Recognition: unknown
Dante: An Open Source Model Pre-Training and Fine-Tuning Tool for the Dafne Federated Framework for Medical Image Segmentation
Pith reviewed 2026-05-07 12:54 UTC · model grok-4.3
The pith
Dante provides an open-source fine-tuning backend that adapts pre-trained medical segmentation models to new MRI domains using gradual unfreezing and LoRA.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Dante serves as the training and fine-tuning backend for the Dafne federated segmentation framework, enabling training from scratch via automatic architecture setup along with two adaptation strategies: gradual unfreezing of layers and Low-Rank Adaptation extended to N-dimensional convolutions by channel-wise factorization. Across cross-domain MRI scenarios covering abdominal organs and brain white matter lesions in full-data and few-shot regimes, these approaches consistently yield faster convergence and higher final accuracy than standard training from scratch, with the magnitude of improvement scaling according to the richness of the initial pre-training data.
What carries the argument
Dante's LoRA extension to N-dimensional convolutional layers via channel-wise factorization together with gradual unfreezing schedules, which allow selective parameter updates while retaining prior representations during domain transfer.
If this is right
- Gradual unfreezing shortens the number of epochs needed to reach high segmentation performance when moving to a new domain.
- LoRA delivers strong Dice coefficients while updating only a small subset of parameters, even in few-shot settings.
- Larger performance gains occur when the source pre-training dataset is more extensive or diverse.
- The same adaptation methods succeed across different anatomical targets such as organs and lesions.
- Dante functions as a reusable, domain-agnostic component inside the Dafne federated framework.
Where Pith is reading between the lines
- New clinical sites could adapt existing models with substantially less local annotation effort.
- The federated integration may allow collaborative adaptation across institutions while keeping raw patient data private.
- The channel-wise LoRA factorization could be applied to other volumetric modalities such as CT or PET.
- Faster domain adaptation may accelerate practical deployment of segmentation tools in varied hospital environments.
Load-bearing premise
The specific cross-domain MRI transfer scenarios for abdominal organs and brain lesions, plus the full-data versus few-shot conditions tested, are representative of real clinical deployment and that the observed performance gains generalize beyond the datasets and architectures used.
What would settle it
A new experiment at an independent clinical site with a different MRI scanner and patient population in which neither gradual unfreezing nor LoRA matches or exceeds the Dice scores and convergence speed of standard fine-tuning on the same segmentation tasks.
Figures
read the original abstract
Adapting pre-trained deep learning segmentation models to new clinical domains is a persistent challenge in medical image analysis, particularly when annotated data at the target site are scarce. Parameter-efficient fine-tuning strategies offer a principled solution by selectively updating a controlled subset of model parameters, preserving previously acquired representations while reducing the risk of overfitting on small datasets. This paper introduces DAfNe TrainEr (Dante), an open-source module integrating with the Dafne federated segmentation ecosystem as a dedicated training and fine-tuning backend. Dante supports training from scratch with automatic architecture configuration, configurable layer freezing schedules, and Low-Rank Adaptation (LoRA) extended to N-dimensional convolutional layers through channel-wise factorization. To validate the module, Gradual Unfreezing (GU) and LoRA are assessed across realistic cross-domain MRI transfer scenarios covering abdominal organ segmentation and brain white matter lesion segmentation, under full-data and few-shot conditions. GU reduced the epochs required to reach 85% of peak performance by up to 63.6% compared to training from scratch, while LoRA achieved Dice Similarity Coefficients up to 0.957 in data-rich scenarios. Both strategies outperformed the baseline across all tested domains, with gains amplified by richer pre-training datasets. These results validate Dante as a domain-agnostic fine-tuning module for medical image segmentation in real clinical deployment conditions. Dante code is available at https://github.com/dafne-imaging/dafne-torch-trainer while Dafne ecosystem project is available at https://github.com/dafne-imaging.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces Dante, an open-source training and fine-tuning backend integrated with the Dafne federated medical image segmentation framework. It supports training from scratch with automatic architecture configuration, configurable layer freezing, and an extension of Low-Rank Adaptation (LoRA) to N-dimensional convolutional layers via channel-wise factorization. Validation experiments assess Gradual Unfreezing (GU) and LoRA on cross-domain MRI transfer tasks for abdominal organ segmentation and brain white matter lesion segmentation, under both full-data and few-shot regimes, reporting up to 63.6% fewer epochs to reach 85% of peak performance with GU and Dice scores up to 0.957 with LoRA.
