Recognition: unknown
Activation in Vesicle-Mediated Signaling Shaped by Batch Arrival Statistics
Pith reviewed 2026-05-08 04:41 UTC · model grok-4.3
The pith
Activation kinetics in vesicle signaling depend on arrival statistics beyond mean rates
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
We derive an exact solution for the full time-dependent probability distribution of a general batch arrival-degradation model using generating functions and a recursion relation. This enables a full analysis of first-passage times to a concentration threshold representing downstream activation. We show that activation kinetics are not determined by mean dynamics alone, but depend sensitively on the temporal statistics of arrival events, batch-size variability, and degradation, with different arrival processes having identical mean rates leading to qualitatively distinct first-passage behavior due to time-asymmetric fluctuations.
What carries the argument
Generating functions combined with a recursion relation to solve for the probability distribution in the batch arrival process with continuous degradation, enabling exact first-passage time analysis.
If this is right
- Activation timing varies with the specific arrival process even at fixed mean rate.
- Batch size variability affects the first-passage time distributions.
- Continuous degradation interacts with burst arrivals to break time-reversal symmetry in the dynamics.
- The framework allows incorporation of vesicle depletion effects for more realistic models.
Where Pith is reading between the lines
- Cells could exploit different release statistics to achieve faster or more reliable activation without changing average secretion rates.
- This mechanism might extend to other biological systems involving bursty molecular releases, such as in gene expression or immune responses.
- Future models could integrate this with spatial diffusion to see how location of vesicle fusions influences activation.
Load-bearing premise
The model assumes that activation is captured entirely by the first-passage time to a fixed concentration threshold in a well-mixed system with continuous degradation, without spatial effects or regulatory feedback.
What would settle it
Comparing measured first-passage time distributions in a vesicle secretion system for Poisson versus bursty arrivals with the same mean rate but different higher moments, to check if they differ as the model predicts.
Figures






read the original abstract
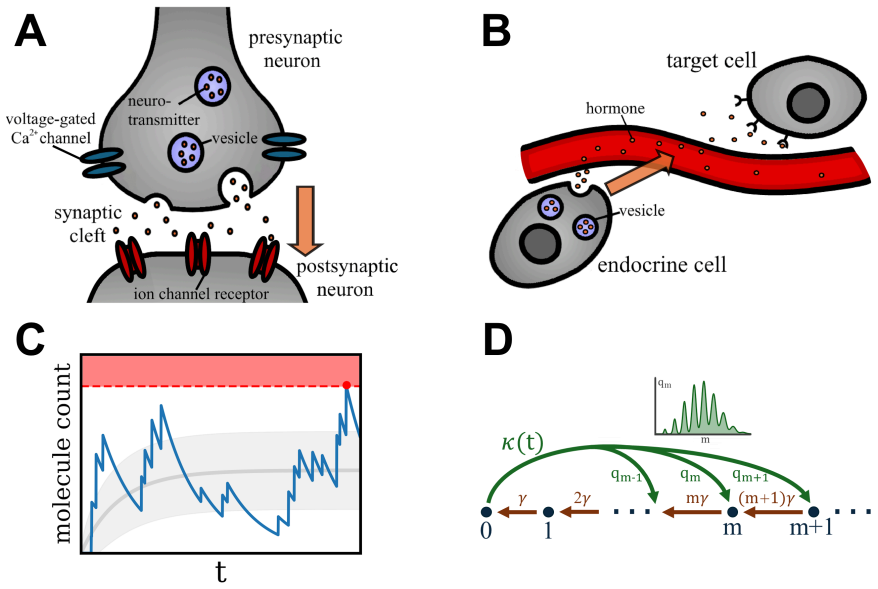
Vesicle-mediated secretion of ions or molecules is a central mechanism of cellular communication, for example in processes such as neurotransmission or hormone release. These events are inherently stochastic: vesicle fusions lead to bursts of variable sizes, releasing discrete packets of transmitters that are subsequently cleared or degraded. The dynamics break time-reversal symmetry due to the interplay of spontaneous bursts and continuous degradation. Using generating functions and a recursion relation, we derive an exact solution for the full time-dependent probability distribution of a general batch arrival-degradation model. This framework also enables a full analysis of first-passage times to a concentration threshold representing downstream activation. We show that activation kinetics are not determined by mean dynamics alone, but depend sensitively on the temporal statistics of arrival events, batch-size variability, and degradation. In particular, different arrival processes with identical mean rates can lead to qualitatively distinct first-passage behavior, reflecting the role of time-asymmetric fluctuations. We also discuss extensions incorporating vesicle depletion. Our results provide a transparent link between stochastic release dynamics and activation timing in vesicle-mediated signaling.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper derives an exact generating-function solution together with a recursion relation for the time-dependent probability distribution of molecule number in a general batch-arrival process subject to continuous linear degradation. It then uses this solution to compute first-passage times to a fixed concentration threshold that represents downstream activation in vesicle-mediated signaling, showing that these times depend on the full temporal statistics and batch-size distribution even when the mean arrival rate is held fixed.
