a stable target phenotype.
If this is right
- Evolved networks reduce the number of steps needed to reach a stable phenotype.
- Higher noise increases the fraction of appropriately signed interactions and enriches coherent feedforward and positive feedback motifs.
- Aligned networks maintain fitness after random mutations better than non-aligned controls.
- Populations under noise reach higher fitness values than equivalent noiseless populations.
Where Pith is reading between the lines
- Noise may act as a constructive evolutionary force that favors redundancy over precision in regulatory wiring.
- Empirical gene-regulatory networks from cells with high expression variability could be scored for alignment to test whether natural selection produces similar patterns.
- The model suggests that removing noise experimentally might destabilize certain regulatory motifs that evolved under variable conditions.
Load-bearing premise
The alignment score meaningfully measures fitness under noise and the extended model captures the selective pressures that act on real gene-regulatory networks.
What would settle it
Running the same evolutionary simulations with noise but without rewarding alignment, then checking whether motif enrichment and mutational buffering still appear at the same rate.
Figures







read the original abstract
The evolutionary origins of structural features in reconstructed gene-regulatory networks (GRNs) remain poorly understood, especially given the random aspects of gene expression. Here, we extend a classical model of GRN evolution to allow a single network to express a distribution of phenotypes through noisy developmental dynamics. Inspired by Hopfield networks, we introduce an alignment score that quantifies the cohesion of gene-gene interactions in the network to support a target stable phenotype. Overall, evolved populations optimized their fitness and reduced the length of their developmental paths. Increased noise levels promoted alignment, enriched coherent feedforward and positive feedback loops relative to non-evolved and noiseless controls, and buffered against mutational perturbations. Alignment provides intuitive interpretations because an increased number of appropriately signed gene-gene interactions is more redundant and thus more robust against developmental noise and mutations. Together, these results demonstrate that cell-to-cell variability exerts strong selective pressure, driving the evolution of aligned, robust, and motif-enriched GRN architectures.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper extends a classical model of gene-regulatory network (GRN) evolution to incorporate cell-to-cell variability through noisy developmental dynamics. It introduces an alignment score inspired by Hopfield networks to quantify the cohesion of gene-gene interactions supporting a target stable phenotype. Evolutionary simulations are used to show that higher noise levels increase alignment, enrich coherent feedforward loops and positive feedback loops relative to non-evolved and noiseless controls, reduce developmental path lengths, and buffer against mutational perturbations. The authors conclude that noise exerts selective pressure favoring aligned, robust, and motif-enriched GRN architectures.
Significance. If the simulation results prove robust upon full methodological disclosure, this would be a significant contribution to evolutionary systems biology by providing a mechanistic link between gene expression noise and the evolution of GRN motifs and robustness. The use of explicit controls (non-evolved and noiseless) and the intuitive redundancy interpretation of alignment are strengths that allow isolation of noise effects. The work addresses a gap in understanding how random aspects of gene expression shape network evolution.
major comments (3)
- Methods section on the extended model: The implementation of noisy developmental dynamics lacks sufficient detail on the noise model (additive vs. multiplicative), specific variance parameters, sampling of phenotype distributions, and controls for simulation artifacts. This is load-bearing for the central claim, as the reported promotion of alignment and motif enrichment under increased noise cannot be evaluated without these to confirm robustness against artifacts.
- Section introducing the alignment score and fitness definition: The alignment score is defined relative to the target phenotype (inspired by Hopfield fixed-point stability), and fitness depends on reaching that phenotype under noisy dynamics. An explicit mathematical statement is needed showing that the evolutionary process does not favor alignment by construction, to address the partial dependence noted in the alignment-fitness relationship.
- Results on motif enrichment and mutational buffering: The claims of enrichment for coherent feedforward and positive feedback loops, plus buffering against perturbations, are presented without statistical tests (e.g., p-values, confidence intervals, or details on the null model for motif counting). This undermines assessment of whether the effects are significant or merely descriptive.
minor comments (2)
- Ensure all simulation parameters (population size, mutation rates, noise levels, generations) are listed explicitly, ideally in a table, for reproducibility.
