Recognition: unknown
I see artifacts: ICA-based EEG artifact removal does not improve deep network decoding across three BCI tasks
Pith reviewed 2026-05-08 07:26 UTC · model grok-4.3
The pith
ICA-based artifact removal in EEG signals does not consistently improve deep network decoding across BCI tasks
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Component-rejected EEG data produced by Infomax or AMICA combined with ICLabel or MARA does not deliver consistently higher decoding accuracy in CNN or LSTM models than the same data without any rejection, across motor imagery, long-term memory, and visual memory tasks.
What carries the argument
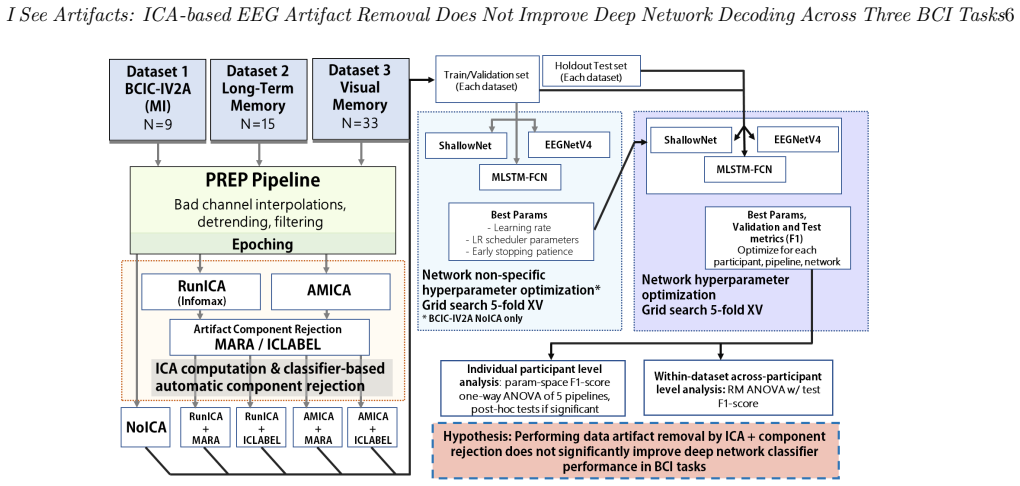
A pipeline matrix that applies ICA decomposition followed by optional component rejection before feeding the signals into neural network classifiers, then compares accuracy with and without the rejection step.
If this is right
- Deep networks can achieve comparable decoding performance on raw EEG without explicit ICA-based cleaning.
- The computational overhead of running ICA may not be justified when the goal is only to improve classification accuracy.
- Within-participant cross-validation results generalize similarly whether or not artifact rejection is applied.
Where Pith is reading between the lines
- In practice, BCI developers could skip ICA preprocessing to reduce latency in real-time systems.
- Attention may shift toward other preprocessing choices or architecture tweaks that more reliably affect performance.
- The finding raises the question of whether ICA still adds value when combined with newer model types or larger datasets.
Load-bearing premise
The three chosen datasets, two ICA algorithms, three rejection strategies, and three network architectures are representative of typical BCI decoding scenarios.
What would settle it
A follow-up experiment on new EEG datasets that finds statistically significant and consistent accuracy gains from the same ICA rejection pipelines would falsify the central claim.
Figures












read the original abstract
In this paper, we conduct a detailed investigation on the effect of independent component (IC)-based noise rejection methods in neural network classifier-based decoding of electroencephalography (EEG) data in different task datasets. We apply a pipeline matrix of two popular different independent component (IC) decomposition methods (Infomax and Adaptive Mixture Independent Component Analysis (AMICA)) with three different component rejection strategies (none, ICLabel, and multiple artifact rejection algorithm [MARA]) on three different EEG datasets (motor imagery, long-term memory formation, and visual memory). We cross-validate processed data from each pipeline with three architectures commonly used for EEG classification (two convolutional neural networks and one long short-term memory-based model. We compare decoding performances on within-participant and within-dataset levels.Our results show that the benefit from using IC-based noise rejection for decoding analyses is at best minor, as component-rejected data did not show consistently better performance than data without rejections; especially given the significant computational resources required for independent component analysis (ICA) computations.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper conducts a systematic empirical evaluation of ICA-based artifact removal for deep neural network-based EEG decoding. It tests two ICA decomposition methods (Infomax and AMICA), three rejection strategies (none, ICLabel, MARA), across three datasets (motor imagery, long-term memory formation, visual memory), and three network architectures (two CNNs and one LSTM). Using within-participant and within-dataset cross-validation, the central finding is that data with component rejection does not consistently outperform data without rejection, indicating that the benefit of IC-based noise rejection is at best minor, particularly in light of the computational demands of ICA.
