Recognition: unknown
Factor State Space Modelling of the Ornstein-Uhlenbeck Process with Measurement Error and its Application
Pith reviewed 2026-05-10 15:00 UTC · model grok-4.3
The pith
A factor structure resolves identifiability for the multivariate Ornstein-Uhlenbeck state space model with measurement error.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The factor OUSSM represents the latent mean-reverting dynamics through a lower-dimensional factor loading matrix applied to an OU process, with the observation equation containing additive measurement error; necessary linear constraints on loadings and variances restore identifiability so that maximum-likelihood estimation recovers the drift, diffusion, and noise parameters consistently.
What carries the argument
The factor loading matrix that compresses the high-dimensional latent OU state while the measurement-error term in the observation equation remains separate.
If this is right
- Multivariate time series with mean reversion can be analyzed without systematic bias from ignored measurement error.
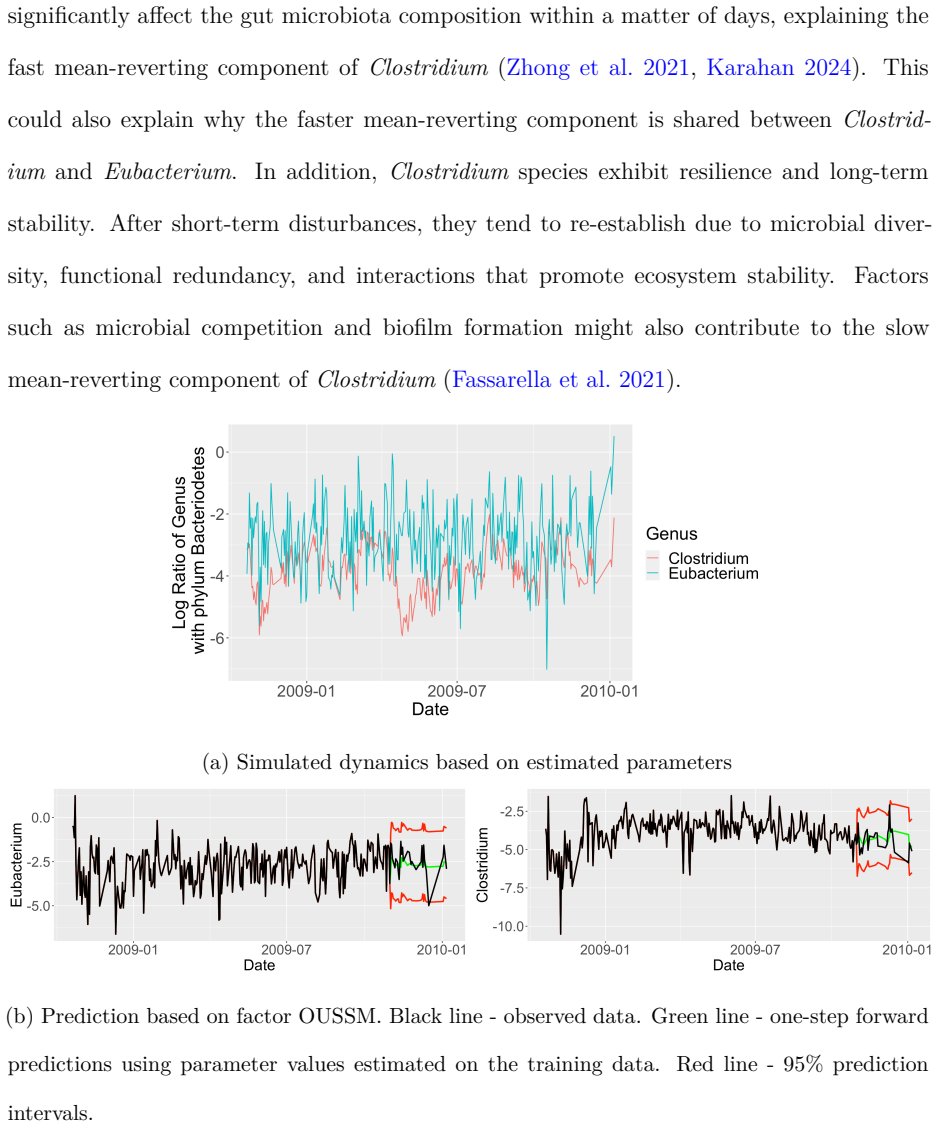
- Latent temporal structures in biological systems such as the gut microbiome become recoverable as distinct factor-driven OU processes.