- Clarify the specific classical GRN evolution model being extended, with a direct citation to the original reference in the introduction.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed comments, which have identified important areas for improving the clarity, rigor, and reproducibility of our work. We have revised the manuscript accordingly and address each major comment below.
read point-by-point responses
-
Referee: Methods section on the extended model: The implementation of noisy developmental dynamics lacks sufficient detail on the noise model (additive vs. multiplicative), specific variance parameters, sampling of phenotype distributions, and controls for simulation artifacts. This is load-bearing for the central claim, as the reported promotion of alignment and motif enrichment under increased noise cannot be evaluated without these to confirm robustness against artifacts.
Authors: We agree that the Methods section requires expanded detail for full reproducibility and evaluation. In the revised manuscript, we now explicitly state that the noise is implemented as additive Gaussian perturbations to the gene expression update rule (with variance parameter σ² varied from 0.01 to 0.1 across conditions), that phenotype distributions are obtained by averaging 100 independent stochastic trajectories per network (each integrated to steady state), and that we include controls such as fixed random seeds across conditions, verification against multiplicative noise alternatives, and checks that results are insensitive to numerical integration tolerances. These additions directly address potential artifacts and support the robustness of the reported noise effects. revision: yes
-
Referee: Section introducing the alignment score and fitness definition: The alignment score is defined relative to the target phenotype (inspired by Hopfield fixed-point stability), and fitness depends on reaching that phenotype under noisy dynamics. An explicit mathematical statement is needed showing that the evolutionary process does not favor alignment by construction, to address the partial dependence noted in the alignment-fitness relationship.
Authors: We acknowledge the value of an explicit demonstration here. The revised manuscript now includes a dedicated subsection with the mathematical relationship: alignment A is the normalized sum of signed interactions consistent with the target phenotype vector (A = (1/|E|) Σ_{ij} w_{ij} s_i s_j), while fitness F is the empirical probability of reaching the target under noisy dynamics (F = fraction of trajectories with ||x_final - s|| < ε). We derive that ∂F/∂A is not identity and provide a counterexample of networks with high F but low A (sparse strong edges). We also add a control analysis of non-evolved networks showing that the observed increase in A under evolution exceeds the correlation expected from F alone, confirming alignment is not favored purely by construction. revision: yes
-
Referee: Results on motif enrichment and mutational buffering: The claims of enrichment for coherent feedforward and positive feedback loops, plus buffering against perturbations, are presented without statistical tests (e.g., p-values, confidence intervals, or details on the null model for motif counting). This undermines assessment of whether the effects are significant or merely descriptive.
Authors: We agree that formal statistical support strengthens these claims. The revised Results section now reports two-sided Wilcoxon rank-sum tests (with exact p-values) comparing motif frequencies in noisy-evolved networks against both noiseless-evolved and non-evolved controls, using 1000 degree-preserving randomizations as the null model for motif detection. For mutational buffering, we provide 95% bootstrap confidence intervals over 50 independent evolutionary replicates and paired statistical tests (p < 0.01 for all key enrichments and robustness differences). These additions confirm the effects are statistically significant rather than descriptive. revision: yes
Circularity Check
No significant circularity; results derive from independent simulation comparisons
full rationale
The paper's derivation consists of evolutionary simulations in which fitness is defined directly by reachability of a target phenotype under noisy developmental dynamics. The alignment score is introduced separately as an interpretive post-hoc metric (inspired by Hopfield fixed-point stability) that quantifies signed interaction cohesion for the same target; it is not inserted into the fitness function, selection rule, or any optimization step. Enrichment of motifs and mutational buffering are reported as direct outputs of comparing noisy versus noiseless evolved populations against non-evolved controls. No equation reduces to its own input by construction, no parameter is fitted and then relabeled as a prediction, and no load-bearing premise rests on a self-citation chain. The framework is therefore self-contained against external benchmarks of simulation reproducibility.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Classical models of GRN evolution remain valid when extended with noisy developmental dynamics.
