Recognition: unknown
Semi-Markov Models with Particle-Based Bayesian Inference for Epidemics
Pith reviewed 2026-05-08 01:59 UTC · model grok-4.3
The pith
Epidemic models can represent sustained transmission waves by letting the infection rate switch between persistent regimes of random length via a semi-Markov process.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The paper establishes that a semi-Markov process can govern the evolution of the transmission rate through persistent regimes of random duration in a state-space epidemic model. This provides an interpretable representation of sustained transmission phases with a parsimonious parameterization. Particle-based Bayesian methods are extended to handle the semi-Markov setting, differential equation-driven latent dynamics, and observation models based on functionals of the latent process, allowing both batch and sequential Monte Carlo inference. Application to UK COVID-19 data demonstrates that combining cases and deaths improves inference precision and stability over using deaths alone.
What carries the argument
The semi-Markov process that governs switches between transmission-rate regimes of random duration, paired with particle filters that update the latent state in the presence of differential-equation dynamics.
If this is right
- The semi-Markov formulation yields an interpretable identification of sustained transmission phases during epidemics such as COVID-19.
- Particle-based inference remains feasible when latent dynamics follow differential equations and observations depend on functionals of the state.
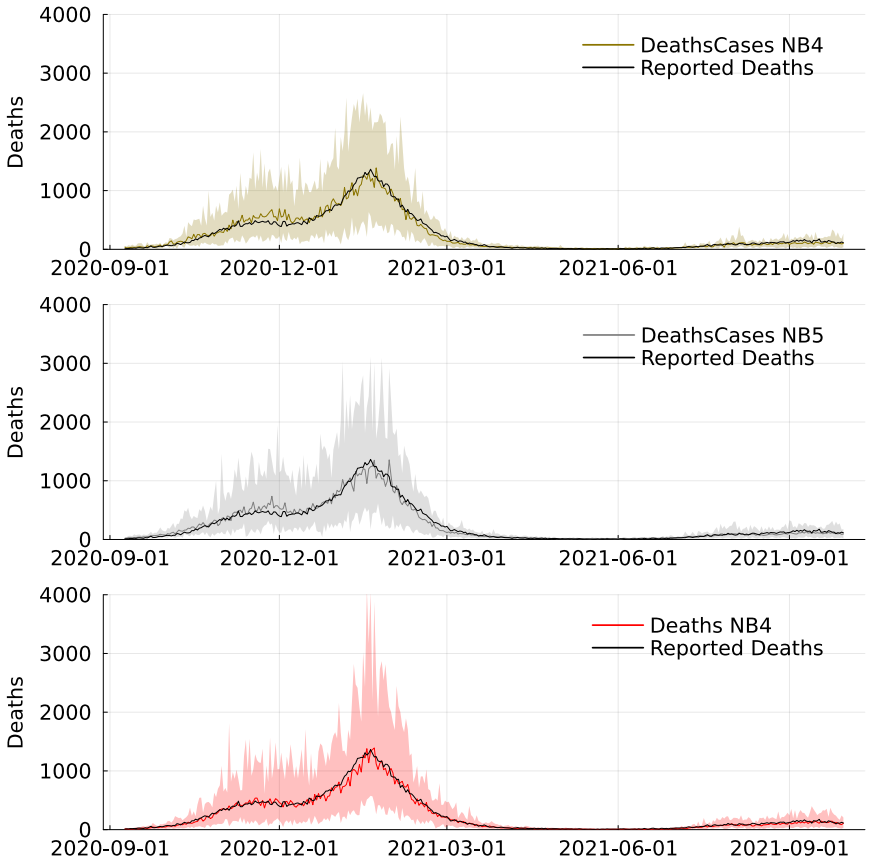
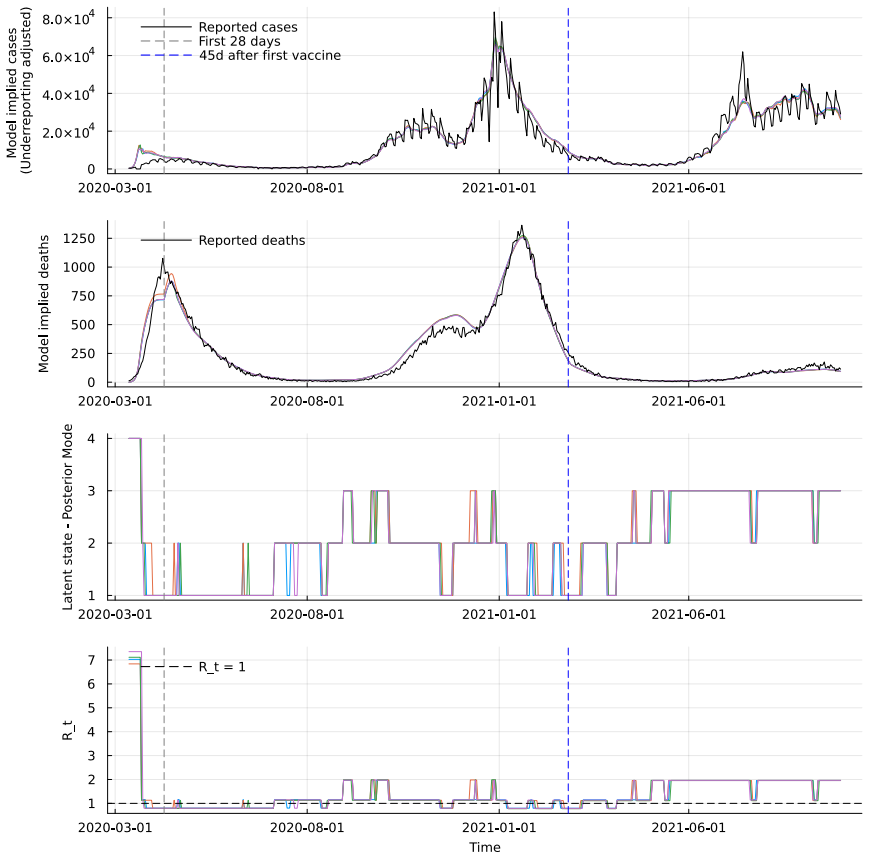
- Combining multiple data sources such as cases and deaths produces more precise and stable estimates of transmission regimes than single sources.
- The framework supports both batch analysis of complete datasets and sequential updating as new observations arrive.
Where Pith is reading between the lines
- If regime durations are well described by the semi-Markov process, the model could support short-term forecasts of how long the current transmission phase is likely to last.
- The same regime-switching structure might be applied to other systems that exhibit abrupt but sustained shifts, such as financial volatility regimes or population dynamics in ecology.
- Adding further data streams such as hospitalizations or mobility indicators could sharpen the detection of regime transitions without increasing model complexity.
Load-bearing premise
