Recognition: 2 theorem links
· Lean TheoremCA-DEL: An Open Multi-Target, Multi-Modal Benchmark for Learning from DNA-Encoded Library Screens
Pith reviewed 2026-05-11 01:45 UTC · model grok-4.3
The pith
CA-DEL supplies a benchmark of noisy DNA-encoded library screens for three carbonic anhydrase isoforms together with real Ki validation data from ChEMBL.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
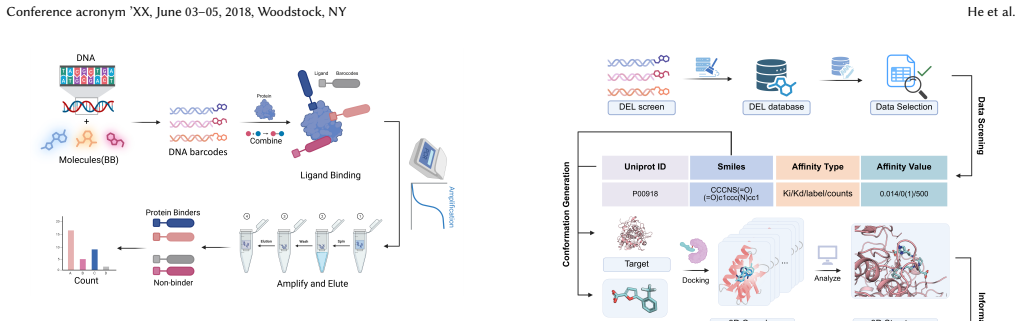
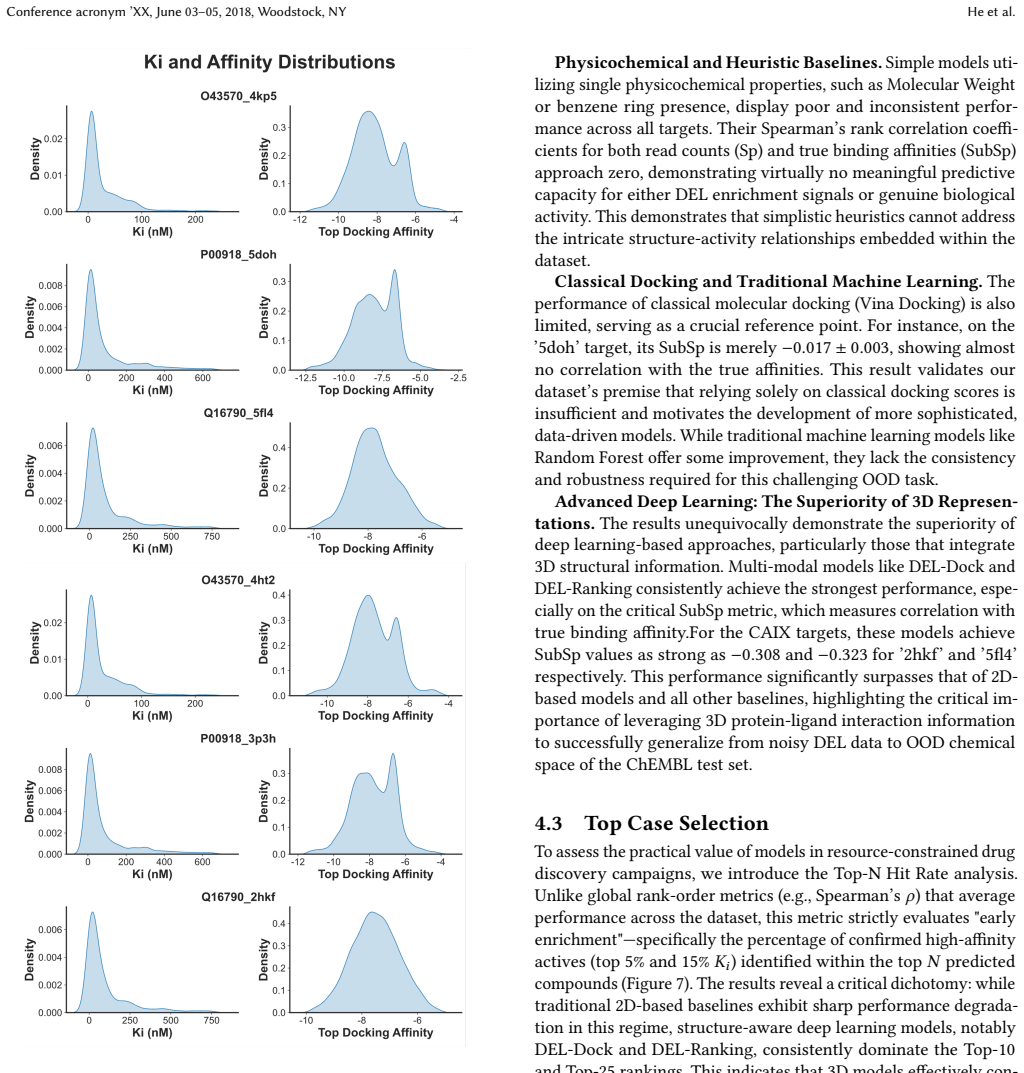
CA-DEL is a multi-target, multi-modal benchmark featuring DEL screens against three homologous carbonic anhydrase isoforms and an integrated validation set of experimentally determined binding affinities (Ki) from ChEMBL, enabling models to be trained on noisy enrichment scores and evaluated on high-fidelity biophysical data.
