Recognition: unknown
Inferring Active Neural Circuits Using Diffusion Scores
Pith reviewed 2026-05-08 01:44 UTC · model grok-4.3
The pith
Denoising score models on consecutive neural snapshots yield products that recover the Jacobian of state transitions, identifying lag-specific directed circuits.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
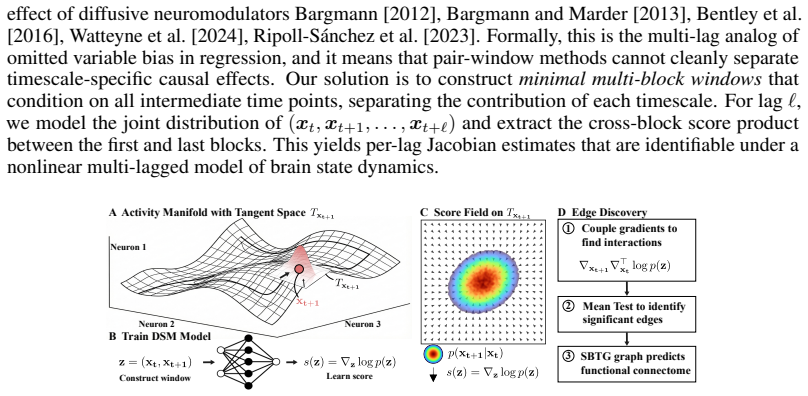
The key discovery is that products of scores estimated across blocks from joint-window denoising models recover the Jacobian of the nonlinear transition map between brain states. By using minimal multi-block windows that condition on intermediate time points, the approach separates lag-specific effects and identifies directed interactions in neuronal population data.
What carries the argument
Cross-block score products computed from denoising score models trained on multi-block activity windows; these products act as tests for directed edges by recovering the Jacobian of the state transition.
Load-bearing premise
The denoising score models trained on the joint windows of activity snapshots must accurately capture the score of the underlying data distribution.
What would settle it
Generate synthetic neural activity data from a known nonlinear dynamical system whose transition Jacobian can be computed exactly, apply the score product method, and verify if the estimated values match the true Jacobian entries up to sampling noise.
Figures


read the original abstract
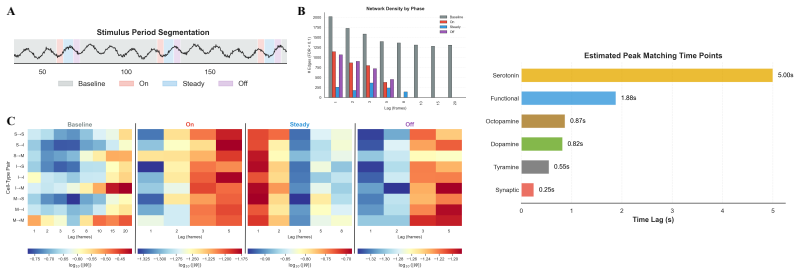
In biological systems, neural circuits compute through directed, short-latency interactions whose effects unfold across multiple time scales and behavioral contexts. We address the problem of inferring these local, lag-specific interactions from sampled neural population activity under varying stimuli, without assuming a parametric form for the underlying dynamics. Our approach leverages denoising score models by estimating joint-window scores over consecutive activity snapshots (i.e., brain states) and converting these scores into calibrated, directed edge tests via cross-block score products. The key insight is that these products recover the Jacobian of the transition map between brain states under nonlinear dynamics. To cleanly separate lag-specific effects, we introduce minimal multi-block windows that condition on intermediate time points, avoiding the omitted-lag bias inherent in pairwise analyses. The resulting method, Score--Block Time Graphs (SBTG), identifies lag-specific directed interactions in sampled neuronal population data. We specifically apply SBTG to whole-brain C. elegans calcium imaging data to recover lag-specific circuit structure not resolved by current methods, including improved alignment with independent connectomes, cell-type-specific temporal organization, and neuromodulatory profiles consistent with known receptor kinetics. These findings highlight the potential for SBTG to serve as a practical ``AI for science'' tool by turning high-dimensional neural population recordings into statistically testable circuit hypotheses.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces Score-Block Time Graphs (SBTG), a method that trains denoising score models on multi-block joint windows of neural population activity snapshots and converts cross-block score products into directed, lag-specific interaction tests. The central claim is that these products recover the Jacobian of the (nonlinear) transition map between brain states without parametric assumptions on the dynamics. Multi-block conditioning is introduced to mitigate omitted-lag bias. The method is applied to whole-brain C. elegans calcium imaging data, with reported improvements in alignment to independent connectomes, cell-type temporal organization, and consistency with neuromodulatory receptor kinetics.