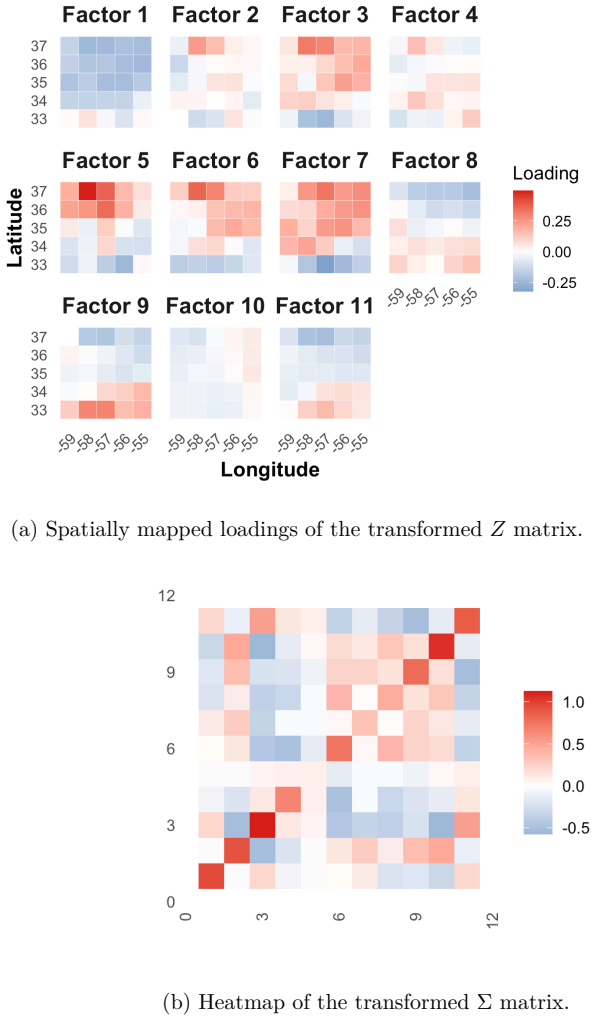
- Environmental series like sea-surface temperature can be decomposed into lower-dimensional mean-reverting components.
- The same constrained estimation procedure applies to any collection of noisy, mean-reverting variables once the factor dimension is chosen.
Where Pith is reading between the lines
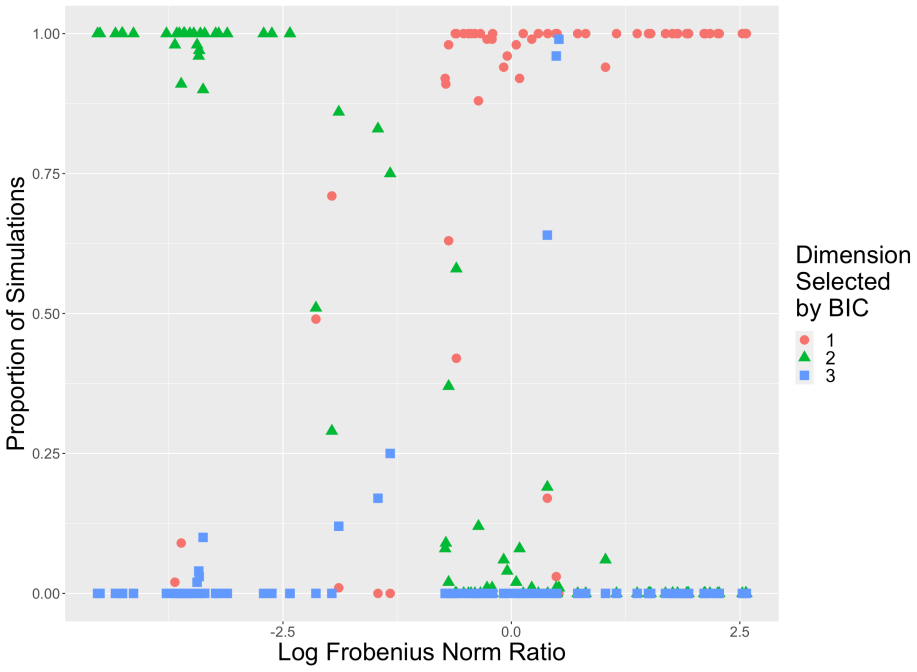
- The method could be paired with model-selection rules for choosing the number of factors when that choice is not obvious from domain knowledge.
