Recognition: no theorem link
Inference of Qualitative Models from Steady-State Data via Weighted MaxSMT
Pith reviewed 2026-05-11 02:11 UTC · model grok-4.3
The pith
A weighted MaxSMT method infers qualitative biological models from steady-state data even when some observations conflict.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
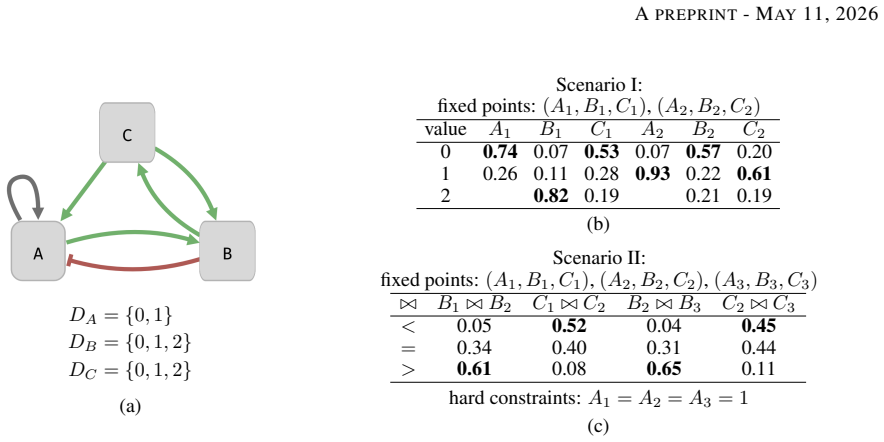
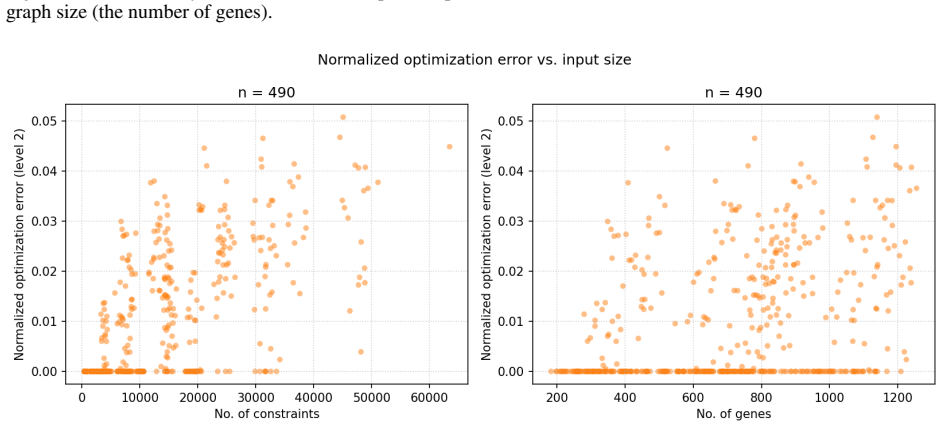
By encoding uncertain biological observations as weighted soft constraints inside a MaxSMT problem, the solver can return a qualitative model that maximises the total weight of satisfied constraints; the same encoding works for both Boolean and multi-valued domains and for both level and ordering constraints, allowing successful inference on prior-knowledge networks of 200–1300 genes describing neural cell differentiation.
What carries the argument
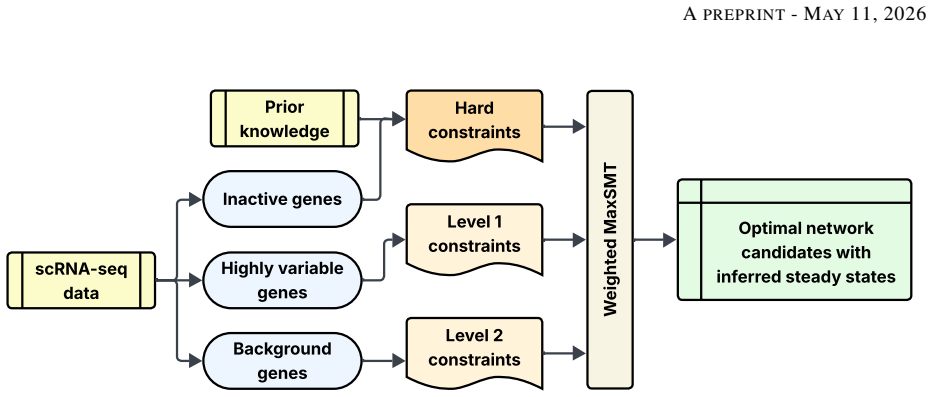
Weighted MaxSMT solver that treats biological observations as soft constraints with explicit weights to select the model maximising satisfied weight.