Significance. This negative result, if substantiated, is significant for the BCI and EEG machine learning community as it challenges the routine application of ICA preprocessing in deep learning pipelines for EEG classification. The strength lies in the broad experimental matrix covering multiple methods, datasets, and models, providing a more robust test than single-dataset studies. The authors deserve credit for the comprehensive design, the use of held-out cross-validation, and the focus on practical implications regarding computational resources. This could encourage reevaluation of preprocessing steps in resource-constrained BCI applications.
major comments (1)
- Results section: The claim that component-rejected data 'did not show consistently better performance' and that the benefit is 'at best minor' is central but unsupported by reported effect sizes, mean accuracy deltas, standard deviations, or statistical tests (e.g., paired t-tests or Wilcoxon tests across pipelines). Without these in tables or text for each dataset-architecture combination, the strength of the negative conclusion cannot be fully assessed.
Simulated Author's Rebuttal
We thank the referee for their constructive review and positive assessment of the manuscript's scope and design. We agree that the central negative finding requires stronger quantitative backing and will revise the Results section to include the requested metrics and tests.
read point-by-point responses
-
Referee: Results section: The claim that component-rejected data 'did not show consistently better performance' and that the benefit is 'at best minor' is central but unsupported by reported effect sizes, mean accuracy deltas, standard deviations, or statistical tests (e.g., paired t-tests or Wilcoxon tests across pipelines). Without these in tables or text for each dataset-architecture combination, the strength of the negative conclusion cannot be fully assessed.
Authors: We accept this criticism. In the revised manuscript we will add a table (main text or supplementary) that reports, for every dataset–architecture pair, the mean accuracy and standard deviation under each preprocessing pipeline (none, ICLabel, MARA) for both Infomax and AMICA decompositions. We will also compute and report mean accuracy deltas relative to the no-rejection baseline and will apply paired non-parametric tests (Wilcoxon signed-rank) across participants or folds, together with effect-size measures (e.g., rank-biserial correlation). These additions will allow readers to evaluate both the magnitude and statistical reliability of any observed differences. revision: yes
Circularity Check
No significant circularity: purely empirical comparison
full rationale
The paper reports an empirical matrix of ICA pipelines (Infomax/AMICA × none/ICLabel/MARA) applied to three EEG datasets, decoded by three neural architectures, with performance measured via within-participant and within-dataset cross-validation. No derivations, equations, fitted parameters renamed as predictions, or self-citation chains appear in the abstract or described methods. The central negative result (component-rejected data not consistently superior) follows directly from comparing measured accuracies on held-out folds; it does not reduce to any input by construction. This is a standard, self-contained empirical study with no load-bearing self-referential steps.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Independent components recovered by Infomax and AMICA correspond to physiologically meaningful sources that can be labeled as artifacts.
- domain assumption Within-participant cross-validation on the chosen datasets provides a fair estimate of decoding performance.
Reference graph
Works this paper leans on
-
[1]
Results 3.1. Component Rejections In the BCI Competition IV-2A dataset, an average of 0.1 artifact components were rejected from AMICA component data rejected with ICLabel, while an average of 4.9 artifacts components were rejected through MARA. In data processed with RunICA, an average of 0.1 components were rejected through ICLabel, and 5.1 through MARA...
-
[2]
10), while all 15 participants had significant p-values in MLSTM-FCN
and Shallownet (p > .05 in ptc. 10), while all 15 participants had significant p-values in MLSTM-FCN. As can be seen in the circos plots, the chosen best performing parameters were highly variable between participants (and somewhat even across pipelines in same participant) for MLSTM-FCN and ShallowNet, while they were somewhat consistent for EEGNetV4 in ...
-
[3]
To see whether the results would be generalizable, we chose 3 different network architectures, including two specifically designed for brain signal classification
Discussion In this study, we pre-processed 3 datasets of epoched EEG signals purposed for BCI classification through a total of 4 (+1 control) different ICA-based automatic pipelines for rejecting artifact components, to investigate whether such pre-processing procedures affect general classifier performance in neural network- based architectures. To see ...
