Recognition: 2 theorem links
· Lean TheoremThe Origin of Life in the Light of Evolution
Pith reviewed 2026-05-12 02:19 UTC · model grok-4.3
The pith
The last universal common ancestor was already a complex, ecologically adapted population far from life's starting point.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
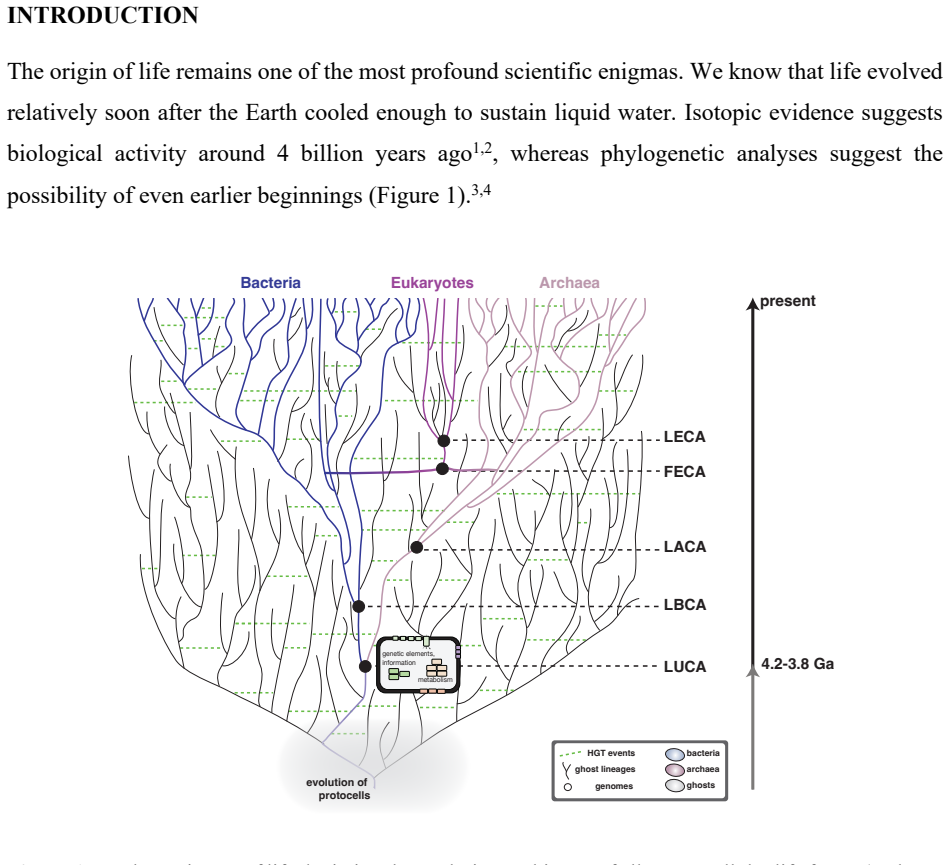
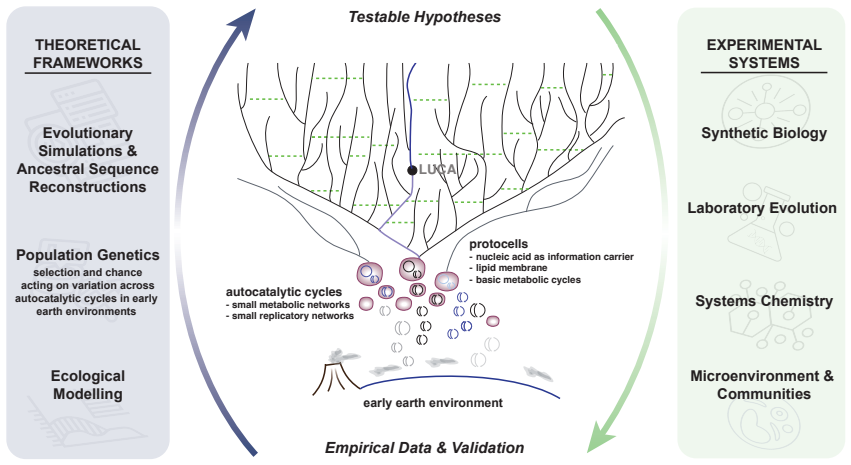
Synthesizing evidence from comparative genomics, phylogenetics, biochemistry, and geoscience, the paper establishes that LUCA was already a complex, ecologically adapted population far removed from the starting point of life, implying a deep pre-LUCA evolutionary history. It argues that origin-of-life research must expand to explore how pre-biological systems evolved under the influence of selection, drift, mutation, horizontal gene transfer, parasites, and compartmentalization. The authors outline an agenda spanning protometabolic and autocatalytic networks, protocells, the emergence of translation, and the transition to DNA genomes, where qualitative models can be formalized by evolution-
What carries the argument
Phylogenetic reconstruction of LUCA from modern genomes, which serves as the evidence that early life had already undergone substantial evolutionary change before the last universal ancestor.