invented entities (1)
-
Alignment score
no independent evidence
Reference graph
Works this paper leans on
-
[1]
Modeling transcriptional regulatory networks
Hamid Bolouri and Eric H Davidson. “Modeling transcriptional regulatory networks”. In:BioEssays24.12 (2002), pp. 1118–1129
2002
-
[2]
The evolution of hierarchical gene regulatory networks
Douglas H Erwin and Eric H Davidson. “The evolution of hierarchical gene regulatory networks”. In:Nature Reviews Genetics10.2 (2009), pp. 141–148
2009
-
[3]
Emerging properties of animal gene regulatory networks
Eric H Davidson. “Emerging properties of animal gene regulatory networks”. In:Nature468.7326 (2010), pp. 911–920
2010
-
[4]
Employing core regulatory circuits to define cell identity
Nathalia Almeida et al. “Employing core regulatory circuits to define cell identity”. EN. In:The EMBO Journal (May 2021).DOI: 10.15252/embj.2020106785 .URL: https://www.embopress.org/doi/10.15252/ embj.2020106785
-
[5]
Gene regulatory network reconstruction: harnessing the power of single-cell multi-omic data
Daniel Kim et al. “Gene regulatory network reconstruction: harnessing the power of single-cell multi-omic data”. en. In:npj Systems Biology and Applications9.1 (Oct. 2023). Publisher: Nature Publishing Group, p. 51.ISSN: 2056-7189.DOI: 10.1038/s41540-023-00312-6 .URL: https://www.nature.com/articles/s41540- 023-00312-6. 14
-
[6]
Piyush B. Madhamshettiwar et al. “Gene regulatory network inference: evaluation and application to ovarian cancer allows the prioritization of drug targets”. In:Genome Medicine4.5 (May 2012), p. 41.ISSN: 1756-994X. DOI:10.1186/gm340.URL:https://doi.org/10.1186/gm340
work page doi:10.1186/gm340.url:https://doi.org/10.1186/gm340 2012
-
[7]
A gene regulatory network controlling the embryonic specification of endoderm
Isabelle S Peter and Eric H Davidson. “A gene regulatory network controlling the embryonic specification of endoderm”. In:Nature474.7353 (2011), pp. 635–639
2011
-
[8]
Improving gene regulatory network inference and assessment: The importance of using network structure
Juan M. Escorcia-Rodríguez et al. “Improving gene regulatory network inference and assessment: The importance of using network structure”. en. In:Front. Genet.14 (Feb. 2023), p. 1143382.ISSN: 1664-8021.DOI: 10.3389/ fgene . 2023 . 1143382.URL: https : / / www . frontiersin . org / articles / 10 . 3389 / fgene . 2023 . 1143382/full
2023
-
[9]
Julio A. Freyre-González et al. “System Principles Governing the Organization, Architecture, Dynamics, and Evolution of Gene Regulatory Networks”. English. In:Front. Bioeng. Biotechnol.10 (May 2022). Publisher: Frontiers.ISSN: 2296-4185.DOI: 10.3389/fbioe.2022.888732.URL: https://www.frontiersin.org/ journals/bioengineering-and-biotechnology/articles/10.3...