-
[4]
Acknowledgments This study was supported by the National Re- search Foundation of Korea under project BK21 FOUR and grants NRF-2022R1A2C2092118, NRF- 2022R1H1A2092007, NRF-2019R1A2C2007612, as well as by Institute of Information & Communica- tions Technology Planning & Evaluation (IITP) grants funded by the Korea government (No. 2017-0-00451, Development ...
-
[5]
Committee report: publication guidelines and recommendations for studies using electroencephalography and magnetoencephalography,
A. Keil, S. Debener, G. Gratton, M. Jungh¨ ofer, E. S. Kappenman, S. J. Luck, P. Luu, G. A. Miller, and C. M. Yee, “Committee report: publication guidelines and recommendations for studies using electroencephalography and magnetoencephalography,” Psychophysiology, vol. 51, no. 1, pp. 1–21, 2014
2014
-
[6]
Brain-computer interface technologies: from signal to action,
A. Ortiz-Rosario and H. Adeli, “Brain-computer interface technologies: from signal to action,”Reviews in the Neurosciences, vol. 24, no. 5, pp. 537–552, 2013
2013
-
[7]
Good practice for conducting and reporting meg research,
J. Gross, S. Baillet, G. R. Barnes, R. N. Henson, A. Hillebrand, O. Jensen, K. Jerbi, V. Litvak, B. Maess, R. Oostenveldet al., “Good practice for conducting and reporting meg research,”Neuroimage, vol. 65, pp. 349– 363, 2013
2013
-
[8]
Preprocessing of eeg,
S.-P. Kim, “Preprocessing of eeg,” inComputational EEG Analysis. Springer, 2018, pp. 15–33
2018
-
[9]
The PREP pipeline: Standardized preprocessing for large-scale EEG analysis,
N. Bigdely-Shamlo, T. Mullen, C. Kothe, K.-M. Su, and K. A. Robbins, “The PREP pipeline: Standardized preprocessing for large-scale EEG analysis,”Frontiers in Neuroinformatics, vol. 9, p. 16, 2015
2015
-
[10]
Automated eeg mega- analysis i: Spectral and amplitude characteristics across studies,
N. Bigdely-Shamlo, J. Touryan, A. Ojeda, C. Kothe, T. Mullen, and K. Robbins, “Automated eeg mega- analysis i: Spectral and amplitude characteristics across studies,”NeuroImage, vol. 207, p. 116361, 2020
2020
-
[11]
Comparison of eeg preprocessing methods to improve the classification of p300 trials,
Z. Cashero, “Comparison of eeg preprocessing methods to improve the classification of p300 trials,”Master of Science Thesis, Colorado State University, 2011
2011
-
[12]
Compar- ison of different wavelet features from eeg signals for classifying human emotions,
M. Murugappan, R. Nagarajan, and S. Yaacob, “Compar- ison of different wavelet features from eeg signals for classifying human emotions,” in2009 IEEE symposium on industrial electronics & applications, vol. 2. IEEE, 2009, pp. 836–841
2009
-
[13]
To explore the potentials of independent component analysis in brain- computer interface of motor imagery,
X. Wu, B. Zhou, Z. Lv, and C. Zhang, “To explore the potentials of independent component analysis in brain- computer interface of motor imagery,”IEEE Journal of Biomedical and Health Informatics, vol. 24, no. 3, pp. 775–787, 2019
2019
-
[14]
The correction of eye blink artefacts in the eeg: a comparison of two prominent methods,
S. Hoffmann and M. Falkenstein, “The correction of eye blink artefacts in the eeg: a comparison of two prominent methods,”PloS one, vol. 3, no. 8, p. e3004, 2008
2008
-
[15]
Removing electroencephalographic artifacts: comparison between ica and pca,
T.-P. Jung, C. Humphries, T.-W. Lee, S. Makeig, M. J. McKeown, V. Iragui, and T. J. Sejnowski, “Removing electroencephalographic artifacts: comparison between ica and pca,” inNeural Networks for Signal Processing VIII. Proceedings of the 1998 IEEE Signal Processing Society Workshop (Cat. No. 98TH8378). IEEE, 1998, pp. 63–72
1998
-
[16]
The Harvard Automated Pro- cessing Pipeline for Electroencephalography (HAPPE): Standardized processing software for developmental and high-artifact data,
L. J. Gabard-Durnam, A. S. Mendez Leal, C. L. Wilkin- son, and A. R. Levin, “The Harvard Automated Pro- cessing Pipeline for Electroencephalography (HAPPE): Standardized processing software for developmental and high-artifact data,”Frontiers in Neuroscience, vol. 12, p. 97, 2018
2018
-
[17]
An automatic pre-processing pipeline for EEG Analysis (APP) based on robust statistics,