If this is right
- Origin-of-life models must incorporate population genetics to explain how selection and drift shaped early communities.
- Scenarios for prebiotic systems must account for horizontal gene transfer and parasites as forces acting on protocell populations.
- Synthetic biology experiments can test specific evolutionary transitions such as the origin of translation and DNA genomes.
- Qualitative chemical models of life's beginnings become viable only when paired with evolution-driven, testable hypotheses.
Where Pith is reading between the lines
- Chemical origin scenarios must produce systems that quickly support populations capable of responding to selection rather than isolated molecules.
- The perspective links the origin of life to later evolutionary questions by showing that cellular complexity appeared early under population dynamics.
- Laboratory evolution of synthetic cells could be used to explore timelines and constraints on pre-LUCA stages.
Load-bearing premise
Comparative genomics and phylogenetics can reliably reconstruct the complexity and ecological adaptations of LUCA without being limited by method assumptions or the lack of direct evidence from that era.
What would settle it
Improved phylogenetic reconstructions or additional genomic data that show LUCA lacked key complex traits such as translation machinery or membrane components would disprove the claim of its advanced state.
Figures
read the original abstract
The origin of life is often framed primarily as a chemical problem, yet life's defining feature is evolution. Advances in geochemistry, prebiotic chemistry, and molecular biology have produced diverse scenarios for the emergence of genomes, metabolism, and cellular compartments on the early Earth, but most of these models lack a population-genetics framework. Here, we argue that origin-of-life research must expand from asking simply how life began to exploring how it evolved from pre-biological systems. Synthesizing evidence from comparative genomics, phylogenetics, biochemistry, and geoscience, we emphasize that the last universal common ancestor (LUCA) was already a complex, ecologically adapted population far removed from the starting point of life, implying a deep pre-LUCA evolutionary history. We highlight how population genetics, ecology, and synthetic biology can constrain origin-of-life scenarios by making explicit the roles of selection, drift, mutation, horizontal gene transfer, parasites, and compartmentalization in shaping early communities. Finally, we outline an evolutionary research agenda spanning protometabolic and autocatalytic networks, protocells, the emergence of translation, and the transition to DNA genomes, in which qualitative models can now be buttressed and formalized by evolution-driven hypotheses subject to testing using theory and laboratory experiments, including those with synthetic cells.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript is a perspective article arguing that origin-of-life research must incorporate an evolutionary framework, particularly population genetics, ecology, and synthetic biology. It synthesizes evidence from comparative genomics and phylogenetics to claim that LUCA was already a complex, ecologically adapted population with substantial pre-LUCA evolutionary history, and it outlines a research agenda for topics including protometabolic networks, protocells, the emergence of translation, and the transition to DNA genomes, emphasizing roles of selection, drift, HGT, parasites, and compartmentalization.
Significance. If the synthesis of existing phylogenetic evidence holds, the paper could meaningfully advance the field by promoting testable, evolution-informed models that bridge prebiotic chemistry with early cellular life. The explicit call for population-genetics constraints on origin scenarios is a logical extension of cited literature and could guide future synthetic biology experiments, though the perspective nature means impact depends on how well the agenda is adopted.
minor comments (2)
- [Abstract] Abstract: the statement that 'qualitative models can now be buttressed and formalized by evolution-driven hypotheses subject to testing' would be strengthened by naming at least one specific model or hypothesis (e.g., a particular autocatalytic network) and indicating a concrete test via theory or experiment.
- [Main synthesis section (implied after abstract)] The synthesis of LUCA complexity from comparative genomics is presented without a dedicated subsection addressing known methodological limitations such as horizontal gene transfer biases or sparse early-Earth taxon sampling; adding a short paragraph on these would improve balance while preserving the central claim.
Simulated Author's Rebuttal
We thank the referee for their positive and accurate summary of our perspective article, which correctly captures our central argument that origin-of-life research must be situated within an evolutionary framework, recognizing LUCA as a complex, ecologically adapted population with substantial pre-LUCA history. We appreciate the recommendation for minor revision and the recognition of the paper's potential to guide future work at the interface of phylogenetics, population genetics, and synthetic biology.