Significance. If the Jacobian-recovery claim is rigorously established, SBTG would constitute a non-parametric advance for extracting directed, lag-resolved circuit hypotheses from high-dimensional neural recordings. The C. elegans application illustrates potential utility by producing biologically plausible structure not resolved by prior methods. The absence of free parameters in the core construction and the use of existing score-matching theory are strengths that would support broader adoption if the derivation holds.
major comments (1)
- [Abstract and §3] Abstract and §3 (Method): the assertion that 'cross-block score products recover the Jacobian of the transition map between brain states under nonlinear dynamics' is load-bearing yet unsupported by any derivation. For a general map x_{t+τ}=f(x_t)+η with nonlinear f and arbitrary noise η, the joint score ∇_{x,y} log p(x,y) equals the conditional score only after marginal subtraction and involves the density ratio p(y|x)/p(x); a simple product of score blocks does not isolate df/dx. The multi-block conditioning removes omitted-lag bias but does not alter the algebraic step required inside each window. A complete proof or explicit counter-example analysis under the paper's noise model is required.
minor comments (3)
- [§2] §2 (Background): the notation for 'joint-window scores' and 'cross-block products' should be introduced with an explicit equation before the claim is stated.
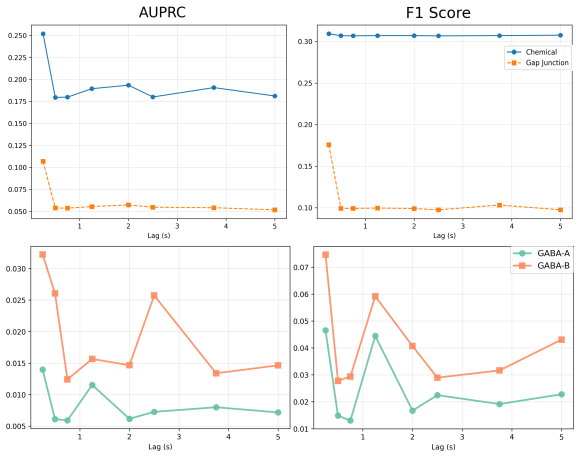
- [Figure 3] Figure 3 (C. elegans results): quantitative metrics (e.g., precision-recall against connectome or edge-overlap statistics) comparing SBTG to baselines are needed to support the claim of 'improved alignment'; visual inspection alone is insufficient.
- [§3.1] The manuscript would benefit from an explicit statement of the noise model assumed when training the score networks (e.g., whether the diffusion process matches the biological noise statistics).
Simulated Author's Rebuttal
We thank the referee for their careful reading and constructive critique. The major comment correctly identifies that the central theoretical claim requires a more explicit derivation, which we address below by committing to a targeted revision.
read point-by-point responses
-
Referee: [Abstract and §3] Abstract and §3 (Method): the assertion that 'cross-block score products recover the Jacobian of the transition map between brain states under nonlinear dynamics' is load-bearing yet unsupported by any derivation. For a general map x_{t+τ}=f(x_t)+η with nonlinear f and arbitrary noise η, the joint score ∇_{x,y} log p(x,y) equals the conditional score only after marginal subtraction and involves the density ratio p(y|x)/p(x); a simple product of score blocks does not isolate df/dx. The multi-block conditioning removes omitted-lag bias but does not alter the algebraic step required inside each window. A complete proof or explicit counter-example analysis under the paper's noise model is required.