Significance. If the exact solution and first-passage analysis hold, the work supplies an analytically tractable framework that demonstrates the insufficiency of mean-field descriptions for activation timing in stochastic secretion systems. This is a clear strength for the field of cellular signaling, where bursty release and degradation are common; the transparent link between arrival statistics and first-passage behavior, plus the sketched extension to vesicle depletion, offers a foundation for quantitative predictions without relying solely on simulation.
major comments (1)
- [Derivation of the recursion and first-passage analysis] The central claim that first-passage times are sensitive to arrival statistics beyond the mean rests on the generating function encoding the joint burst-time and burst-size distribution. The manuscript should explicitly verify that the recursion preserves normalization and non-negativity for at least one non-Poisson batch process (e.g., geometric batch sizes) at a concrete parameter set, to confirm that the claimed qualitative distinction in first-passage behavior is not an artifact of truncation or numerical inversion.
minor comments (2)
- [Abstract and results section on first-passage times] The abstract states that the framework 'enables a full analysis' of first-passage times, but the main text should clarify whether the first-passage distribution is obtained in closed form or via numerical inversion of the generating function; this affects how easily the results can be used by experimentalists.
- [Model definition and generating-function section] Notation for the batch-size probability generating function and the degradation rate should be introduced once with a single consistent symbol set; repeated re-definition across sections makes the recursion harder to follow.
Simulated Author's Rebuttal
We thank the referee for the positive assessment of our work and the recommendation for minor revision. We address the single major comment below and will incorporate the requested verification.
read point-by-point responses
-
Referee: [Derivation of the recursion and first-passage analysis] The central claim that first-passage times are sensitive to arrival statistics beyond the mean rests on the generating function encoding the joint burst-time and burst-size distribution. The manuscript should explicitly verify that the recursion preserves normalization and non-negativity for at least one non-Poisson batch process (e.g., geometric batch sizes) at a concrete parameter set, to confirm that the claimed qualitative distinction in first-passage behavior is not an artifact of truncation or numerical inversion.
Authors: We agree that an explicit numerical verification for a non-Poisson process would strengthen the manuscript. In the revised version we will add a short appendix (or subsection) that applies the recursion to geometric batch sizes with a concrete parameter set (mean batch size 4, arrival rate 0.8, degradation rate 0.5). We will report that the probability mass remains normalized to machine precision (sum p_n(t) = 1 within 10^{-12}) and that all probabilities stay non-negative up to the times used for first-passage calculations. This check will be performed both with the exact recursion and with the numerical inversion procedure employed in the paper, thereby confirming that the reported distinctions in first-passage behavior are not numerical artifacts. revision: yes
Circularity Check
No significant circularity in derivation chain
full rationale
The paper states a general batch-arrival process with continuous linear degradation, then applies standard generating-function methods plus a recursion to obtain the exact time-dependent distribution and first-passage times. The claim that different arrival processes with identical means produce distinct first-passage behavior follows directly from the fact that the generating function encodes the full joint statistics of burst times and sizes; this is a mathematical consequence of the stated model, not a reduction to fitted parameters or a self-citation loop. No load-bearing step is shown to be equivalent to its inputs by construction, and the framework is presented as self-contained.