Significance. If the empirical claims hold under broader testing, Dante would provide a practical, open-source module for parameter-efficient adaptation of segmentation models in data-scarce clinical settings. The explicit release of code at the cited GitHub repositories is a clear strength that supports reproducibility and community adoption within the Dafne ecosystem.
major comments (1)
- [Abstract] Abstract: The final sentence asserts that 'These results validate Dante as a domain-agnostic fine-tuning module for medical image segmentation in real clinical deployment conditions.' This claim is not supported by the reported experiments, which are confined to two MRI-specific tasks (abdominal organs and brain white-matter lesions) using controlled cross-domain splits on fixed architectures; no results are shown for CT/PET modalities, scanner/protocol variation, or the federated multi-site training and privacy constraints that define the Dafne framework.
minor comments (2)
- [Abstract] The abstract and results sections do not report dataset sizes, exact baseline implementations, or statistical significance tests for the reported Dice and epoch reductions, making it difficult to assess the magnitude and reliability of the gains.
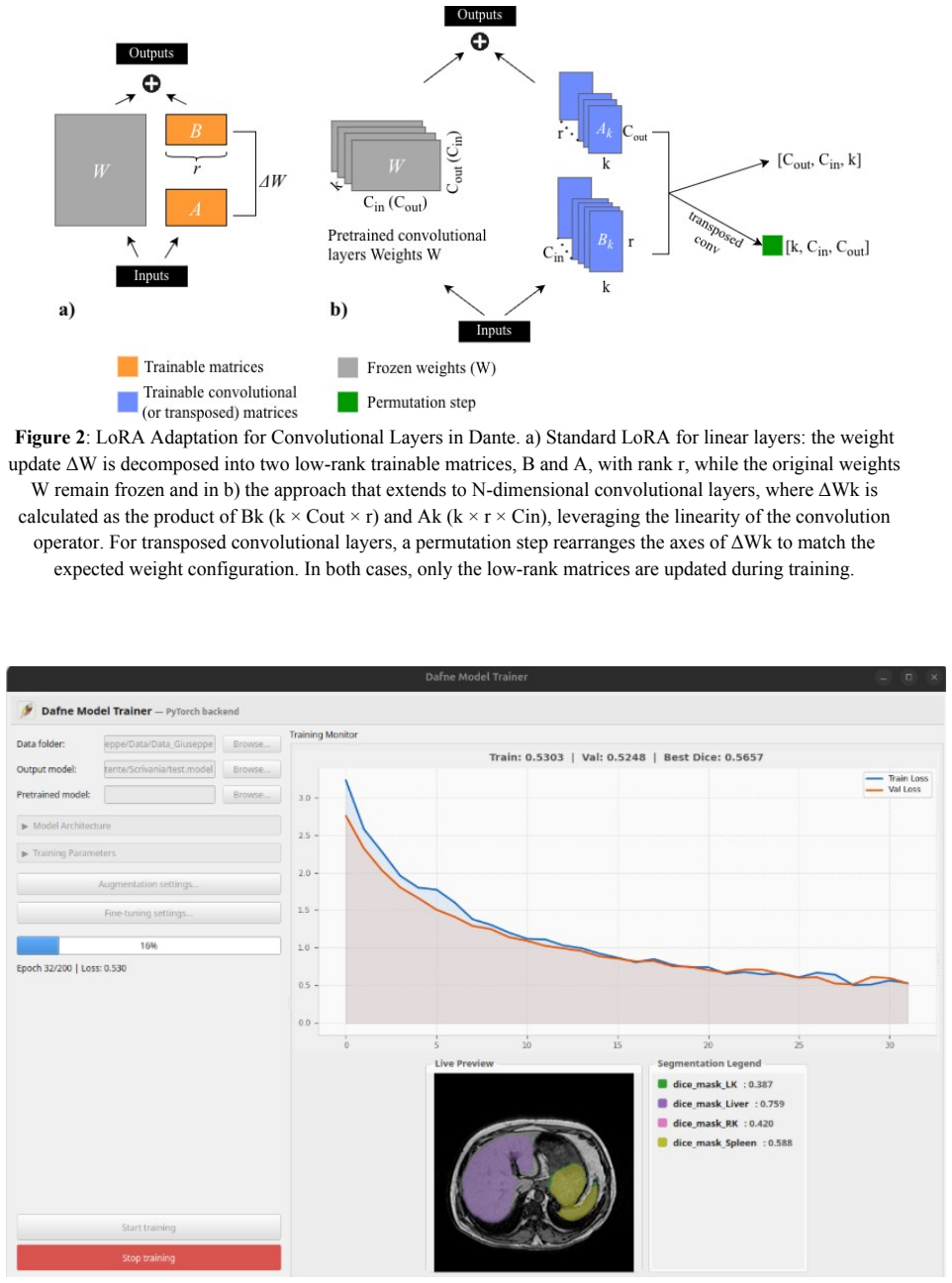
- Notation for the LoRA extension to N-D convolutions (channel-wise factorization) is introduced without an explicit equation or pseudocode in the methods description, which would aid reproducibility.
Simulated Author's Rebuttal
We thank the referee for their constructive feedback. We agree that the abstract's concluding claim overreaches the scope of the presented experiments and will revise it accordingly.
read point-by-point responses
-
Referee: [Abstract] Abstract: The final sentence asserts that 'These results validate Dante as a domain-agnostic fine-tuning module for medical image segmentation in real clinical deployment conditions.' This claim is not supported by the reported experiments, which are confined to two MRI-specific tasks (abdominal organs and brain white-matter lesions) using controlled cross-domain splits on fixed architectures; no results are shown for CT/PET modalities, scanner/protocol variation, or the federated multi-site training and privacy constraints that define the Dafne framework.