If this is right
- Large prior-knowledge networks can be used without first discarding conflicting observations.
- Both Boolean and multi-valued qualitative models become feasible under the same inference procedure.
- Ordering constraints derived from differential expression can be combined directly with level constraints.
- Neural cell differentiation networks up to 1300 genes become tractable targets for automated model construction.
Where Pith is reading between the lines
- The weighting scheme could be learned from replicate experiments rather than assigned by hand.
- The same soft-constraint approach may extend to time-series data once suitable encodings for transitions are added.
- Model quality could be validated by checking whether the inferred network reproduces held-out steady-state measurements.
Load-bearing premise
Assigning weights to soft constraints can reliably separate minor measurement errors from fundamental flaws in the model structure.
What would settle it
A synthetic data set in which the true underlying model is known, controlled errors are injected, and the weighted solver returns a model that deviates from the true structure on the most reliable observations.
Figures


read the original abstract
Qualitative models provide crucial instruments for modelling complex biological systems. While advances in automated reasoning and symbolic encodings have enabled rigorous inference of these models from data, the process remains highly fragile. First, biological measurement errors inevitably propagate into formal model specifications. Second, when a specification becomes unsatisfiable, distinguishing between fundamental design flaws and minor technical errors is notoriously difficult. This uncertainty often leads to under-specification, as it is unclear which observations are still ``safe'' to incorporate. To overcome these challenges, we introduce a robust inference method based on weighted MaxSMT. By encoding uncertain biological observations as weighted soft constraints, our approach enables the solver to identify a model best reflecting the observations, even with some conflicting constraints. Our method allows for Boolean and multi-valued variable domains, alongside observations derived from discretisation (level constraints) and differential expression (ordering constraints). We show our approach can be used to successfully infer neural cell differentiation models from prior-knowledge networks with 200--1300 genes using ordering constraints on all included genes.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces a weighted MaxSMT encoding for inferring qualitative (Boolean or multi-valued) models from steady-state biological data. Uncertain observations are represented as weighted soft constraints to tolerate conflicts arising from measurement error, while hard constraints encode prior knowledge; the solver returns a model maximizing the weighted satisfaction. The approach is demonstrated on prior-knowledge networks for neural cell differentiation, successfully inferring models from ordering constraints over all genes in networks of 200–1300 nodes.
Significance. If the weighting scheme can be shown to be data-driven and stable, the method would provide a principled way to scale qualitative model inference to genome-scale networks while explicitly handling the common problem of conflicting observations. This would strengthen automated reasoning tools in systems biology by reducing the need for manual under-specification of data.
major comments (2)
- [Methods (constraint encoding and weight assignment)] The weighting scheme for soft constraints is not specified in sufficient detail (see the description of the MaxSMT encoding). It is unclear whether weights are derived from measurable properties of the data (e.g., replicate variance or discretization confidence) or assigned by trial-and-error; without this, the claim that the solver distinguishes minor technical errors from fundamental model flaws cannot be evaluated and the reported success on 200–1300-gene networks may be sensitive to arbitrary penalty choices.