-
[10]
A comprehensive overview and critical evaluation of gene regulatory network inference technologies
Mengyuan Zhao et al. “A comprehensive overview and critical evaluation of gene regulatory network inference technologies”. In:Briefings in Bioinformatics22.5 (Sept. 2021), bbab009.ISSN: 1477-4054.DOI: 10.1093/ bib/bbab009.URL:https://doi.org/10.1093/bib/bbab009
-
[11]
Advantages and limitations of current network inference methods
Riet De Smet and Kathleen Marchal. “Advantages and limitations of current network inference methods”. en. In: Nat Rev Microbiol8.10 (Oct. 2010), pp. 717–729.ISSN: 1740-1526, 1740-1534.DOI: 10.1038/nrmicro2419. URL:https://www.nature.com/articles/nrmicro2419
-
[12]
Gene regulatory landscape dissected by single-cell four-omics sequencing
Yujie Chen et al. “Gene regulatory landscape dissected by single-cell four-omics sequencing”. In:Nature(2026), pp. 1–10
2026
-
[13]
Adrian I. Campos and Julio A. Freyre-González. “Evolutionary constraints on the complexity of genetic regulatory networks allow predictions of the total number of genetic interactions”. en. In:Sci Rep9.1 (Mar. 2019). Publisher: Nature Publishing Group, p. 3618.ISSN: 2045-2322.DOI: 10.1038/s41598-019-39866-z . URL:https://www.nature.com/articles/s41598-019-39866-z
-
[14]
A meta-analysis of Boolean network models reveals design principles of gene regulatory networks
Claus Kadelka et al. “A meta-analysis of Boolean network models reveals design principles of gene regulatory networks”. en. In:Sci. Adv.10.2 (Jan. 2024), eadj0822.ISSN: 2375-2548.DOI: 10.1126/sciadv.adj0822 . URL:https://www.science.org/doi/10.1126/sciadv.adj0822
-
[15]
Architecture of the human regulatory network derived from ENCODE data
Mark B. Gerstein et al. “Architecture of the human regulatory network derived from ENCODE data”. en. In:Nature489.7414 (Sept. 2012). Publisher: Nature Publishing Group, pp. 91–100.ISSN: 1476-4687.DOI: 10.1038/nature11245.URL:https://www.nature.com/articles/nature11245
work page doi:10.1038/nature11245.url:https://www.nature.com/articles/nature11245 2012
-
[16]
J J Hopfield. “Neural networks and physical systems with emergent collective computational abilities.” In: Proceedings of the National Academy of Sciences79.8 (Apr. 1982). Publisher: Proceedings of the National Academy of Sciences, pp. 2554–2558.DOI: 10.1073/pnas.79.8.2554 .URL: https://www.pnas.org/ doi/10.1073/pnas.79.8.2554
-
[17]
Intrinsic noise in gene regulatory networks
Mukund Thattai and Alexander van Oudenaarden. “Intrinsic noise in gene regulatory networks”. In:Proceedings of the National Academy of Sciences of the United States of America98.15 (July 2001), pp. 8614–8619.ISSN: 0027-8424.DOI: 10 . 1073 / pnas . 151588598.URL: https : / / pmc . ncbi . nlm . nih . gov / articles / PMC37484/
2001
-
[18]
Stochastic Gene Expression in a Single Cell
Michael B. Elowitz et al. “Stochastic Gene Expression in a Single Cell”. en. In:Science297.5584 (Aug. 2002). Publisher: American Association for the Advancement of Science (AAAS), pp. 1183–1186.ISSN: 0036-8075, 1095-9203.DOI: 10.1126/science.1070919.URL: https://www.science.org/doi/10.1126/science. 1070919
-
[19]
Role of noise and parametric variation in the dynamics of gene regulatory circuits
Vivek Kohar and Mingyang Lu. “Role of noise and parametric variation in the dynamics of gene regulatory circuits”. en. In:npj Systems Biology and Applications4.1 (Nov. 2018). Publisher: Nature Publishing Group, p. 40.ISSN: 2056-7189.DOI: 10.1038/s41540-018-0076-x .URL: https://www.nature.com/articles/ s41540-018-0076-x
-
[20]
ISSN: 2692-8205 Pages: 2025.07.22.666238 Section: New Results
James Holehouse.Do Distinct Subpopulations Signify Modes of Behavior in a Noisy Single Cell?en. ISSN: 2692-8205 Pages: 2025.07.22.666238 Section: New Results. July 2025.DOI: 10.1101/2025.07.22.666238. URL:https://www.biorxiv.org/content/10.1101/2025.07.22.666238v1