Authors: We agree that the manuscript would be strengthened by an explicit derivation. The current §3 motivates the cross-block product via score-matching properties but does not spell out the algebraic cancellation of the density-ratio and marginal terms under the assumed additive noise model. In the revision we will insert a dedicated subsection deriving the relation step-by-step: starting from the joint score of the multi-block window, applying the chain rule for the conditional density, and showing how the product isolates the Jacobian of the nonlinear transition map after the multi-block conditioning subtracts the omitted-lag contributions. We will also add a short synthetic-data experiment with a known nonlinear f to illustrate recovery and bound the approximation error. This directly addresses the referee's concern about the missing algebraic step. revision: yes
Circularity Check
No significant circularity in derivation chain
full rationale
The paper grounds its method in established denoising score-matching theory to estimate joint-window scores from data snapshots, then applies cross-block products as a derived operation to recover the Jacobian of the transition map. This step is presented as a mathematical consequence of score properties rather than a self-definition, fitted parameter renamed as prediction, or reduction to the target result by construction. Multi-block conditioning is introduced explicitly to mitigate omitted-lag bias and is not tautological with the Jacobian claim. No load-bearing self-citation chain or ansatz smuggling is required for the core conversion; the approach remains self-contained against external score-matching benchmarks.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Denoising score models trained on joint activity windows estimate the true score of the data distribution
- ad hoc to paper Cross-block score products recover the Jacobian of the transition map under nonlinear dynamics
Reference graph
Works this paper leans on
-
[1]
Journal of Machine Learning Research , volume =
Estimation of Non-Normalized Statistical Models by Score Matching , author =. Journal of Machine Learning Research , volume =
-
[2]
Neural Computation , volume =
A Connection Between Score Matching and Denoising Autoencoders , author =. Neural Computation , volume =. 2011 , doi =
2011
-
[3]
Generative Modeling by Estimating Gradients of the Data Distribution , author =. Advances in Neural Information Processing Systems , year =. 1907.05600 , archivePrefix =
-
[4]
Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & data mining , pages=
Optuna: A next-generation hyperparameter optimization framework , author=. Proceedings of the 25th ACM SIGKDD international conference on knowledge discovery & data mining , pages=
-
[5]
Score-Based Generative Modeling through Stochastic Differential Equations
Score-Based Generative Modeling through Stochastic Differential Equations , author =. International Conference on Learning Representations , year =. 2011.13456 , archivePrefix =
work page internal anchor Pith review arXiv 2011
-
[6]
Nature , volume =
Survey of Spiking in the Mouse Visual System Reveals Functional Hierarchy , author =. Nature , volume =. 2021 , doi =
2021
-
[7]
Neuron , volume =
Cortical Areas Interact through a Communication Subspace , author =. Neuron , volume =. 2019 , doi =
2019
-
[8]
Nature , volume =
Network Anatomy and In Vivo Physiology of Visual Cortical Neurons , author =. Nature , volume =. 2011 , doi =
2011
-
[9]
Journal of Computational Neuroscience , volume =
Transfer Entropy---A Model-Free Measure of Effective Connectivity for the Neurosciences , author =. Journal of Computational Neuroscience , volume =. 2011 , doi =
2011
-
[10]
Econometrica , volume =
Investigating Causal Relations by Econometric Models and Cross-Spectral Methods , author =. Econometrica , volume =. 1969 , doi =
1969
-
[11]
2005 , isbn =
New Introduction to Multiple Time Series Analysis , author =. 2005 , isbn =
2005
-
[12]
Biometrika , volume =
Spectra of Some Self-Exciting and Mutually Exciting Point Processes , author =. Biometrika , volume =. 1971 , doi =
1971
-
[13]
Discovering Contemporaneous and Lagged Causal Relations in Autocorrelated Nonlinear Time Series Datasets , author =. arXiv preprint , year =. 2003.03685 , archivePrefix =