J. R. da Cruz, V. Chicherov, M. H. Herzog, and P. Figueiredo, “An automatic pre-processing pipeline for EEG Analysis (APP) based on robust statistics,” Clinical Neurophysiology, vol. 129, no. 7, pp. 1427–1437, 2018
2018
-
[18]
Automagic: Standardized preprocessing of big EEG data,
A. Pedroni, A. Bahreini, and N. Langer, “Automagic: Standardized preprocessing of big EEG data,”Neu- roImage, 2019
2019
-
[19]
Robust artifactual independent component classification for BCI practitioners,
I. Winkler, S. Brandl, F. Horn, E. Waldburger, C. Allefeld, and M. Tangermann, “Robust artifactual independent component classification for BCI practitioners,”Jour- nal of Neural Engineering, vol. 11, no. 3, p. 035013, 2014
2014
-
[20]
Iclabel: An automated electroencephalographic inde- pendent component classifier, dataset, and website,
L. Pion-Tonachini, K. Kreutz-Delgado, and S. Makeig, “Iclabel: An automated electroencephalographic inde- pendent component classifier, dataset, and website,” NeuroImage, vol. 198, pp. 181–197, 2019
2019
-
[21]
Investigation into the origin of the noise of surface electrodes,
E. Huigen, A. Peper, and C. Grimbergen, “Investigation into the origin of the noise of surface electrodes,” Medical and biological engineering and computing, vol. 40, pp. 332–338, 2002
2002
-
[22]
Adaptive digital notch filter design on the unit circle for the removal of powerline noise from biomedical signals,
M. Ferdjallah and R. E. Barr, “Adaptive digital notch filter design on the unit circle for the removal of powerline noise from biomedical signals,”IEEE transactions on biomedical engineering, vol. 41, no. 6, pp. 529–536, 1994
1994
-
[23]
Filters: when, why, and how (not) to use them,
A. de Cheveign´ e and I. Nelken, “Filters: when, why, and how (not) to use them,”Neuron, vol. 102, no. 2, pp. 280–293, 2019
2019
-
[24]
Removal of ocular artifacts from eeg using adaptive thresholding of wavelet coefficients,
V. Krishnaveni, S. Jayaraman, L. Anitha, and K. Ra- madoss, “Removal of ocular artifacts from eeg using adaptive thresholding of wavelet coefficients,”Journal of neural engineering, vol. 3, no. 4, p. 338, 2006
2006
-
[25]
Faster: fully automated statistical thresholding for eeg artifact rejection,
H. Nolan, R. Whelan, and R. B. Reilly, “Faster: fully automated statistical thresholding for eeg artifact rejection,”Journal of neuroscience methods, vol. 192, I See Artifacts: ICA-based EEG Artifact Removal Does Not Improve Deep Network Decoding Across Three BCI Tasks22 no. 1, pp. 152–162, 2010
2010
-
[26]
A generic eeg artifact removal algorithm based on the multi-channel wiener filter,
B. Somers, T. Francart, and A. Bertrand, “A generic eeg artifact removal algorithm based on the multi-channel wiener filter,”Journal of neural engineering, vol. 15, no. 3, p. 036007, 2018
2018
-
[27]
Optimal filters for erp research i: A general approach for selecting filter settings,
G. Zhang, D. R. Garrett, and S. J. Luck, “Optimal filters for erp research i: A general approach for selecting filter settings,”Psychophysiology, p. e14531, 2024
2024
-
[28]
Eeg artifact removal—state-of-the-art and guidelines,
J. A. Urig¨ uen and B. Garcia-Zapirain, “Eeg artifact removal—state-of-the-art and guidelines,”Journal of neural engineering, vol. 12, no. 3, p. 031001, 2015
2015
-
[29]
Univariate versus multivariate tests in repeated-measures experiments
M. L. Davidson, “Univariate versus multivariate tests in repeated-measures experiments.”Psychological Bul- letin, vol. 77, no. 6, p. 446, 1972
1972
-
[30]
Deconstructing multivariate decoding for the study of brain function,
M. N. Hebart and C. I. Baker, “Deconstructing multivariate decoding for the study of brain function,” Neuroimage, vol. 180, pp. 4–18, 2018
2018
-
[31]
The same analysis approach: Practical protection against the pitfalls of novel neuroimaging analysis methods,
K. G¨ orgen, M. N. Hebart, C. Allefeld, and J.-D. Haynes, “The same analysis approach: Practical protection against the pitfalls of novel neuroimaging analysis methods,”Neuroimage, vol. 180, pp. 19–30, 2018
2018
-
[32]
Does data cleaning improve brain state classification?