Circularity Check
No significant circularity detected
full rationale
The manuscript is a perspective article synthesizing prior literature on LUCA complexity from comparative genomics and phylogenetics without new derivations, equations, fitted parameters, or formal models. Central claims rest on established external reconstructions rather than internal reductions to self-definitions or author-overlapping citations. No load-bearing steps equate to inputs by construction, and the argument remains self-contained against independent benchmarks.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Comparative genomics and phylogenetics can reconstruct key features of LUCA and pre-LUCA life despite the absence of direct fossils or molecular evidence from that period.
- domain assumption Population-genetic processes (selection, drift, HGT, parasites) operated in prebiotic and protocell systems in ways that can be modeled and tested.
Lean theorems connected to this paper
-
IndisputableMonolith/Foundation/RealityFromDistinction.leanreality_from_one_distinction unclearThe LUCA was already a complex, ecologically adapted population far removed from the starting point of life, implying a deep pre-LUCA evolutionary history... population genetics, ecology, and synthetic biology can constrain origin-of-life scenarios
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearautocatalytic cycles... selection and chance acting on variation across autocatalytic cycles... compartmentalization
Reference graph
Works this paper leans on
-
[1]
LUCA used both types of phospholipids to produce mixed membranes, with alternative biosynthetic pathways subsequently lost in bacteria or archaea. Notably, mixed lipids are capable of producing protocell-like vesicles77, and E. coli can grow, albeit with a drop in fitness, with a mixture of typical bacterial phospholipids and up to 30% archaeal phospholip...
work page 2015
-
[2]
Crapitto, A.J., Campbell, A., Harris, A.J., and Goldman, A.D. (2022). A consensus view of the proteome of the last universal common ancestor. Ecol Evol 12, e8930. 23. Lombard, J., López-García, P., and Moreira, D. (2012). The early evolution of lipid membranes and the three domains of life. Nat Rev Microbiol 10, 507-515. 24. Adam, P.S., Borrel, G., and Gr...
work page 2022
-
[3]
Adamski, P., Eleveld, M., Sood, A., Kun, Á., Szilágyi, A., Czárán, T., Szathmáry, E., and Otto, S. (2020). From self-replication to replicator systems en route to de novo life. Nat Rev Chem 4, 386-403. 42. Babajanyan, S.G., Wolf, Y.I., Khachatryan, A., Allahverdyan, A., Lopez-Garcia, P., and Koonin, E.V. (2023). Coevolution of reproducers and replicators ...
work page 2020
-
[4]
Papastavrou, N., Horning, D.P., and Joyce, G.F. (2024). RNA-catalyzed evolution of catalytic RNA. Proceedings of the National Academy of Sciences 121, e2321592121. 62. Uhlmann, E., Peyman, A., Breipohl, G., and Will, D.W. (1998). PNA: Synthetic polyamide nucleic acids with unusual binding properties. Angew Chem Int Ed Engl 37, 2796-2823. 63. Benner, S.A.,...
work page 2024
-
[5]
Caforio, A., Siliakus, M.F., Exterkate, M., Jain, S., Jumde, V.R., Andringa, R.L.H., Kengen, S.W.M., Minnaard, A.J., Driessen, A.J.M., and van der Oost, J. (2018). Converting Escherichia coli into an archaebacterium with a hybrid heterochiral membrane. Proc Natl Acad Sci U S A 115, 3704-3709. 79. Łapińska, U., Glover, G., Kahveci, Z., Irwin, N.A.T., Milne...
-
[6]
Roger, A.J., Muñoz-Gómez, S.A., and Kamikawa, R. (2017). The Origin and Diversification of Mitochondria. Curr Biol 27, 1177-1192. 96. Aravind, L., Anantharaman, V., and Koonin, E.V. (2002). Monophyly of class I aminoacyl tRNA synthetase, USPA, ETFP, photolyase, and PP-ATPase nucleotide-binding domains: implications for protein evolution in the RNA. Protei...
work page 2017
-
[7]
Kaur, H., Rauscher, S.A., Werner, E., Song, Y., Yi, J., Kazone, W., Martin, W.F., Tuysuz, H., and Moran, J. (2024). A prebiotic Krebs cycle analog generates amino acids with H(2) and NH(3) over nickel. Chem 10, 1528-1540. 116. Muchowska, K.B., Varma, S.J., Chevallot-Beroux, E., Lethuillier-Karl, L., Li, G., and Moran, J. (2017). Metals promote sequences o...
work page 2024
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.