- If the factor assumption is too restrictive for a given dataset, the recovered latent paths would show systematic residuals that a full-dimensional OUSSM might avoid.
- Forecasting skill in microbiome or climate applications would improve only if the latent factors truly capture the dominant reversion dynamics rather than merely fitting noise.
Load-bearing premise
The imposed factor structure and constraints preserve the true latent mean-reversion rates and do not create spurious temporal correlations in the data.
What would settle it
Simulate multivariate OU trajectories with known parameters and added measurement noise, fit the factor OUSSM under the stated constraints, and check whether the recovered drift matrix and reversion speeds deviate substantially from the generating values.
Figures




read the original abstract
Standard Ornstein-Uhlenbeck (OU) models often yield biased parameter estimates when measurement error is ignored. While the Ornstein-Uhlenbeck State Space Model (OUSSM) addresses this in univariate settings, multidimensional extensions remain limited. This paper introduces the factor OUSSM to model multi-dimensional, mean-reverting systems with observational noise. We resolve critical identifiability challenges in parameter estimation by establishing necessary constraints and validating the method through extensive simulations. We demonstrate the model's versatility by analyzing human gut microbiome dynamics and North Atlantic Sea Surface Temperature (SST) data. The results reveal distinct latent temporal structures in both biological and environmental systems, establishing the factor OUSSM as a robust framework for multivariate time series analysis.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces the factor Ornstein-Uhlenbeck State Space Model (factor OUSSM) to extend univariate OUSSM to multivariate mean-reverting processes with observational noise. It resolves identifiability challenges via parameter constraints on the factor loadings, drift, and diffusion matrices, validates the estimator through simulations, and applies the model to human gut microbiome time series and North Atlantic SST data to extract latent temporal structures.
Significance. If the imposed factor structure and identifiability constraints can be shown not to materially bias the latent mean-reversion dynamics, the framework offers a practical dimensionality-reduction approach for high-dimensional OU processes with measurement error. This combination of state-space modeling and factor analysis addresses a recurring need in applied statistics for biological and environmental time series.