Epidemic transmission dynamics are adequately captured by discrete persistent regimes of random duration rather than continuous evolution or more intricate structures.
What would settle it
Observing epidemic time series in which transmission rate changes occur smoothly without identifiable persistent regimes, or finding that particle-based inference produces unstable or inconsistent regime estimates on such data, would falsify the central modeling claim.
Figures

read the original abstract
The COVID-19 pandemic has been characterised by multiple waves of transmission driven by interventions and emerging variants, challenging epidemic models that assume gradually evolving transmission dynamics. We propose a class of state-space models in which the transmission rate evolves through persistent regimes of random duration, governed by a semi-Markov process. This formulation yields an interpretable representation of sustained transmission phases and retains a parsimonious parameterisation. Particle-based Bayesian methods are well established for standard state-space models, but their use in semi-Markov settings has received comparatively limited attention. In epidemic applications, inference is further complicated by differential equation-driven latent dynamics and observation models defined through functionals of the latent process. We develop an inferential framework that accommodates these features, combining particle-based state updates with gradient-based parameter updates and enabling batch and sequential inference via particle and sequential Monte Carlo. We apply the proposed methodology to COVID-19 data from the United Kingdom and show that combining reported cases and deaths leads to more precise and stable inference compared to using deaths alone. These results illustrate the practical value of semi-Markov transmission models for epidemic analysis under complex observation schemes.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes a class of state-space models in which the transmission rate evolves through persistent regimes of random duration governed by a semi-Markov process. This yields an interpretable representation of sustained transmission phases while retaining parsimonious parameterization. The authors develop a particle-based Bayesian inferential framework that combines particle state updates with gradient-based parameter updates to handle semi-Markov dynamics, differential-equation latent processes, and observation functionals; both batch and sequential inference are enabled. The method is applied to UK COVID-19 data, where combining reported cases and deaths produces more precise and stable inference than deaths alone.