Authors: We acknowledge that this sentence overstates the generalizability of the results. The validation is limited to MRI cross-domain transfer tasks on two segmentation problems and does not include other modalities, scanner variations, or explicit federated multi-site experiments. We will revise the abstract to state that the results demonstrate the effectiveness of Dante for parameter-efficient adaptation in cross-domain MRI segmentation, while noting that broader validation across modalities and federated settings remains future work. The revised sentence will read: 'These results support the use of Dante for efficient fine-tuning in cross-domain MRI medical image segmentation tasks.' This change will be made in the resubmitted version. revision: yes
Circularity Check
No circularity: empirical validation relies on new experiments
full rationale
The paper describes the Dante module and its features (GU, LoRA for N-D conv layers) then reports direct empirical results on cross-domain MRI transfers for abdominal organs and brain lesions under full-data and few-shot regimes. Performance metrics (epoch reduction up to 63.6%, Dice up to 0.957) are obtained from fresh training runs and comparisons against baselines on held-out splits; no equations, fitted parameters renamed as predictions, self-definitional constructs, or load-bearing self-citations appear in the derivation. The validation chain is self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
Reference graph
Works this paper leans on
-
[1]
Moglia A, Leccardi M, Cavicchioli M, Maccarini A, Marcon M, Mainardi L, et al. Generalist models in medical image segmentation: A survey and performance comparison with task-specific approaches. Information Fusion. 2026 Mar;127:103709. doi:10.1016/j.inffus.2025.103709
-
[2]
Noh S, Lee BD. A narrative review of foundation models for medical image segmentation: zero-shot performance evaluation on diverse modalities. Quant Imaging Med Surg. 2025 Jun;15(6):5825–58. doi:10.21037/qims-2024-2826
-
[3]
Deep Learning-Based 3D and 2D Approaches for Skeletal Muscle Segmentation on Low-Dose CT Images
Timpano G, Veltri P, Vizza P, Cascini GL, Manti F. Deep Learning-Based 3D and 2D Approaches for Skeletal Muscle Segmentation on Low-Dose CT Images. J Digit Imaging Inform med. 2025 Aug 27. doi:10.1007/s10278-025-01646-9
-
[4]
Ma J, He Y, Li F, Han L, You C, Wang B. Segment anything in medical images. Nat Commun. 2024 Jan 22;15(1):654. doi:10.1038/s41467-024-44824-z
-
[5]
TotalSegmentator: Robust Segmentation of 104 Anatomic Structures in CT Images
Wasserthal J, Breit HC, Meyer MT, Pradella M, Hinck D, Sauter AW, et al. TotalSegmentator: Robust Segmentation of 104 Anatomic Structures in CT Images. Radiology: Artificial Intelligence. 2023 Sep 1;5(5):e230024. doi:10.1148/ryai.230024
-
[6]
Domain Adaptation for Medical Image Analysis: A Survey