S. L. Meisler, M. J. Kahana, and Y. Ezzyat, “Does data cleaning improve brain state classification?”Journal of neuroscience methods, vol. 328, p. 108421, 2019
2019
-
[33]
Ica: a potential tool for bci systems,
A. Kachenoura, L. Albera, L. Senhadji, and P. Comon, “Ica: a potential tool for bci systems,”IEEE Signal Processing Magazine, vol. 25, no. 1, pp. 57–68, 2007
2007
-
[34]
Comparing results of algorithms implementing blind source separation of eeg data,
A. Delorme, J. Plamer, R. Oostenveld, J. Onton, and S. Makeig, “Comparing results of algorithms implementing blind source separation of eeg data,” Swartz foundation and NIH grant, 2007
2007
-
[35]
Independent component analysis as a tool to eliminate artifacts in eeg: a quantitative study,
J. Iriarte, E. Urrestarazu, M. Valencia, M. Alegre, A. Malanda, C. Viteri, and J. Artieda, “Independent component analysis as a tool to eliminate artifacts in eeg: a quantitative study,”Journal of clinical neurophysiology, vol. 20, no. 4, pp. 249–257, 2003
2003
-
[36]
Extraction of ocular artefacts from eeg using independent component analysis,
R. N. Vig´ ario, “Extraction of ocular artefacts from eeg using independent component analysis,”Electroen- cephalography and clinical neurophysiology, vol. 103, no. 3, pp. 395–404, 1997
1997
-
[37]
Enhanced detection of artifacts in eeg data using higher- order statistics and independent component analysis,
A. Delorme, T. Sejnowski, and S. Makeig, “Enhanced detection of artifacts in eeg data using higher- order statistics and independent component analysis,” Neuroimage, vol. 34, no. 4, pp. 1443–1449, 2007
2007
-
[38]
Independent component approach to the analysis of eeg and meg recordings,
R. Vig´ ario, J. Sarela, V. Jousmiki, M. Hamalainen, and E. Oja, “Independent component approach to the analysis of eeg and meg recordings,”IEEE transactions on biomedical engineering, vol. 47, no. 5, pp. 589–593, 2000
2000
-
[39]
Recovering eeg brain signals: Artifact suppression with wavelet enhanced independent component analysis,
N. P. Castellanos and V. A. Makarov, “Recovering eeg brain signals: Artifact suppression with wavelet enhanced independent component analysis,”Journal of neuroscience methods, vol. 158, no. 2, pp. 300–312, 2006
2006
-
[40]
Independent component analysis of electroencephalo- graphic data,
S. Makeig, A. J. Bell, T.-P. Jung, and T. J. Sejnowski, “Independent component analysis of electroencephalo- graphic data,” inAdvances in neural information pro- cessing systems, 1996, pp. 145–151
1996
-
[41]
Eeglab: an open source toolbox for analysis of single-trial eeg dynamics including independent component analysis,
A. Delorme and S. Makeig, “Eeglab: an open source toolbox for analysis of single-trial eeg dynamics including independent component analysis,”Journal of neuroscience methods, vol. 134, no. 1, pp. 9–21, 2004
2004
-
[42]
Fieldtrip: Open source software for advanced analysis of meg, eeg, and invasive electrophysiological data,
R. Oostenveld, P. Fries, E. Maris, and J.-M. Schoffelen, “Fieldtrip: Open source software for advanced analysis of meg, eeg, and invasive electrophysiological data,” Computational Intelligence and Neuroscience, vol. 2011, p. 1, 2011
2011
-
[43]
Mne software for processing meg and eeg data,
A. Gramfort, M. Luessi, E. Larson, D. A. Engemann, D. Strohmeier, C. Brodbeck, L. Parkkonen, and M. S. H¨ am¨ al¨ ainen, “Mne software for processing meg and eeg data,”Neuroimage, vol. 86, pp. 446–460, 2014
2014
-
[44]
An information- maximization approach to blind separation and blind deconvolution,
A. J. Bell and T. J. Sejnowski, “An information- maximization approach to blind separation and blind deconvolution,”Neural computation, vol. 7, no. 6, pp. 1129–1159, 1995