Significance. If the numerical reliability of the particle approximations holds, the semi-Markov formulation offers a useful middle ground between fully continuous transmission-rate models and discrete change-point models, directly addressing the sustained phases observed during the COVID-19 pandemic. The technical extension of particle methods to non-Markovian regime durations together with ODE-driven compartments is a non-trivial contribution. The empirical demonstration that joint case-death data improves posterior stability is practically relevant for epidemic surveillance, though its generalizability would be strengthened by additional validation.
major comments (2)
- [Results (UK application) and §3.2 (particle filter update)] The central claim that the particle-based framework reliably approximates the joint posterior over regime sequences, sojourn times, and continuous latent states is load-bearing, yet the manuscript provides no explicit analysis of effective sample size decay or weight variance induced by the auxiliary elapsed-time variable in the semi-Markov kernel. This variable modulates both the switching hazard and the numerical integration of the transmission-rate ODE; when observations are functionals of integrated compartments, the effective likelihood is only available after each integration step, amplifying degeneracy risk. No ESS plots or degeneracy diagnostics appear alongside the UK posterior results.
- [§3.1 (model specification) and numerical implementation details] The ODE integration step-size choice (Euler or Runge-Kutta) and its potential bias on the likelihood and posterior are not examined. Because the semi-Markov elapsed-time variable affects the instantaneous transmission rate at every integration step, even moderate step-size errors could systematically distort the regime-duration posterior; this is especially pertinent when the reported stability of the UK posteriors is used to support the method's practical value.
minor comments (2)
- [§2] The notation distinguishing the semi-Markov kernel from the embedded Markov chain could be made more explicit in the model section to aid readers unfamiliar with semi-Markov processes.
- [Results figures] Figure captions for the UK posterior plots should explicitly state the number of particles, the ESS threshold used for resampling, and whether the displayed trajectories are from a single run or aggregated.
Simulated Author's Rebuttal
We thank the referee for their positive summary of the paper's contributions and for the detailed major comments, which identify important areas for strengthening the presentation of numerical reliability. We respond to each comment below and will make the indicated revisions to the manuscript.
read point-by-point responses
-
Referee: [Results (UK application) and §3.2 (particle filter update)] The central claim that the particle-based framework reliably approximates the joint posterior over regime sequences, sojourn times, and continuous latent states is load-bearing, yet the manuscript provides no explicit analysis of effective sample size decay or weight variance induced by the auxiliary elapsed-time variable in the semi-Markov kernel. This variable modulates both the switching hazard and the numerical integration of the transmission-rate ODE; when observations are functionals of integrated compartments, the effective likelihood is only available after each integration step, amplifying degeneracy risk. No ESS plots or degeneracy diagnostics appear alongside the UK posterior results.
Authors: We agree that explicit reporting of effective sample size (ESS) trajectories and weight variance is necessary to substantiate the reliability of the particle approximations, especially with the auxiliary elapsed-time variable. Although these quantities were monitored during code development and showed no problematic degeneracy for the UK data, the manuscript did not include the corresponding plots or discussion. In the revised manuscript we will add ESS plots and degeneracy diagnostics alongside the UK posterior results, confirming that the semi-Markov kernel does not induce excessive weight variance or rapid ESS decay. This addition will directly support the central claim without altering any existing results. revision: yes
-
Referee: [§3.1 (model specification) and numerical implementation details] The ODE integration step-size choice (Euler or Runge-Kutta) and its potential bias on the likelihood and posterior are not examined. Because the semi-Markov elapsed-time variable affects the instantaneous transmission rate at every integration step, even moderate step-size errors could systematically distort the regime-duration posterior; this is especially pertinent when the reported stability of the UK posteriors is used to support the method's practical value.