Axiom & Free-Parameter Ledger
axioms (2)
- standard math Generating functions combined with a recursion relation yield the exact time-dependent probability distribution for the batch arrival-degradation model.
- domain assumption First-passage time to a fixed concentration threshold represents downstream activation.
Reference graph
Works this paper leans on
-
[1]
S¨ udhof T C 2004Annu. Rev. Neurosci.27509–547
-
[2]
S¨ udhof T C and Rizo J 2011Cold Spring Harbor perspectives in biology3a005637
-
[3]
Rizo J and Xu J 2015Annual review of biophysics44339–367
-
[4]
Lang J 1999European journal of biochemistry2593–17
-
[5]
Bergles D E, Diamond J S and Jahr C E 1999Current opinion in neurobiology9293–298
-
[6]
Brabant G, Prank K and Schofl C 1992Trends in Endocrinology & Metabolism3183–190
-
[7]
Veldhuis J D, Keenan D M and Pincus S M 2008Endocrine reviews29823–864
-
[8]
Ernst A, Sch¨ utte C, Sigrist S J and Winkelmann S 2022Mathematical Biosciences343108760
-
[9]
Ernst A, Unger N, Sch¨ utte C, Walter A M and Winkelmann S 2023Mathematical Biosciences 362109023
-
[10]
Johnston D and Wu S M S 1994Foundations of cellular neurophysiology(MIT press)
-
[11]
Lodish H F 2008Molecular cell biology(Macmillan)
-
[12]
Ghusinga K R and Singh A 2015 Theoretical predictions on the first-passage time for a gene expression model2015 54th IEEE Conference on Decision and Control (CDC)(IEEE) pp 3864–3869
2015
-
[13]
Gerstner W, Kistler W M, Naud R and Paninski L 2014Neuronal dynamics: From single neurons to networks and models of cognition(Cambridge University Press)
-
[14]
Alberts B, Heald R, Johnson A, Morgan D, Raff M, Roberts K and Walter P 2022Molecular biology of the cell: seventh international student edition with registration card(WW Norton & Company)
-
[15]
Gillespie C S and Renshaw E 2005Proceedings of the Royal Society A: Mathematical, Physical and Engineering Sciences4611563–1581
-
[16]
Daw A and Pender J 2019Queueing Systems91367–401 13
-
[17]
Bhattacharya R N and Waymire E C 2021Random walk, Brownian motion, and martingales vol 52 (Springer)
-
[18]
Paulsson J 2005Physics of life reviews2157–175
-
[19]
Shahrezaei V and Swain P S 2008Proceedings of the National Academy of Sciences105 17256–17261
-
[20]
Tunnacliffe E and Chubb J R 2020Trends in Genetics36288–297
-
[21]
Rijal K, Prasad A, Singh A and Das D 2022Physical Review Letters128048101
-
[22]
Ham L, Coomer M A, ¨Ocal K, Grima R and Stumpf M P 2024Nature Communications15 5286
-
[23]
Szavits-Nossan J and Grima R 2024Biophysical Journal1231034–1057
-
[24]
Elowitz M B, Levine A J, Siggia E D and Swain P S 2002Science2971183–1186
-
[25]
Ozbudak E M, Thattai M, Kurtser I, Grossman A D and Van Oudenaarden A 2002Nature genetics3169–73
-
[26]
Raser J M and O’Shea E K 2004science3041811–1814
-
[27]
Raser J M and O’shea E K 2005Science3092010–2013
2013
-
[28]
Golding I, Paulsson J, Zawilski S M and Cox E C 2005Cell1231025–1036
-
[29]
Cai L, Friedman N and Xie X S 2006Nature440358–362
-
[30]
Raj A and Van Oudenaarden A 2008Cell135216–226
-
[31]
Cao Z and Grima R 2020Proceedings of the National Academy of Sciences1174682–4692