1995
-
[45]
A unifying information-theoretic framework for inde- pendent component analysis,
T.-W. Lee, M. Girolami, A. J. Bell, and T. J. Sejnowski, “A unifying information-theoretic framework for inde- pendent component analysis,”Computers & Mathemat- ics with Applications, vol. 39, no. 11, pp. 1–21, 2000
2000
-
[46]
Independent component anal- ysis: algorithms and applications,
A. Hyv¨ arinen and E. Oja, “Independent component anal- ysis: algorithms and applications,”Neural networks, vol. 13, no. 4-5, pp. 411–430, 2000
2000
-
[47]
Cichocki and S.-i
A. Cichocki and S.-i. Amari,Adaptive blind signal and image processing: learning algorithms and applications. John Wiley & Sons, 2002
2002
-
[48]
Robust blind source separation algorithms using cumulants,
S. Cruces, L. Castedo, and A. Cichocki, “Robust blind source separation algorithms using cumulants,” Neurocomputing, vol. 49, no. 1-4, pp. 87–118, 2002
2002
-
[49]
Amica: An adaptive mixture of independent component analyzers with shared components,
J. A. Palmer, K. Kreutz-Delgado, and S. Makeig, “Amica: An adaptive mixture of independent component analyzers with shared components,”Swartz Center for Computatonal Neursoscience, University of California San Diego, Tech. Rep, 2012
2012
-
[50]
How sensitive are eeg results to preprocessing methods: a benchmarking study,
K. A. Robbins, J. Touryan, T. Mullen, C. Kothe, and N. Bigdely-Shamlo, “How sensitive are eeg results to preprocessing methods: a benchmarking study,” IEEE transactions on neural systems and rehabilitation engineering, vol. 28, no. 5, pp. 1081–1090, 2020
2020
-
[51]
Automatic classification of artifactual ica-components for artifact removal in eeg signals,
I. Winkler, S. Haufe, and M. Tangermann, “Automatic classification of artifactual ica-components for artifact removal in eeg signals,”Behavioral and brain functions, vol. 7, no. 1, pp. 1–15, 2011
2011
-
[52]
Artifact removal techniques with signal reconstruction,
C. A. E. Kothe and T.-P. Jung, “Artifact removal techniques with signal reconstruction,” Apr. 28 2016, uS Patent App. 14/895,440
2016
-
[53]
Eeg is better left alone,
A. Delorme, “Eeg is better left alone,”Scientific reports, vol. 13, no. 1, p. 2372, 2023
2023
-
[54]
Removal of ocular artifacts using ica and adaptive filter for motor imagery-based bci,
C. S. Kim, J. Sun, D. Liu, Q. Wang, and S. G. Paek, “Removal of ocular artifacts using ica and adaptive filter for motor imagery-based bci,”IEEE/CAA journal of automatica sinica, 2017
2017
-
[55]
A comparsion of artifact rejection methods for a bci using event related potentials,
M. Kim and S.-P. Kim, “A comparsion of artifact rejection methods for a bci using event related potentials,” in 2018 6th International Conference on Brain-Computer Interface (BCI). IEEE, 2018, pp. 1–4
2018
-
[56]
On the influence of high-pass filtering on ICA-based artifact reduction in EEG-ERP,
I. Winkler, S. Debener, K.-R. M¨ uller, and M. Tangermann, “On the influence of high-pass filtering on ICA-based artifact reduction in EEG-ERP,” in2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC). IEEE, 2015, pp. 4101–4105
2015
-
[57]
Deep learning for electroencephalogram (EEG) classification tasks: A review,
A. Craik, Y. He, and J. L. Contreras-Vidal, “Deep learning for electroencephalogram (EEG) classification tasks: A review,”Journal of Neural Engineering, vol. 16, no. 3, p. 031001, 2019
2019
-
[58]
Deep learning techniques for classification of electroencephalogram (eeg) motor imagery (mi) signals: a review,