Authors: We acknowledge that a dedicated examination of ODE integrator step-size bias was omitted and that the elapsed-time variable makes this check particularly relevant. The implementation uses a fixed-step fourth-order Runge-Kutta scheme with a step size of 0.1 days, chosen to be an order of magnitude smaller than the daily observation interval. In the revision we will explicitly state the integrator and step size, and we will add a brief sensitivity study showing that regime-duration posteriors for the UK data remain stable under moderate step-size perturbations (e.g., 0.05 and 0.2 days). This will address the concern about possible systematic distortion while preserving the reported stability conclusions. revision: yes
Circularity Check
No circularity: model proposal and inference framework are independently constructed and applied to external data.
full rationale
The paper defines a new semi-Markov state-space model for evolving transmission rates in persistent random-duration regimes, develops particle-based Bayesian inference (combining particle filters with gradient-based parameter updates) to handle the resulting latent ODE dynamics and observation functionals, and applies it to UK COVID-19 case and death data with a comparison to a deaths-only baseline. No derivation step reduces by construction to its inputs, no fitted quantities are relabeled as predictions, and no load-bearing self-citations or uniqueness theorems from the authors' prior work are invoked. The central claims rest on the explicit model specification and empirical performance on independent data.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Transmission rate evolves through persistent regimes of random duration governed by a semi-Markov process.
Reference graph
Works this paper leans on
-
[1]
Andrieu, A
C. Andrieu, A. Doucet, and R. Holenstein. Particle markov chain monte carlo methods. Journal of the Royal Statistical Society Series B, 72 0 (3): 0 269--342, 2010
2010
-
[2]
Barmpounakis and N
P. Barmpounakis and N. Demiris. Multiphasic stochastic epidemic models. Journal of the Royal Statistical Society Series C: Applied Statistics, 574 0 (2): 0 491–505, 2025
2025
-
[3]
Brazeau, R
N.F. Brazeau, R. Verity, S. Jenks, H. Fu, C. Whittaker, et al. Estimating the covid-19 infection fatality ratio accounting for seroreversion using statistical modelling. Communications Medicine, 54 0 (2), 2022
2022
-
[4]
Bulla and J
I. Bulla and J. Bulla. Stylized facts of financial time series and hidden semi-markov models. Computational Statistics & Data Analysis, 51: 0 2192--2209, 2006
2006
-
[5]
Chatzilena, N
A. Chatzilena, N. Demiris, and K. Kalogeropoulos. A modelling framework for the analysis of the transmission of sars-cov2. Statistics in Medicine, 43 0 (23): 0 4542--4558, 2024
2024
-
[6]
N. Chopin. A sequential particle filter method for static models. Biometrika, 89 0 (3): 0 539--552
-
[7]
Chopin and O
N. Chopin and O. Papaspiliopoulos. An Introduction to Sequential Monte Carlo. Springer International Publishing, 2020
2020
-
[8]
SMC 2 : an efficient algorithm for sequential analysis of state space models
N Chopin, P E Jacob, and O Papaspiliopoulos. SMC 2 : an efficient algorithm for sequential analysis of state space models . Journal of the Royal Statistical Society: Series B: Statistical Methodology, 75 0 (3): 0 397--426, 2012
2012
-
[9]
Corenflos, J
A. Corenflos, J. Thornton, A. Doucet, and G. Deligiannidis. Differentiable particle filtering via entropy-regularized optimal transport. In Proceedings of the 38th International Conference on Machine Learning, volume 139, 2021
2021
-
[10]
C. Dai, J. Heng, P. E. Jacob, and N. Whiteley. An invitation to sequential monte carlo samplers. Journal of the American Statistical Association, 117: 0 1587--1600, 2022
2022
-
[11]
Del Moral, A
P. Del Moral, A. Doucet, and A. Jasra. Sequential monte carlo samplers. Journal of the Royal Statistical Society: Series B: Statistical Methodology, 68 0 (3): 0 411--436, 2006
2006
-
[12]
Diekmann, H
O. Diekmann, H. Heesterbeek, and T. Britton. Mathematical Tools for Understanding Infectious Disease Dynamics . Princeton University Press, 11 2012
2012
-
[13]
Dureau, K