major comments (2)
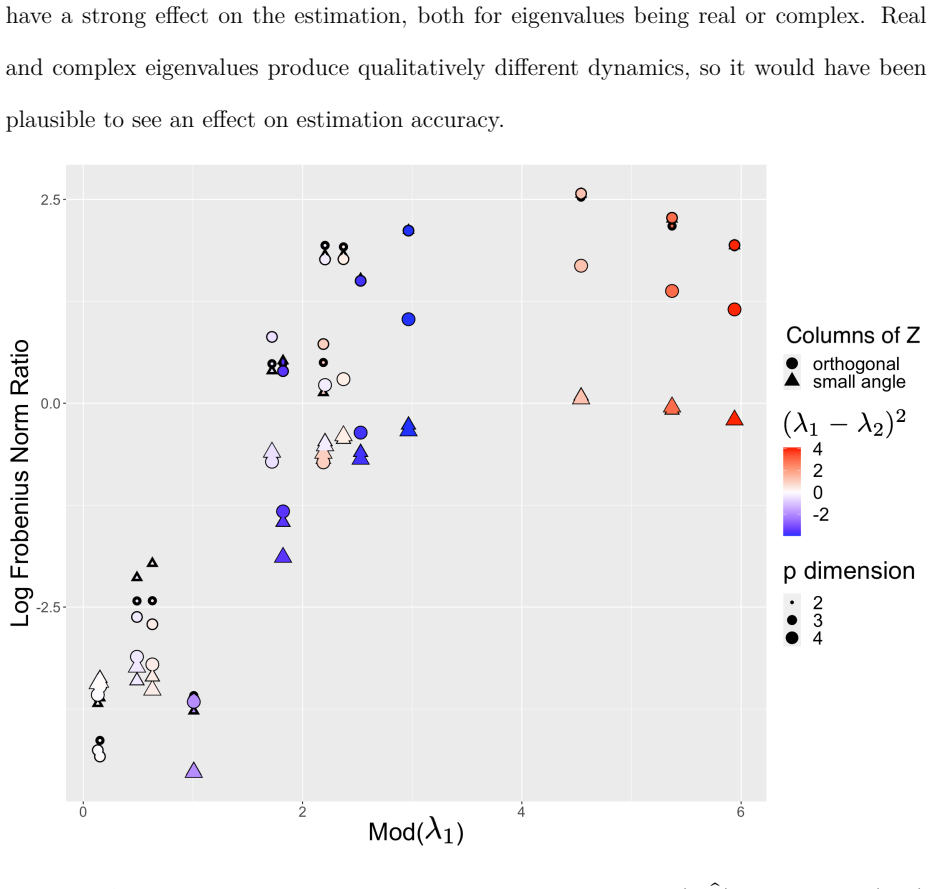
- [Simulation study] The simulation validation (described in the abstract as confirming the estimator) is generated from the same constrained factor OUSSM. This design cannot detect distortion of the latent OU trajectories or mean-reversion rates when the true data-generating process lies outside the allowable space of the reduced-rank loading matrix and constraints.
- [Applications] The applications to microbiome and SST data claim revelation of distinct latent temporal structures, yet no comparison to unconstrained multivariate OU formulations or sensitivity checks on the factor rank is reported. This leaves the central assumption—that the low-rank representation is faithful rather than artifactual—untested and load-bearing for the empirical conclusions.
minor comments (2)
- The abstract would benefit from a concise statement of the dimension of the factor space and the explicit form of the identifiability constraints (e.g., on the drift matrix or observation equation).
- Notation distinguishing the factor loading matrix from the OU drift and diffusion parameters should be introduced early and used consistently.
Simulated Author's Rebuttal
We thank the referee for the constructive comments, which highlight important aspects of model validation and empirical robustness. We address each major comment below and commit to revisions that strengthen the manuscript without altering its core contributions.
read point-by-point responses
-
Referee: [Simulation study] The simulation validation (described in the abstract as confirming the estimator) is generated from the same constrained factor OUSSM. This design cannot detect distortion of the latent OU trajectories or mean-reversion rates when the true data-generating process lies outside the allowable space of the reduced-rank loading matrix and constraints.
Authors: We agree that the simulations validate estimator performance under correct specification of the factor OUSSM and its identifiability constraints. This is the standard approach for new estimators, but it does not address potential bias under misspecification. In the revised manuscript we will add a new simulation subsection generating data from full-rank OU processes (with and without measurement error) and from processes violating the loading constraints. We will report bias and coverage for the recovered mean-reversion rates and latent trajectories, together with a discussion of when the factor constraints materially affect inference. revision: yes
-
Referee: [Applications] The applications to microbiome and SST data claim revelation of distinct latent temporal structures, yet no comparison to unconstrained multivariate OU formulations or sensitivity checks on the factor rank is reported. This leaves the central assumption—that the low-rank representation is faithful rather than artifactual—untested and load-bearing for the empirical conclusions.