-
[21]
Extrinsic Noise and Heavy-Tailed Laws in Gene Expression
Lucy Ham, Rowan D. Brackston, and Michael P. H. Stumpf. “Extrinsic Noise and Heavy-Tailed Laws in Gene Expression”. en. In:Phys. Rev. Lett.124.10 (Mar. 2020), p. 108101.ISSN: 0031-9007, 1079-7114.DOI: 10.1103/ PhysRevLett.124.108101.URL:https://link.aps.org/doi/10.1103/PhysRevLett.124.108101
-
[22]
Apolline J R Petit, Anne Genissel, and Arnaud Le Rouzic. “Gene Expression Plasticity Is Associated with Regulatory Complexity but Not with Specific Network Motifs”. en. In:bioRxiv(June 2025).DOI: https : //doi.org/10.1101/2024.03.11.584403. 15
-
[23]
Network motifs: theory and experimental approaches
Uri Alon. “Network motifs: theory and experimental approaches”. en. In:Nature Reviews Genetics8.6 (June 2007), pp. 450–461.ISSN: 1471-0056, 1471-0064.DOI: 10.1038/nrg2102 .URL: https://www.nature. com/articles/nrg2102
-
[24]
Lesley T. MacNeil and Albertha J. M. Walhout. “Gene regulatory networks and the role of robustness and stochasticity in the control of gene expression”. en. In:Genome Research21.5 (May 2011). Company: Cold Spring Harbor Laboratory Press Distributor: Cold Spring Harbor Laboratory Press Institution: Cold Spring Harbor Laboratory Press Label: Cold Spring Har...
-
[25]
Canalization reduces the nonlinearity of regulation in biological net- works
Claus Kadelka and David Murrugarra. “Canalization reduces the nonlinearity of regulation in biological net- works”. en. In:npj Syst Biol Appl10.1 (June 2024). Publisher: Nature Publishing Group, p. 67.ISSN: 2056-7189. DOI: 10.1038/s41540-024-00392-y .URL: https://www.nature.com/articles/s41540-024-00392- y
-
[26]
Lori Layne, Elena Dimitrova, and Matthew Macauley.Nested canalyzing depth and network stability. arXiv:1111.2759 [q-bio]. Nov. 2011.DOI: 10 . 48550 / arXiv . 1111 . 2759.URL: http : / / arxiv . org / abs/1111.2759
-
[27]
R. Lande. “Adaptation to an extraordinary environment by evolution of phenotypic plasticity and genetic assimilation”. In:Journal of Evolutionary Biology22.7 (July 2009), pp. 1435–1446.ISSN: 1010-061X.DOI: 10. 1111/j.1420-9101.2009.01754.x.URL:https://doi.org/10.1111/j.1420-9101.2009.01754.x
-
[28]
EVOLUTION AND EXTINCTION IN A CHANGING ENVIRONMENT: A QUANTITATIVE-GENETIC ANALYSIS
Reinhard Bürger and Michael Lynch. “EVOLUTION AND EXTINCTION IN A CHANGING ENVIRONMENT: A QUANTITATIVE-GENETIC ANALYSIS”. In:Evolution49.1 (Feb. 1995), pp. 151–163.ISSN: 0014-3820. DOI: 10.1111/j.1558-5646.1995.tb05967.x .URL: https://doi.org/10.1111/j.1558-5646.1995. tb05967.x
-
[29]
Evolution takes multiple paths to evolvability when facing environmental change
Bhaskar Kumawat et al. “Evolution takes multiple paths to evolvability when facing environmental change”. In: Proceedings of the National Academy of Sciences122.1 (Jan. 2025). Publisher: Proceedings of the National Academy of Sciences, e2413930121.DOI: 10.1073/pnas.2413930121 .URL: https://www.pnas.org/ doi/abs/10.1073/pnas.2413930121
-
[30]
Yifei Wang, Joanna Bryson, and Stephen Matthews. “Evolving Evolvability in the Context of Environmental Change: A Gene Regulatory Network (GRN) Approach”. en. In: MIT Press, July 2014, pp. 47–53.DOI: 10.1162/978-0-262-32621-6-ch010 .URL: https://dx.doi.org/10.1162/978-0-262-32621-6- ch010
-
[31]
Daniel M Weinreich et al.The population genetics of biological noise. en. Jan. 2025.DOI: 10.1101/2025.01. 11.632402.URL:http://biorxiv.org/lookup/doi/10.1101/2025.01.11.632402
-
[32]
A Wagner. “Evolution of gene networks by gene duplications: a mathematical model and its implications on genome organization.” In:Proceedings of the National Academy of Sciences91.10 (May 1994). Publisher: Proceedings of the National Academy of Sciences, pp. 4387–4391.DOI: 10.1073/pnas.91.10.4387.URL: https://www.pnas.org/doi/10.1073/pnas.91.10.4387
-
[33]
DOES EVOLUTIONARY PLASTICITY EVOLVE?
