HyGAS: an Open, Sensor-Agnostic Platform for Multi-Satellite Methane Plume Retrieval, Uncertainty Propagation, and Emission-Rate Estimation
HyGAS standardizes retrieval from raw data or products, uncertainty tracking, and flux estimation for comparable results across PRISMA, EnMA
abstract
click to expand
The rapid expansion of spaceborne methane observing capabilities at the facility-scale (fostered both by public missions and commercial constellations) has created a need for harmonised, reproducible, and uncertainty-aware processing chains that support both monitoring workflows and fair inter-sensor comparisons. This paper presents HyGAS (Hyperspectral Gas Analysis Suite), an open and sensor-agnostic framework that standardises methane processing across multiple imaging spectrometers. HyGAS currently supports end-to-end processing from Level-1 radiance to methane enhancement for PRISMA, EnMAP, and Tanager-1, and it supports ingestion of Level-2 methane enhancement products from EMIT and GHGSat, which are subsequently processed through common downstream modules for background selection, plume segmentation, Integrated Mass Enhancement (IME), and emission-rate inversion. HyGAS prioritises operational robustness via (i) matched-filter variants designed to mitigate background heterogeneity and pushbroom artefacts, (ii) explicit decomposition and propagation of uncertainty from instrument noise and scene-driven clutter to IME and flux, and (iii) a scale-aware segmentation strategy defined in physical units and rescaled by ground sampling distance to improve multi-sensor comparability. Representative sample outputs are reported for PRISMA, EnMAP, and Tanager-1.
Keywords: Methane emissions, hyperspectral satellites, Tanager-1, PRISMA, EnMAP, GHGSat, EMIT, Tanager, oil and gas, landfills, remote sensing, atmospheric science, greenhouse gas monitoring, spectral analysis, emission quantification, satellite synergy, environmental monitoring.
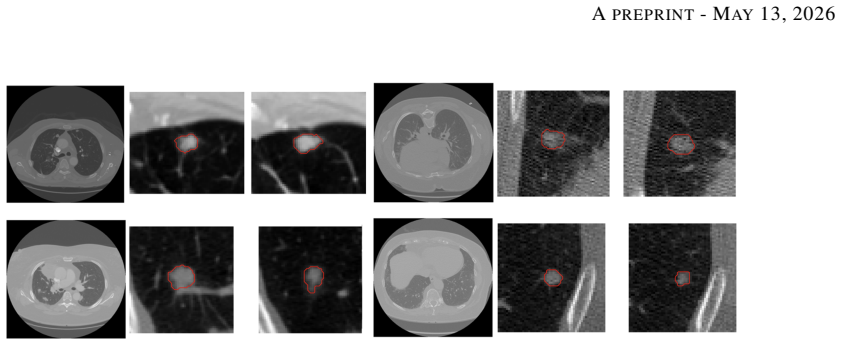 full image
full image