Authors: We acknowledge that the current applications would be strengthened by explicit sensitivity checks and, where feasible, comparisons. In the revision we will add results for factor ranks k = 2, 3, 4, 5 on both datasets, showing stability of the extracted mean-reversion rates and latent factor interpretations. Direct comparison to fully unconstrained high-dimensional OU-SSM is limited by identifiability and computational cost—the very issues that motivate the factor structure—but we will include a low-dimensional unconstrained benchmark on variable subsets and discuss the trade-offs. These additions will make the faithfulness of the low-rank assumption more transparent. revision: yes
Circularity Check
No significant circularity; derivation self-contained against external benchmarks
full rationale
The abstract and reader's summary present the factor OUSSM as introducing a reduced-rank state-space extension of the multivariate OU process, with identifiability constraints derived from the model structure to resolve rotational freedom in the drift and loading matrices. No quoted equations or self-citation chains are supplied that reduce any central prediction (e.g., latent trajectories or reversion rates) to a fitted parameter by construction. Simulations are described as validation on data generated from the same model class, which is standard practice and does not constitute circularity under the rules; real-data applications (microbiome, SST) are treated as external checks. Because no load-bearing step is shown to collapse to self-definition, fitted-input renaming, or an unverified self-citation theorem, the derivation chain remains independent of its own outputs.
Axiom & Free-Parameter Ledger
Reference graph
Works this paper leans on
-
[1]
Skew Ornstein--Uhlenbeck processes and their financial applications , journal =
Suxin Wang and Shiyu Song and Yongjin Wang , keywords =. Skew Ornstein--Uhlenbeck processes and their financial applications , journal =. 2015 , issn =. doi:https://doi.org/10.1016/j.cam.2014.06.023 , url =
-
[2]
Available at pauillac.inria.fr/algo/csolve/ou.pdf , year=
Ornstein--Uhlenbeck process , author=. Available at pauillac.inria.fr/algo/csolve/ou.pdf , year=
-
[3]
Application of
Kenney, Toby and Gao, Junqiu and Gu, Hong , journal=. Application of. 2020 , publisher=
2020
-
[4]
Physica A: Statistical Mechanics and its Applications , volume=
Ornstein--Uhlenbeck process in a human body weight fluctuation , author=. Physica A: Statistical Mechanics and its Applications , volume=. 2021 , publisher=
2021
-
[5]
Insurance: Mathematics and Economics , volume=
Optimal proportional reinsurance and investment in a stock market with Ornstein--Uhlenbeck process , author=. Insurance: Mathematics and Economics , volume=. 2011 , publisher=
2011
-
[6]
Physica A: Statistical Mechanics and its Applications , volume=
A stochastic differential equation SIS epidemic model incorporating Ornstein--Uhlenbeck process , author=. Physica A: Statistical Mechanics and its Applications , volume=. 2018 , publisher=
2018
-
[7]
Stochastic Environmental Research and Risk Assessment , volume=
Linear Gaussian state-space model with irregular sampling: application to sea surface temperature , author=. Stochastic Environmental Research and Risk Assessment , volume=. 2011 , publisher=
2011
-
[8]
2012 , publisher=
Time series analysis by state space methods , author=. 2012 , publisher=
2012
-
[9]
Ecology , volume=
Density-dependent state-space model for population-abundance data with unequal time intervals , author=. Ecology , volume=. 2014 , publisher=
2014
-
[10]
Ecological Monographs , volume=
A guide to state--space modeling of ecological time series , author=. Ecological Monographs , volume=. 2021 , publisher=
2021
-
[11]
Genome biology , volume=
Moving pictures of the human microbiome , author=. Genome biology , volume=. 2011 , publisher=
2011
-
[12]
science , volume=