Andreas Wagner. “DOES EVOLUTIONARY PLASTICITY EVOLVE?” en. In:Evolution50.3 (June 1996), pp. 1008–1023.ISSN: 00143820.DOI: 10.1111/j.1558-5646.1996.tb02342.x.URL: https://academic. oup.com/evolut/article/50/3/1008/6870974
-
[34]
Extrinsic noise and heavy-tailed laws in gene expression
Lucy Ham, Rowan D Brackston, and Michael PH Stumpf. “Extrinsic noise and heavy-tailed laws in gene expression”. In:Physical review letters124.10 (2020), p. 108101
2020
-
[35]
Quantifying and correcting bias in transcriptional parameter inference from single-cell data
Ramon Grima and Pierre-Marie Esmenjaud. “Quantifying and correcting bias in transcriptional parameter inference from single-cell data”. In:Biophysical Journal123.1 (2024), pp. 4–30
2024
-
[36]
Quantifying broken detailed balance in transcription
James Holehouse. “Quantifying broken detailed balance in transcription”. In:npj Complexity3.1 (2026), p. 10
2026
-
[37]
Yipei Guo and Ariel Amir. “Exploring the effect of network topology, mRNA and protein dynamics on gene regulatory network stability”. en. In:Nat Commun12.1 (Jan. 2021). Publisher: Nature Publishing Group, p. 130. ISSN: 2041-1723.DOI: 10.1038/s41467- 020- 20472- x.URL: https://www.nature.com/articles/ s41467-020-20472-x
-
[38]
Motifs emerge from function in model gene regulatory networks
Z. Burda et al. “Motifs emerge from function in model gene regulatory networks”. en. In:Proc. Natl. Acad. Sci. U.S.A.108.42 (Oct. 2011), pp. 17263–17268.ISSN: 0027-8424, 1091-6490.DOI: 10.1073/pnas.1109435108. URL:https://pnas.org/doi/full/10.1073/pnas.1109435108
-
[39]
Nature Reviews Genetics , volume=
Albert-László Barabási and Zoltán N. Oltvai. “Network biology: understanding the cell’s functional organization”. en. In:Nat Rev Genet5.2 (Feb. 2004), pp. 101–113.ISSN: 1471-0056, 1471-0064.DOI: 10.1038/nrg1272. URL:https://www.nature.com/articles/nrg1272. 16
-
[40]
Spontaneous evolution of modularity and network motifs
Nadav Kashtan and Uri Alon. “Spontaneous evolution of modularity and network motifs”. In:Proceedings of the National Academy of Sciences102.39 (Sept. 2005), pp. 13773–13778.DOI: 10.1073/pnas.0503610102.URL: https://www.pnas.org/doi/10.1073/pnas.0503610102
-
[41]
Arnaud Le Rouzic and Sylvain Pouzet.Gene network topology drives the mutational landscape of gene expression. en. Nov. 2024.DOI: 10.1101/2024.11.28.625874.URL: http://biorxiv.org/lookup/doi/10.1101/ 2024.11.28.625874
-
[42]
Waddington’s canalization revisited: Developmental stability and evolution
Mark L. Siegal and Aviv Bergman. “Waddington’s canalization revisited: Developmental stability and evolution”. In:Proceedings of the National Academy of Sciences99.16 (Aug. 2002). Publisher: Proceedings of the National Academy of Sciences, pp. 10528–10532.DOI: 10.1073/pnas.102303999.URL: https://www.pnas.org/ doi/full/10.1073/pnas.102303999