Guan H, Liu M. Domain Adaptation for Medical Image Analysis: A Survey. IEEE Trans Biomed Eng. 2022 Mar;69(3):1173–85. doi:10.1109/TBME.2021.3117407
-
[7]
Karimi D, Warfield SK, Gholipour A. Transfer learning in medical image segmentation: New insights from analysis of the dynamics of model parameters and learned representations. Artificial Intelligence in Medicine. 2021 Jun;116:102078. doi:10.1016/j.artmed.2021.102078
-
[8]
Santini F, Wasserthal J, Agosti A, Deligianni X, Keene KR, Kan HE, et al. Deep Anatomical Federated Network (Dafne): An Open Client-Server Framework for Continuous, Collaborative Improvement of Deep Learning–based Medical Image Segmentation. Radiology: Artificial Intelligence. 2025 May 1;7(3):e240097. doi:10.1148/ryai.240097
-
[9]
LoRA: Low-Rank Adaptation of Large Language Models
Hu EJ, Shen Y, Wallis P, Allen-Zhu Z, Li Y, Wang S, et al. LoRA: Low-Rank Adaptation of Large Language Models [Internet]. arXiv; 2021 [cited 2026 Mar 10]. Available from: http://arxiv.org/abs/2106.09685 doi:10.48550/arXiv.2106.09685
work page internal anchor Pith review doi:10.48550/arxiv.2106.09685 2021
-
[10]
arXiv preprint arXiv:2304.13785 (2023)
Zhang K, Liu D. Customized Segment Anything Model for Medical Image Segmentation [Internet]. arXiv; 2023 [cited 2026 Mar 10]. Available from: https://arxiv.org/abs/2304.13785 doi:10.48550/ARXIV.2304.13785
-
[11]
Zhong Z, Tang Z, He T, Fang H, Yuan C. Convolution Meets LoRA: Parameter Efficient Finetuning for Segment Anything Model [Internet]. arXiv; 2024 [cited 2026 Mar 10]. Available from: https://arxiv.org/abs/2401.17868 doi:10.48550/ARXIV.2401.17868
-
[12]
Parameter-efficient fine-tuning for medical image analysis: The missed opportunity,
Dutt R, Ericsson L, Sanchez P, Tsaftaris SA, Hospedales T. Parameter-Efficient Fine-Tuning for Medical Image Analysis: The Missed Opportunity [Internet]. arXiv; 2023 [cited 2026 Mar 10]. Available from: https://arxiv.org/abs/2305.08252 doi:10.48550/ARXIV.2305.08252
-
[13]
Silva-Rodríguez J, Dolz J, Ben Ayed I. Towards Foundation Models and Few-Shot Parameter-Efficient Fine-Tuning for Volumetric Organ Segmentation. Medical Image Analysis. 2025 Jul;103:103596. doi:10.1016/j.media.2025.103596
-
[14]
Zhuang Y, Mathai TS, Mukherjee P, Khoury B, Kim B, Hou B, et al. MRISegmenter: A Fully Accurate and Robust Automated Multiorgan and Structure Segmentation Tool for T1-weighted Abdominal MRI. Radiology. 2025 Apr 1;315(1):e241979. doi:10.1148/radiol.241979
-
[15]
arXiv preprint arXiv:1411.1792 , year=
Yosinski J, Clune J, Bengio Y, Lipson H. How transferable are features in deep neural networks? [Internet]. 2014. doi:10.48550/ARXIV.1411.1792