J. Dureau, K. Kalogeropoulos, and M. Baguelin. Capturing the time-varying drivers of an epidemic using stochastic dynamical systems. Biostatistics, 14 0 (3): 0 541--555, 2013
2013
-
[14]
Flaxman, S
S. Flaxman, S. Mishra, A. Gandy, H.J.T. Unwin, T.A. Mellan, et al. Estimating the effects of non-pharmaceutical interventions on covid-19 in europe. Nature, 584 0 (257-261), 2020
2020
-
[15]
On the marginal likelihood and cross-validation
E Fong and C C Holmes. On the marginal likelihood and cross-validation. Biometrika, 107 0 (2): 0 489--496, 06 2020
2020
-
[16]
Gelman, J
A. Gelman, J. Hwang, and A. Vehtari. Understanding predictive information criteria for bayesian models. Statistics and Computing, 24 0 (6): 0 1997--1016, 2014
1997
-
[17]
Gov.uk coronavirus (covid-19) in the uk, 2022
GOV.UK . Gov.uk coronavirus (covid-19) in the uk, 2022
2022
-
[18]
M. D. Hoffman and A. Gelman. The no-u-turn sampler: Adaptively setting path lengths in hamiltonian monte carlo. The Journal of Machine Learning Research, 15 0 (1): 0 1593–1623, 2014
2014
-
[19]
Iannucci, P
G. Iannucci, P. Barmpounakis, A. Beskos, and N. Demiris. On a reinforcement learning methodology for epidemic control, with application to covid-19, 2025
2025
-
[20]
G. Kastner. Dealing with stochastic volatility in time series using the r package stochvol. Journal of Statistical Software, Articles, 69 0 (5): 0 1--30, 2016
2016
-
[21]
Levin, W.P
A.T. Levin, W.P. Hanage, N. Owusu-Boaitey, K.B. Cochran, S.P. Walsh, et al. Assessing the age specificity of infection fatality rates for covid-19: systematic review, meta-analysis, and public policy implications, 2020
2020
-
[22]
Lindsten and T
F. Lindsten and T. Schön. Backward simulation methods for monte carlo statistical inference. Foundations and Trends in Machine Learning, 6: 0 1--142, 01 2013
2013
-
[23]
J. Liu, W. Xie, Y. Wang, Y. Xiong, S. Chen, et al. A comparative overview of covid-19, mers and sars: Review article. International journal of surgery, 81 0 (1-8), 2020
2020
-
[24]
R.M. Neal. Mcmc using hamiltonian dynamics. In Steve Brooks, Andrew Gelman, Galin L. Jones, and Xiao-Li Meng, editors, Handbook of Markov Chain Monte Carlo, pages 113--162. Chapman and Hall/CRC Press, 2010
2010
-
[25]
Covid-19, sars and mers: are they closely related? Clinical Microbiology and Infection, 26 0 (6), 2020
N Petrosillo, G Viceconte, O Ergonul, G Ippolito, and E Petersen. Covid-19, sars and mers: are they closely related? Clinical Microbiology and Infection, 26 0 (6), 2020
2020
-
[26]
Rackauckas and Q
C. Rackauckas and Q. Nie. Differentialequations.jl--a performant and feature-rich ecosystem for solving differential equations in julia. Journal of Open Research Software, 5 0 (1): 0 15, 2017
2017
-
[27]
Spiegelhalter, N.G
D.J. Spiegelhalter, N.G. Best, B.P. Carlin, and A. Van Der Linde. Bayesian measures of model complexity and fit. Journal of the Royal Statistical Society: Series B: Statistical Methodology, 64 0 (4): 0 583--639, 2002
2002
-
[28]
Vehtari, A
A. Vehtari, A. Gelman, and A. Gabry. Practical bayesian model evaluation using leave-one-out cross-validation and WAIC . Statistics and Computing, 27 0 (5): 0 1413--1432, 2016
2016
-
[29]
Verity, L.C
R. Verity, L.C. Okell, I. Dorigatti, P. Winskill, C. Whittaker, et al. Estimates of the severity of coronavirus disease 2019: a model-based analysis. The Lancet infectious diseases, 20 0 (6): 0 669--677, 2020
2019
-
[30]
Watanabe
S. Watanabe. Asymptotic equivalence of bayes cross validation and widely applicable information criterion in singular learning theory. Journal of Machine Learning Research, 11: 0 3571--3594, 2010
2010
-
[31]
Appropriate models for the management of infectious diseases, 2005
HJ Wearing, P Rohani, and MJ Keeling. Appropriate models for the management of infectious diseases, 2005
2005
-
[32]
X. Xu, T. Kypraios, and P. O'Neill. Bayesian non-parametric inference for stochastic epidemic models using Gaussian Processes . Biostatistics, 17 0 (4): 0 619--633, 2018
2018
-
[33]
S. Yu. Hidden semi-markov models. Artificial Intelligence, 174 0 (2): 0 215--243, 2010
2010
-
[34]
S. Yu. Hidden Semi-Markov Models: Theory, Algorithms and Applications. Elsevier, 2016
2016
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.