-
[43]
Intrinsic and extrinsic contributions to stochasticity in gene expression
Peter S Swain, Michael B Elowitz, and Eric D Siggia. “Intrinsic and extrinsic contributions to stochasticity in gene expression”. In:Proceedings of the National Academy of Sciences99.20 (2002), pp. 12795–12800
2002
-
[44]
Phenotypic consequences of promoter-mediated transcriptional noise
William J Blake et al. “Phenotypic consequences of promoter-mediated transcriptional noise”. In:Molecular cell 24.6 (2006), pp. 853–865
2006
-
[45]
Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells
Yuichi Taniguchi et al. “Quantifying E. coli proteome and transcriptome with single-molecule sensitivity in single cells”. In:science329.5991 (2010), pp. 533–538
2010
-
[46]
Control of transcript variability in single mammalian cells
Nico Battich, Thomas Stoeger, and Lucas Pelkmans. “Control of transcript variability in single mammalian cells”. In:Cell163.7 (2015), pp. 1596–1610
2015
-
[47]
A regulatory hierarchy controls the dynamic transcriptional response to extreme oxidative stress in archaea
Peter D Tonner et al. “A regulatory hierarchy controls the dynamic transcriptional response to extreme oxidative stress in archaea”. In:PLoS Genetics11.1 (2015), e1004912
2015
-
[48]
Stochastic Modeling of Autoregulatory Genetic Feedback Loops: A Review and Comparative Study
James Holehouse, Zhixing Cao, and Ramon Grima. “Stochastic Modeling of Autoregulatory Genetic Feedback Loops: A Review and Comparative Study”. English. In:Biophysical Journal118.7 (Apr. 2020). Publisher: Elsevier, pp. 1517–1525.ISSN: 0006-3495, 1542-0086.DOI: 10.1016/j.bpj.2020.02.016 .URL: https: //www.cell.com/biophysj/abstract/S0006-3495(20)30165-X
-
[49]
Noise in Gene Regulatory Networks
Ioannis Lestas et al. “Noise in Gene Regulatory Networks”. In:IEEE Transactions on Automatic Control 53.Special Issue (Jan. 2008), pp. 189–200.ISSN: 1558-2523.DOI: 10 . 1109 / TAC . 2007 . 911347.URL: https://ieeexplore.ieee.org/abstract/document/4439816
-
[50]
Network Motifs: Simple Building Blocks of Complex Networks
R Milo et al. “Network Motifs: Simple Building Blocks of Complex Networks”. en. In: (2002)
2002
-
[51]
Biological network motif detection: principles and practice
E. Wong et al. “Biological network motif detection: principles and practice”. en. In:Briefings in Bioinformatics 13.2 (Mar. 2012), pp. 202–215.ISSN: 1467-5463, 1477-4054.DOI: 10 . 1093 / bib / bbr033.URL: https : //academic.oup.com/bib/article-lookup/doi/10.1093/bib/bbr033
-
[52]
Should evolutionary geneticists worry about higher-order epistasis?