-
[32]
Wang Y, Szavits-Nossan J, Cao Z and Grima R 2025Physical Review Letters135068401
-
[33]
Del Castillo J and Katz B 1954The Journal of physiology124560
-
[34]
Katz B 1969Liverpool University Press5–39
-
[35]
Hatamie A, He X, Ewing A and Rorsman P 2024ACS Measurement Science Au4629–637
-
[36]
Malagon G, Miki T, Llano I, Neher E and Marty A 2016Journal of Neuroscience36 4010–4025
-
[37]
Silver R A 2003Journal of neuroscience methods130127–141
-
[38]
Rudolph S, Tsai M C, von Gersdorff H and Wadiche J I 2015Trends in neurosciences38 428–438
-
[39]
Zucker R S and Regehr W G 2002Annual review of physiology64355–405
-
[40]
Regehr W G 2012Cold Spring Harbor perspectives in biology4a005702
-
[41]
Pulido C, Trigo F F, Llano I and Marty A 2015Neuron85159–172
-
[42]
Rosenbaum R, Rubin J and Doiron B 2012PLoS computational biology8e1002557
-
[43]
Loebel A, Silberberg G, Helbig D, Markram H, Tsodyks M and Richardson M 2009Frontiers in computational neuroscience3689
-
[44]
Fuhrmann G, Segev I, Markram H and Tsodyks M 2002Journal of neurophysiology87 140–148
-
[45]
Rijal K, M¨ uller N I, Friauf E, Singh A, Prasad A and Das D 2024Physical review letters132 228401
-
[46]
Ali S Y, Prasad A and Das D 2025Physical Review E111014405 14
-
[47]
Gambrell O, Vahdat Z and Singh A 2024 Feedforward regulation of interneuronal communication2024 IEEE 63rd Conference on Decision and Control (CDC)(IEEE) pp 607–612
2024
-
[48]
Gambrell O and Singh A 2025 Analysis of neuronal firing in stochastic models with adaptive thresholds2025 European Control Conference (ECC)(IEEE) pp 582–587
2025
-
[49]
Gambrell O and Singh A 2025 Modulation of interneuronal communication by presynaptic feedback mechanisms2025 IEEE 64th Conference on Decision and Control (CDC)(IEEE) pp 974–979
2025
-
[50]
Zhang C and Peskin C S 2015Proceedings of the National Academy of Sciences112 14954–14959
-
[51]
Zhang C and Peskin C S 2020Communications on Pure and Applied Mathematics733–62
-
[52]
Vahdat Z, Gambrell O, Fisch J, Friauf E and Singh A 2025PLOS Computational Biology21 e1013067
-
[53]
Gambrell O and Singh A 2025 Consequences of decoy site repair on stochastic fluctuations in neurotransmission2025 American Control Conference (ACC)(IEEE) pp 1406–1411
2025
-
[54]
Redner S 2001A guide to first-passage processes(Cambridge university press)
-
[55]
Chou T and D’Orsogna M R 2014 First passage problems in biologyFirst-passage phenomena and their applications(World Scientific) pp 306–345
2014
-
[56]
Bressloff P C 2014Stochastic processes in cell biologyvol 41 (Springer)
-
[57]
Metzler R, Redner S and Oshanin G 2014First-passage phenomena and their applications vol 35 (World Scientific)
-
[58]
B´ enichou O and Voituriez R 2014Physics Reports539225–284
-
[59]
Ghusinga K R, Dennehy J J and Singh A 2017Proceedings of the National Academy of Sciences114693–698
-
[60]
Van Kampen N G 1992Stochastic processes in physics and chemistryvol 1 (Elsevier)
-
[61]
Comtet L 2012Advanced Combinatorics: The art of finite and infinite expansions(Springer Science & Business Media)
-
[62]
Quastel D 1997Biophysical Journal72728–753
-