H. Altaheri, G. Muhammad, M. Alsulaiman, S. U. Amin, G. A. Altuwaijri, W. Abdul, M. A. Bencherif, and M. Faisal, “Deep learning techniques for classification of electroencephalogram (eeg) motor imagery (mi) signals: a review,”Neural Computing and Applications, pp. 1– 42, 2021
2021
-
[59]
Inter-subject transfer learning with an end-to-end deep convolutional neural network for EEG- based ,
F. Fahimi, Z. Zhang, W. B. Goh, T.-S. Lee, K. K. Ang, and C. Guan, “Inter-subject transfer learning with an end-to-end deep convolutional neural network for EEG- based ,”Journal of neural engineering, vol. 16, no. 2, p. 026007, 2019
2019
-
[60]
Learning eeg topographical representation for classification via convolutional neural network,
M. Xu, J. Yao, Z. Zhang, R. Li, B. Yang, C. Li, J. Li, and J. Zhang, “Learning eeg topographical representation for classification via convolutional neural network,” Pattern Recognition, vol. 105, p. 107390, 2020. I See Artifacts: ICA-based EEG Artifact Removal Does Not Improve Deep Network Decoding Across Three BCI Tasks23
2020
-
[61]
Mi-eegnet: A novel convolutional neural network for motor imagery classification,
M. Riyad, M. Khalil, and A. Adib, “Mi-eegnet: A novel convolutional neural network for motor imagery classification,”Journal of Neuroscience Methods, vol. 353, p. 109037, 2021
2021
-
[62]
Op- timizing motor intention detection with deep learn- ing: towards management of intraoperative awareness,
O. Avilov, S. Rimbert, A. Popov, and L. Bougrain, “Op- timizing motor intention detection with deep learn- ing: towards management of intraoperative awareness,” IEEE Transactions on Biomedical Engineering, vol. 68, no. 10, pp. 3087–3097, 2021
2021
-
[63]
Multiclass classifica- tion of spatially filtered motor imagery eeg signals us- ing convolutional neural network for bci based applica- tions,
N. Shajil, S. Mohan, P. Srinivasan, J. Arivudaiyanambi, and A. Arasappan Murrugesan, “Multiclass classifica- tion of spatially filtered motor imagery eeg signals us- ing convolutional neural network for bci based applica- tions,”Journal of Medical and Biological Engineering, vol. 40, no. 5, pp. 663–672, 2020
2020
-
[64]
Study on the effect of different electrode channel combinations of motor imagery eeg signals on classification accuracy,
K. Zhu, S. Wang, D. Zheng, and M. Dai, “Study on the effect of different electrode channel combinations of motor imagery eeg signals on classification accuracy,” The Journal of Engineering, vol. 2019, no. 23, pp. 8641– 8645, 2019
2019
-
[65]
Mu rhythm (de) synchronization and eeg single-trial classification of different motor imagery tasks,
G. Pfurtscheller, C. Brunner, A. Schl¨ ogl, and F. L. Da Silva, “Mu rhythm (de) synchronization and eeg single-trial classification of different motor imagery tasks,”NeuroImage, vol. 31, no. 1, pp. 153–159, 2006
2006
-
[66]
Iterative gaussianization: from ica to random rotations,
V. Laparra, G. Camps-Valls, and J. Malo, “Iterative gaussianization: from ica to random rotations,”IEEE transactions on neural networks, vol. 22, no. 4, pp. 537– 549, 2011
2011
-
[67]
Eeg- based prediction of successful memory formation during vocabulary learning,
T. Kang, Y. Chen, S. Fazli, and C. Wallraven, “Eeg- based prediction of successful memory formation during vocabulary learning,”IEEE Transactions on Neural Systems and Rehabilitation Engineering, vol. 28, no. 11, pp. 2377–2389, 2020
2020
-
[68]
Eegmanypipelines: A large-scale, grass-root multi-analyst study of eeg analysis practices in the wild,
D. Tr¨ ubutschek, Y.-F. Yang, C. Gianelli, E. Cesnaite, N. L. Fischer, M. C. Vinding, T. Marshall, J. Algermissen, A. Pascarella, T. Puoliv¨ aliet al., “Eegmanypipelines: A large-scale, grass-root multi-analyst study of eeg analysis practices in the wild,” 2022