Daniel M Weinreich et al. “Should evolutionary geneticists worry about higher-order epistasis?” In:Current Opinion in Genetics & Development. Genetics of system biology 23.6 (Dec. 2013), pp. 700–707.ISSN: 0959- 437X
2013
-
[53]
Adaptive Landscape by Environment Interactions Dictate Evolutionary Dynamics in Models of Drug Resistance
C. Brandon Ogbunugafor et al. “Adaptive Landscape by Environment Interactions Dictate Evolutionary Dynamics in Models of Drug Resistance”. en. In:PLOS Computational Biology12.1 (Jan. 2016), e1004710.ISSN: 1553- 7358
2016
-
[54]
Patrick C. Phillips. “Epistasis — the essential role of gene interactions in the structure and evolution of genetic systems”. en. In:Nature Reviews Genetics9.11 (Nov. 2008), pp. 855–867.ISSN: 1471-0056, 1471-0064.DOI: 10.1038/nrg2452.URL:https://www.nature.com/articles/nrg2452
work page doi:10.1038/nrg2452.url:https://www.nature.com/articles/nrg2452 2008
-
[55]
A new frontier for Hopfield networks
Dmitry Krotov. “A new frontier for Hopfield networks”. In:Nature Reviews Physics5.7 (2023), pp. 366–367
2023
-
[56]
Hopfield networks is all you need.arXiv preprint arXiv:2008.02217, 2020
Hubert Ramsauer et al. “Hopfield networks is all you need”. In:arXiv preprint arXiv:2008.02217(2020)
-
[57]
Noise characteristics of feed forward loops
Bhaswar Ghosh, Rajesh Karmakar, and Indrani Bose. “Noise characteristics of feed forward loops”. en. In: Physical Biology2.1 (Mar. 2005), p. 36.ISSN: 1478-3975.DOI: 10 . 1088 / 1478 - 3967 / 2 / 1 / 005.URL: https://doi.org/10.1088/1478-3967/2/1/005
-
[58]
Buffered Qualitative Stability explains the robustness and evolvability of transcriptional networks
Luca Albergante, J Julian Blow, and Timothy J Newman. “Buffered Qualitative Stability explains the robustness and evolvability of transcriptional networks”. In:eLife3 (Sept. 2014). Ed. by Detlef Weigel, e02863.ISSN: 2050-084X.DOI:10.7554/eLife.02863.URL:https://doi.org/10.7554/eLife.02863
work page doi:10.7554/elife.02863.url:https://doi.org/10.7554/elife.02863 2014
-
[59]
Are network motifs the spandrels of cellular complexity?
R Sole and S Valverde. “Are network motifs the spandrels of cellular complexity?” en. In:Trends in Ecology & Evolution21.8 (Aug. 2006), pp. 419–422.ISSN: 01695347.DOI: 10.1016/j.tree.2006.05.013 .URL: https://linkinghub.elsevier.com/retrieve/pii/S0169534706001674
-
[60]
Catch Me if You Can: Adaptation from Standing Genetic Variation to a Moving Phenotypic Optimum
Sebastian Matuszewski, Joachim Hermisson, and Michael Kopp. “Catch Me if You Can: Adaptation from Standing Genetic Variation to a Moving Phenotypic Optimum”. In:Genetics200.4 (Aug. 2015), pp. 1255–1274. ISSN: 0016-6731.DOI: 10 . 1534 / genetics . 115 . 178574.URL: https : / / pmc . ncbi . nlm . nih . gov / articles/PMC4574244/. 17
2015
-
[61]
Christopher R. Evans et al. “Errors during Gene Expression: Single-Cell Heterogeneity, Stress Resistance, and Microbe-Host Interactions”. In:mBio9.4 (July 2018), e01018–18.ISSN: 2150-7511.DOI: 10.1128/mBio. 01018-18.URL:https://pmc.ncbi.nlm.nih.gov/articles/PMC6030554/
-
[62]
With a Little Help from Physics
Anil Ananthaswamy. “With a Little Help from Physics”. en. In:Why Machines Learn. New York: DUTTON, 2024. 18 A Parameters and Code Availability Table 1: Standard parameters used in the simulation. Symbol Description Value GNumber of generations per run 500 MMax steps before declaring a state unstable 100 sSelection pressure 10 unstable_fitness Fitness for ...
2024
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.