[63]
Hennig M H 2013Frontiers in computational neuroscience745
-
[64]
Edwards R H 2007Neuron55835–858
-
[65]
Gordleeva S, Dembitskaya Y, Kazantsev V and Postnikov E B 2023Scientific Reports13 15660
-
[66]
Dayan P and Abbott L F 2005Theoretical neuroscience: computational and mathematical modeling of neural systems(MIT press)
-
[67]
B¨ orgers C 2017An introduction to modeling neuronal dynamicsvol 66 (Springer)
-
[68]
Norris J R 1998Markov chains2 (Cambridge university press)
-
[69]
Br´ emont J 2026Physical Review Letters136107101
-
[70]
Friedman N, Cai L and Xie X S 2006Physical review letters97168302
-
[71]
Voits 1, and Ulrich S
Franks K M, Stevens C F and Sejnowski T J 2003Journal of Neuroscience233186–3195 15 Supplemental Material for Activation in Vesicle-Mediated Signaling Shaped by Batch Arrival Statistics Jan Hauke 1, Julian B. Voits 1, and Ulrich S. Schwarz 1,2∗ 1Institute for Theoretical Physics, University of Heidelberg, Germany 2BioQuant-Center for Quantitative Biology,...
2026
-
[72]
Conditional Post-Release Distribution 5
-
[73]
Fixed-Interval Train: Steady State 8 E
Steady-State Post-Release Distribution 7 D. Fixed-Interval Train: Steady State 8 E. Fixed-Interval Train: Normal Approximation 8 II. Extended Model with Vesicle Depletion 9 A. Release Size Distribution 9 B. General Poisson Train: Mean Molecule Count 10
-
[74]
Gamma-Interval Train: Post-Release Distribution 11
Post-Release Mean 11 C. Gamma-Interval Train: Post-Release Distribution 11
-
[75]
Conditional Post-Release Distribution 12
-
[76]
Steady-State Post-Release Distribution 13 III. Sampling of Stochastic Trajectories 16 References 16 ∗ Corresponding author: schwarz@thphys.uni-heidelberg.de 2 In Section I, we derive results for the main model discussed in the paper, where the molecular count is the only stochastic variable and the release size distribution is constant. In Section II, we ...
-
[77]
For the fixed-interval caseα→ ∞, we simply havep −(F) n|n0 = Binom(n|n0, e−1/λ)
Conditional Post-Release Distribution We start out by considering the pre-release distributionp − n|n0 =⟨Binom(n|n 0, e−τ)⟩τ∼f(t) conditioned on last post- release countn 0. For the fixed-interval caseα→ ∞, we simply havep −(F) n|n0 = Binom(n|n0, e−1/λ). For the Poisson train (α= 1), it was shown in the main text thatp −(P) n|n0 = BetaBinom(n|n0, λ,1). Ot...
-
[78]
To be precise, this process is a discrete-time Markov chain, where the state space is the post- release molecule count and each time-step corresponds to a release event
Steady-State Post-Release Distribution The conditional post-release probabilitiesp + n|n0 derived above can be interpreted as transition probabilities of a stochastic process. To be precise, this process is a discrete-time Markov chain, where the state space is the post- release molecule count and each time-step corresponds to a release event. After many ...
-
[79]
The release size distribution is approximately normal
-
[80]
The variance of the release size is much greater than the mean molecule count,⟨m⟩/σ 2 m ≪(1−e −1/λ)
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.