2022
-
[69]
Blind source separation,
G. R. Naik, W. Wanget al., “Blind source separation,” Berlin: Springer, vol. 10, pp. 978–3, 2014
2014
-
[70]
Heartbeat classification using morphological and dynamic features of ecg signals,
C. Ye, B. V. Kumar, and M. T. Coimbra, “Heartbeat classification using morphological and dynamic features of ecg signals,”IEEE Transactions on Biomedical Engineering, vol. 59, no. 10, pp. 2930–2941, 2012
2012
-
[71]
Application of independent component analysis with adaptive density model to complex- valued fmri data,
H. Li, N. M. Correa, P. A. Rodriguez, V. D. Calhoun, and T. Adali, “Application of independent component analysis with adaptive density model to complex- valued fmri data,”IEEE Transactions on biomedical engineering, vol. 58, no. 10, pp. 2794–2803, 2011
2011
-
[72]
Temporally constrained ica: an application to artifact rejection in electromag- netic brain signal analysis,
C. J. James and O. J. Gibson, “Temporally constrained ica: an application to artifact rejection in electromag- netic brain signal analysis,”IEEE Transactions on biomedical engineering, vol. 50, no. 9, pp. 1108–1116, 2003
2003
-
[73]
Removing electroencephalographic artifacts by blind source separation,
T.-P. Jung, S. Makeig, C. Humphries, T.-W. Lee, M. J. Mckeown, V. Iragui, and T. J. Sejnowski, “Removing electroencephalographic artifacts by blind source separation,”Psychophysiology, vol. 37, no. 2, pp. 163–178, 2000
2000
-
[74]
Blind source separation, wavelet denoising and discriminant analysis for eeg artefacts and noise cancelling,
R. R. V´ azquez, H. Velez-Perez, R. Ranta, V. L. Dorr, D. Maquin, and L. Maillard, “Blind source separation, wavelet denoising and discriminant analysis for eeg artefacts and noise cancelling,”Biomedical signal processing and control, vol. 7, no. 4, pp. 389–400, 2012
2012
-
[75]
An overview of independent component analysis and its applications,
G. R. Naik and D. K. Kumar, “An overview of independent component analysis and its applications,”Informatica, vol. 35, no. 1, 2011
2011
-
[76]
Ica-based eeg denoising: a comparative analysis of fifteen methods,
L. Albera, A. Kachenoura, P. Comon, A. Karfoul, F. Wendling, L. Senhadji, and I. Merlet, “Ica-based eeg denoising: a comparative analysis of fifteen methods,” Bulletin of the Polish Academy of Sciences: Technical Sciences, vol. 60, no. 3 Special issue on Data Mining in Bioengineering, pp. 407–418, 2012
2012
-
[77]
Independent eeg sources are dipolar,
A. Delorme, J. Palmer, J. Onton, R. Oostenveld, and S. Makeig, “Independent eeg sources are dipolar,”PloS one, vol. 7, no. 2, p. e30135, 2012
2012
-
[78]
A blind source separation technique using second-order statistics,
A. Belouchrani, K. Abed-Meraim, J.-F. Cardoso, and E. Moulines, “A blind source separation technique using second-order statistics,”IEEE Transactions on signal processing, vol. 45, no. 2, pp. 434–444, 1997
1997
-
[79]
Blind beamforming for non-gaussian signals,
J.-F. Cardoso and A. Souloumiac, “Blind beamforming for non-gaussian signals,” inIEE proceedings F (radar and signal processing), vol. 140, no. 6. IET, 1993, pp. 362– 370
1993
-
[80]
Modeling and estimation of dependent subspaces with non-radially symmetric and skewed densities,
J. A. Palmer, K. Kreutz-Delgado, B. D. Rao, and S. Makeig, “Modeling and estimation of dependent subspaces with non-radially symmetric and skewed densities,” inIndependent Component Analysis and Signal Separation: 7th International Conference, ICA 2007, London, UK, September 9-12, 2007. Proceedings
2007
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.