-
[14]
DAGs with NO TEARS: Continuous Optimization for Structure Learning , author =. Advances in Neural Information Processing Systems , year =. 1803.01422 , archivePrefix =
-
[15]
2019 , editor =
Yu, Yue and Chen, Jie and Gao, Tian and Yu, Mo , booktitle =. 2019 , editor =
2019
-
[16]
Ozcelik, Furkan and VanRullen, Rufin , title =. Scientific Reports , year =. doi:10.1038/s41598-023-42891-8 , url =
-
[17]
2020 , eprint=
DYNOTEARS: Structure Learning from Time-Series Data , author=. 2020 , eprint=
2020
-
[18]
2024 , eprint=
Latent Diffusion for Neural Spiking Data , author=. 2024 , eprint=
2024
-
[19]
2023 , eprint=
Scalable Causal Discovery with Score Matching , author=. 2023 , eprint=
2023
-
[20]
2023 , eprint=
Sample Complexity Bounds for Score-Matching: Causal Discovery and Generative Modeling , author=. 2023 , eprint=
2023
-
[21]
Journal of the Royal Statistical Society: Series B , volume =
Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing , author =. Journal of the Royal Statistical Society: Series B , volume =
-
[22]
The Annals of Statistics , volume =
The Control of the False Discovery Rate in Multiple Testing under Dependency , author =. The Annals of Statistics , volume =
-
[23]
2023 , eprint=
Score-based Causal Representation Learning with Interventions , author=. 2023 , eprint=
2023
-
[24]
Journal of Multivariate Analysis , volume =
A Well-Conditioned Estimator for Large-Dimensional Covariance Matrices , author =. Journal of Multivariate Analysis , volume =. 2004 , doi =
2004
-
[25]
High Dimensional Probability V: The Luminy Volume , pages =
Bernstein Inequality and Moderate Deviations under Strong Mixing Conditions , author =. High Dimensional Probability V: The Luminy Volume , pages =. 2009 , publisher =
2009
-
[26]
Bioessays , volume=
Beyond the connectome: how neuromodulators shape neural circuits , author=. Bioessays , volume=. 2012 , publisher=
2012
-
[27]
Nature methods , volume=
From the connectome to brain function , author=. Nature methods , volume=. 2013 , publisher=
2013
-
[28]
elegans wired and wireless connectome: insights into principles of nervous system structure and function , author=
C. elegans wired and wireless connectome: insights into principles of nervous system structure and function , author=. Journal of Biosciences , volume=. 2025 , publisher=
2025
-
[29]
Current Opinion in Neurobiology , volume=
Dynamic functional connectivity in the static connectome of Caenorhabditis elegans , author=. Current Opinion in Neurobiology , volume=. 2022 , publisher=
2022
-
[30]
Genetics , volume=
Neuropeptide signaling network of Caenorhabditis elegans: from structure to behavior , author=. Genetics , volume=. 2024 , publisher=
2024
-
[31]
elegans , author=
The neuropeptidergic connectome of C. elegans , author=. Neuron , volume=. 2023 , publisher=
2023
-
[32]
elegans mating , author=
Natural sensory context drives diverse brain-wide activity during C. elegans mating , author=. Cell , volume=. 2021 , publisher=
2021
-
[33]
elegans , author=
A set of hub neurons and non-local connectivity features support global brain dynamics in C. elegans , author=. Current Biology , volume=. 2022 , publisher=
2022
-
[34]
elegans serotonergic system at whole-brain scale , author=
Dissecting the functional organization of the C. elegans serotonergic system at whole-brain scale , author=. Cell , volume=. 2023 , publisher=
2023
-
[35]
Cell , year=
Infrequent strong connections constrain connectomic predictions of neuronal function , author=. Cell , year=
-
[36]
Science , volume=
The neurobiology of slow synaptic transmission , author=. Science , volume=. 2001 , publisher=
2001
-
[37]
elegans connectome and signaling network , author=
Diverging network architecture of the C. elegans connectome and signaling network , author=. PRX Life , volume=. 2025 , publisher=
2025
-
[38]
elegans , author=
Bridging the gap between the connectome and whole-brain activity in C. elegans , author=. bioRxiv , pages=
-
[39]
The structure of the nervous system of the nematode Caenorhabditis elegans: the mind of a worm , author=. Phil. Trans. R. Soc. Lond , volume=
-
[40]
Nature , volume=
Connectomes across development reveal principles of brain maturation , author=. Nature , volume=. 2021 , publisher=