-
[16]
nnU-Net: Self-adapting Framework for U-Net-Based Medical Image Segmentation [Internet]
Isensee F, Petersen J, Klein A, Zimmerer D, Jaeger PF, Kohl S, et al. nnU-Net: Self-adapting Framework for U-Net-Based Medical Image Segmentation [Internet]. arXiv; 2018 [cited 2026 Mar 15]. Available from: http://arxiv.org/abs/1809.10486 doi:10.48550/arXiv.1809.10486
-
[17]
MONAI: An open-source framework for deep learning in healthcare
Cardoso MJ, Li W, Brown R, Ma N, Kerfoot E, Wang Y, et al. MONAI: An open-source framework for deep learning in healthcare [Internet]. arXiv; 2022 [cited 2026 Mar 19]. Available from: http://arxiv.org/abs/2211.02701 doi:10.48550/arXiv.2211.02701
work page internal anchor Pith review doi:10.48550/arxiv.2211.02701 2022
-
[18]
Three mechanisms of weight decay regularization, 2018
Zhang G, Wang C, Xu B, Grosse R. Three Mechanisms of Weight Decay Regularization [Internet]. arXiv; 2018 [cited 2026 May 5]. Available from: http://arxiv.org/abs/1810.12281 doi:10.48550/arXiv.1810.12281
-
[19]
AMOS: A Large-Scale Abdominal Multi- Organ Benchmark for Versatile Medical Image Segmentation
Ji Y, Bai H, GE C, Yang J, Zhu Y, Zhang R, et al. AMOS: A Large-Scale Abdominal Multi- Organ Benchmark for Versatile Medical Image Segmentation. In: Koyejo S, Mohamed S, Agarwal A, Belgrave D, Cho K, Oh A, editors. Advances in Neural Information Processing Systems [Internet]. Curran Associates, Inc.; 2022. p. 36722–32. Available from: https://proceedings....
2022
-
[20]
CHAOS - Combined (CT-MR) Healthy Abdominal Organ Segmentation Challenge Data [Internet]
Kavur AE, Selver MA, Dicle O, Barış M, Gezer NS. CHAOS - Combined (CT-MR) Healthy Abdominal Organ Segmentation Challenge Data [Internet]. Zenodo; 2019 [cited 2026 Mar 12]. Available from: https://zenodo.org/record/3362844 doi:10.5281/ZENODO.3362844
-
[21]
Guarnera F, Rondinella A, Crispino E, Russo G, Di Lorenzo C, Maimone D, et al. MSLesSeg: baseline and benchmarking of a new Multiple Sclerosis Lesion Segmentation dataset. Sci Data. 2025 May 31;12(1):920. doi:10.1038/s41597-025-05250-y
-
[22]
Longitudinal multiple sclerosis lesion segmentation data resource
Carass A, Roy S, Jog A, Cuzzocreo JL, Magrath E, Gherman A, et al. Longitudinal multiple sclerosis lesion segmentation data resource. Data in Brief. 2017 Jun;12:346–50. doi:10.1016/j.dib.2017.04.004 Supplementary materials Figure S1. Validation loss and DICE curves in the AMOS subset training. The loss and DICE curves on the validation dataset indicate th...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.