Bacterial community variation in human body habitats across space and time , author=. science , volume=. 2009 , publisher=
2009
-
[13]
Scientific reports , volume=
Impact of sequencing depth on the characterization of the microbiome and resistome , author=. Scientific reports , volume=. 2018 , publisher=
2018
-
[14]
Frontiers in microbiology , volume=
Analysis of microbiome data in the presence of excess zeros , author=. Frontiers in microbiology , volume=. 2017 , publisher=
2017
-
[15]
PeerJ , volume=
Desulfovibrio is not always associated with adverse health effects in the Guangdong Gut Microbiome Project , author=. PeerJ , volume=. 2021 , publisher=
2021
-
[16]
Journal of allergy and clinical immunology , volume=
Dietary fiber and SCFAs in the regulation of mucosal immunity , author=. Journal of allergy and clinical immunology , volume=. 2023 , publisher=
2023
-
[17]
Gut microbes , volume=
Gut microbes from the phylogenetically diverse genus Eubacterium and their various contributions to gut health , author=. Gut microbes , volume=. 2020 , publisher=
2020
-
[18]
Nutrients , volume=
Dietary short-term fiber interventions in arthritis patients increase systemic SCFA levels and regulate inflammation , author=. Nutrients , volume=. 2020 , publisher=
2020
-
[19]
Frontiers in microbiology , volume=
Intestinal short chain fatty acids and their link with diet and human health , author=. Frontiers in microbiology , volume=. 2016 , publisher=
2016
-
[20]
Gut , volume=
Gut microbiome stability and resilience: elucidating the response to perturbations in order to modulate gut health , author=. Gut , volume=. 2021 , publisher=
2021
-
[21]
Frontiers in Microbiology , volume=
Effect of fiber and fecal microbiota transplantation donor on recipient mice gut microbiota , author=. Frontiers in Microbiology , volume=. 2021 , publisher=
2021
-
[22]
Nature Cell and Science , volume=
Environmental Factors Affecting the Gut Microbiota and Their Consequences , author=. Nature Cell and Science , volume=. doi:10.61474/ncs.2024.00009. , year=
-
[23]
2009 , publisher=
Handbook of stochastic methods: for the natural and social sciences, volume Springer series in syneretics , author=. 2009 , publisher=
2009
-
[24]
Ecology , volume=
Continuous-time correlated random walk model for animal telemetry data , author=. Ecology , volume=. 2008 , publisher=
2008
-
[25]
Microbiome , volume=
Dynamic interaction network inference from longitudinal microbiome data , author=. Microbiome , volume=. 2019 , publisher=
2019
-
[26]
Frontiers in Genetics , volume=
ARZIMM: a novel analytic platform for the inference of microbial interactions and community stability from longitudinal microbiome study , author=. Frontiers in Genetics , volume=. 2022 , publisher=
2022
-
[27]
Journal of Theoretical Biology , volume=
A phylogenetic comparative method for studying multivariate adaptation , author=. Journal of Theoretical Biology , volume=. 2012 , publisher=
2012
-
[28]
arXiv preprint arXiv:1805.10050 , year=
Bayesian estimation for large scale multivariate Ornstein-Uhlenbeck model of brain connectivity , author=. arXiv preprint arXiv:1805.10050 , year=
-
[29]
doi:10.24381/cds.adbb2d47 , note =
ERA5: Fifth generation of ECMWF atmospheric reanalyses of the global climate , year =. doi:10.24381/cds.adbb2d47 , url =
-
[30]
Journal of Climate , volume=
Low-frequency SST and upper-ocean heat content variability in the North Atlantic , author=. Journal of Climate , volume=
-
[31]
Journal of Geophysical Research: Atmospheres , volume=
A review of North Atlantic modes of natural variability and their driving mechanisms , author=. Journal of Geophysical Research: Atmospheres , volume=. 2009 , publisher=
2009
-
[32]
Frontiers in Marine Science , volume=
Multi-scale variability features of global sea surface temperature over the past century , author=. Frontiers in Marine Science , volume=. 2023 , publisher=
2023
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.