2021
-
[41]
elegans males and hermaphrodites , author=
A neurotransmitter atlas of C. elegans males and hermaphrodites , author=. Elife , volume=. 2024 , publisher=
2024
-
[42]
WormBook: The online review of C
The neuronal genome of Caenorhabditis elegans , author=. WormBook: The online review of C. elegans biology [Internet] , year=
-
[43]
Nature , volume=
Ultrasensitive fluorescent proteins for imaging neuronal activity , author=. Nature , volume=. 2013 , publisher=
2013
-
[44]
elegans , author=
Biogenic amine neurotransmitters in C. elegans , author=. WormBook: The Online Review of C. elegans Biology [Internet] , year=
-
[45]
Neuron , volume=
A tyramine-gated chloride channel coordinates distinct motor programs of a Caenorhabditis elegans escape response , author=. Neuron , volume=. 2009 , publisher=
2009
-
[46]
Genetics , volume=
Chemosensory signal transduction in Caenorhabditis elegans , author=. Genetics , volume=. 2021 , publisher=
2021
-
[47]
elegans chemosensory neurons are preserved in behavioral dynamics , author=
Temporal responses of C. elegans chemosensory neurons are preserved in behavioral dynamics , author=. Neuron , volume=. 2014 , publisher=
2014
-
[48]
PLoS computational biology , volume=
Structural properties of the Caenorhabditis elegans neuronal network , author=. PLoS computational biology , volume=. 2011 , publisher=
2011
-
[49]
elegans , author=
Dopamine counteracts octopamine signalling in a neural circuit mediating food response in C. elegans , author=. The EMBO journal , volume=. 2009 , publisher=
2009
-
[50]
Journal of Experimental Zoology , volume=
Effects of starvation and neuroactive drugs on feeding in Caenorhabditis elegans , author=. Journal of Experimental Zoology , volume=. 1990 , publisher=
1990
-
[51]
and Jarrell, Travis A
Cook, Steven J. and Jarrell, Travis A. and Brittin, Christopher A. and Wang, Yi and Bloniarz, Adam E. and Yakovlev, Maksim A. and Nguyen, Ken C. Q. and Tang, Leo T.-H. and Bayer, Emily A. and Duerr, Janet S. and Bülow, H. Robert and Hobert, Oliver and Hall, David H. and Emmons, Scott W. , journal =. Whole-animal connectomes of both. 2019 , doi =
2019
-
[52]
PLoS computational biology , volume=
The multilayer connectome of Caenorhabditis elegans , author=. PLoS computational biology , volume=. 2016 , publisher=
2016
-
[53]
, journal =
Randi, Francesco and Sharma, Anuj and Dvali, Saba and Yuan, Jiarui and Yemini, Eviatar and Shen, Yao and Paninski, Liam and Leifer, Andrew M. , journal =. A neural circuit atlas of sensory integration in. 2023 , doi =
2023
-
[54]
Cell , volume =
Yemini, Eviatar and Lin, Albert and Nejatbakhsh, Alireza and Varol, Erdem and Sun, Ruoxi and M. Cell , volume =. 2021 , doi =
2021
-
[55]
Muralidhara, Inchara and Hardege, Iris , year = 2025, month = oct, journal =. The Dopaminergic System of. doi:10.1098/rsos.250843 , abstract =. https://royalsocietypublishing.org/rsos/article-pdf/doi/10.1098/rsos.250843/2831125/rsos.250843.pdf , pages =
-
[56]
Proceedings of the 35th International Conference on Machine Learning , year =
Neural Relational Inference for Interacting Systems , author =. Proceedings of the 35th International Conference on Machine Learning , year =
-
[57]
and Ostojic, Srdjan , booktitle =
Valente, Adrian and Pillow, Jonathan W. and Ostojic, Srdjan , booktitle =. Extracting computational mechanisms from neural data using low-rank
-
[58]
Lu, Ziyu and Zhang, Wuwei and Le, Trung and Wang, Hao and Sümbül, Uygar and SheaBrown, Eric and Mi, Lu , booktitle =
-
[59]
Econometrica , volume =
A Simple, Positive Semi-Definite, Heteroskedasticity and Autocorrelation Consistent Covariance Matrix , author =. Econometrica , volume =
-
[60]
Econometrica , volume =
Heteroskedasticity and Autocorrelation Consistent Covariance Matrix Estimation , author =. Econometrica , volume =
-
[61]
The Review of Economic Studies , volume =
Automatic Lag Selection in Covariance Matrix Estimation , author =. The Review of Economic Studies , volume =
-
[62]
Estimation of a Structural Vector Autoregression Model Using Non-Gaussianity , journal =
Aapo Hyv. Estimation of a Structural Vector Autoregression Model Using Non-Gaussianity , journal =. 2010 , volume =
2010
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.