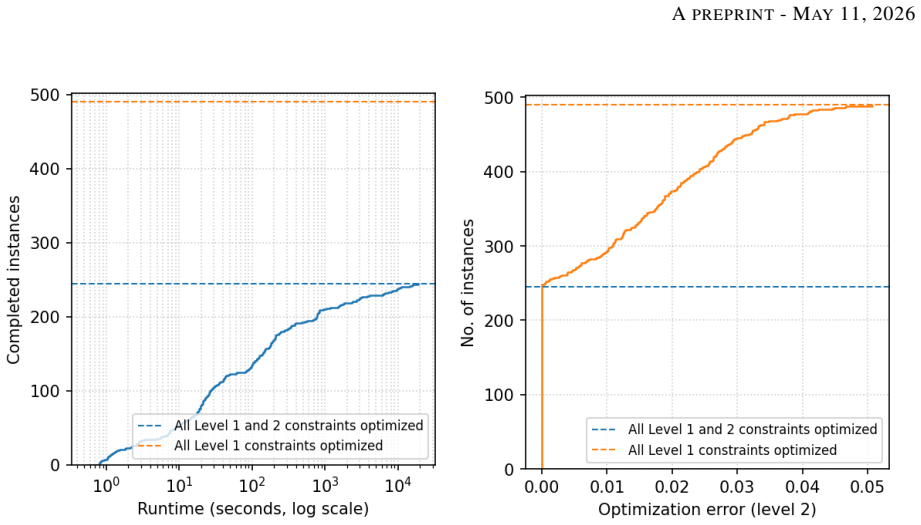
- [Results (large-network experiments)] The validation of inferred models on the neural differentiation networks lacks quantitative metrics, baseline comparisons, or perturbation tests. The abstract states that models were “successfully inferred,” but without reporting how conflicts were resolved, what fraction of constraints were relaxed, or stability under modest weight changes, it is impossible to assess whether the results reflect biological signal or solver artifacts.
minor comments (1)
- Notation for the weighted objective function and the distinction between level constraints and ordering constraints could be made more explicit with a small example table.
Simulated Author's Rebuttal
We thank the referee for these constructive comments, which highlight important areas for clarification and strengthening. We address each point below and indicate the revisions we will make to the manuscript.
read point-by-point responses
-
Referee: The weighting scheme for soft constraints is not specified in sufficient detail (see the description of the MaxSMT encoding). It is unclear whether weights are derived from measurable properties of the data (e.g., replicate variance or discretization confidence) or assigned by trial-and-error; without this, the claim that the solver distinguishes minor technical errors from fundamental model flaws cannot be evaluated and the reported success on 200–1300-gene networks may be sensitive to arbitrary penalty choices.
Authors: We agree that the weighting procedure requires more explicit description. The manuscript states that weights reflect observation reliability (higher for more confident data), but does not detail the assignment rule. In the revision we will expand the Methods section on the MaxSMT encoding to specify that weights are derived from measurable data properties when available (e.g., replicate variance for differential-expression ordering constraints or discretization confidence for level constraints) and will include a concrete example from the neural-differentiation data set showing how these values are computed and normalized. This makes the scheme data-driven and reproducible rather than arbitrary. revision: yes
-
Referee: The validation of inferred models on the neural differentiation networks lacks quantitative metrics, baseline comparisons, or perturbation tests. The abstract states that models were “successfully inferred,” but without reporting how conflicts were resolved, what fraction of constraints were relaxed, or stability under modest weight changes, it is impossible to assess whether the results reflect biological signal or solver artifacts.
Authors: We acknowledge that the current Results section reports only qualitative success on the 200–1300-gene networks. In the revised manuscript we will add quantitative metrics: the fraction of soft constraints satisfied, the number and total weight of relaxed constraints, and a stability analysis under modest (±10 %) weight perturbations for the smaller networks. We will also include a baseline comparison against unweighted MaxSMT (which fails on conflicting data) and report the solver runtime and optimality gap. Full perturbation tests on the largest networks are computationally expensive, but we will provide them for a representative 200-gene subset and discuss scalability limits. revision: partial
Circularity Check
No significant circularity; novel encoding introduced independently of prior fitted results.
full rationale
The paper introduces a weighted MaxSMT encoding to handle uncertain biological observations as soft constraints, allowing inference of qualitative models even with conflicts. The central demonstration—that models can be inferred from 200–1300-gene networks using ordering constraints—is presented as an empirical application of the new method rather than a reduction to self-defined parameters, prior fitted values, or load-bearing self-citations. No equations or steps in the abstract reduce the claimed success to tautological inputs by construction, and the approach remains self-contained against external benchmarks for model inference.
Axiom & Free-Parameter Ledger
free parameters (1)
- weights for soft constraints
axioms (1)
- domain assumption Qualitative models with Boolean or multi-valued domains can adequately represent steady-state biological behavior.
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.