What carries the argument
The CA-DEL dataset, which pairs sequencing read counts from DEL screens with ChEMBL Ki measurements for the same compounds across three isoforms to support training on indirect signals and testing on direct affinities.
If this is right
- Models can be trained on large-scale noisy DEL data and then tested for whether they predict real binding affinities on the same molecules.
- The benchmark allows direct assessment of whether learned patterns generalize from indirect enrichment scores to measured Ki values.
- Models must learn to distinguish binding preferences among three closely related isoforms rather than generic activity.
- The resource supplies a public, multi-modal starting point for developing methods that handle the noise typical of DEL screens.
Where Pith is reading between the lines
- Similar benchmarks could be built for other protein families to test whether DEL-derived signals transfer to real affinities beyond carbonic anhydrases.
- The sim-to-real split may highlight which molecular features survive the noise and therefore deserve priority in early drug design.
- If models succeed here, the same training-plus-validation pattern could reduce reliance on initial wet-lab affinity measurements for many targets.
Load-bearing premise
The noisy sequencing read counts from DEL screens contain learnable signal about true molecular binding that can generalize to independent ChEMBL Ki measurements for the same compounds.
What would settle it
A model trained only on the CA-DEL DEL read counts that shows no improvement over random or simple baselines when predicting the ChEMBL Ki values for held-out compounds.
Figures





read the original abstract
The success of machine learning in drug discovery hinges on learning the relationship between a chemical structure and its biological activity. While DNA-Encoded Library (DEL) technology can generate the massive datasets required for this task, its primary signal -- sequencing read counts -- is an indirect and often noisy proxy for true molecular binding affinity. To address the scarcity of public benchmarks for developing robust models that can overcome this data challenge, we introduce CA-DEL, a multi-dimensional public benchmark featuring screens against three homologous carbonic anhydrase isoforms. While recent benchmarks like KinDEL have introduced 3D poses for kinase targets, CA-DEL distinguishes itself by focusing on the selectivity challenge among homologous Carbonic Anhydrase isoforms (CAII, CAIX, CAXII). Unlike benchmarks relying solely on noisy enrichment scores, CA-DEL integrates a rigorous validation set of experimentally determined binding affinities ($K_i$) from ChEMBL, establishing a critical Sim-to-Real evaluation paradigm: training on noisy DEL screens and testing on high-fidelity biophysical data.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces CA-DEL, a public multi-target benchmark dataset from DNA-encoded library (DEL) screens against three homologous carbonic anhydrase isoforms (CAII, CAIX, CAXII). It supplies noisy sequencing read counts as the primary DEL signal and pairs a subset of compounds with independent, high-fidelity Ki binding affinities drawn from ChEMBL, thereby creating a Sim-to-Real train/test split for evaluating whether models trained on noisy DEL data can generalize to precise biophysical measurements, with emphasis on selectivity among close homologs.
Significance. If the compound overlap is non-trivial and the DEL read counts contain extractable signal, CA-DEL would supply a rare public resource for studying the transfer from high-throughput noisy screens to low-throughput gold-standard assays. The focus on isoform selectivity among homologous CAs is a timely and practically relevant angle that distinguishes it from kinase-centric benchmarks; the open release of paired multi-modal data could accelerate development of models that are robust to the characteristic noise of DEL technology.
major comments (2)
- [Abstract] Abstract: the central claim that CA-DEL 'establishes a critical Sim-to-Real evaluation paradigm' rests on the existence of paired DEL read counts and ChEMBL Ki values for the same compounds, yet the manuscript supplies no quantitative statistics on the size of this overlap, the distribution of read counts, the dynamic range of Ki values, or any measure of assay concordance between the two modalities. Without these numbers it is impossible to judge whether the benchmark actually supports learnable generalization.
- [Methods / Data description] The description of the DEL screens (presumed §2–3) does not report any characterization of sequencing depth, enrichment-score noise, or false-positive rates typical of DEL data; these details are load-bearing for the claim that the read counts constitute a usable training signal rather than pure noise.
minor comments (2)
- [Abstract] The abstract refers to 'multi-dimensional' and 'multi-modal' aspects without defining the exact modalities or dimensions provided in the released dataset.
- [Data availability] No mention is made of data-release format, license, or accession identifiers that would allow immediate reuse.
Simulated Author's Rebuttal
We thank the referee for their thoughtful review and valuable suggestions. We have carefully considered each comment and revised the manuscript to provide the requested quantitative details and characterizations, thereby strengthening the presentation of the CA-DEL benchmark.
read point-by-point responses
-
Referee: [Abstract] Abstract: the central claim that CA-DEL 'establishes a critical Sim-to-Real evaluation paradigm' rests on the existence of paired DEL read counts and ChEMBL Ki values for the same compounds, yet the manuscript supplies no quantitative statistics on the size of this overlap, the distribution of read counts, the dynamic range of Ki values, or any measure of assay concordance between the two modalities. Without these numbers it is impossible to judge whether the benchmark actually supports learnable generalization.
Authors: We agree that quantitative statistics on the compound overlap, read count distributions, Ki value ranges, and assay concordance are necessary to substantiate the Sim-to-Real evaluation paradigm. The revised manuscript now includes these details in an expanded abstract and a new subsection in the data description, with specific numbers on overlap size, distributions, dynamic ranges, and concordance measures such as correlation coefficients between DEL signals and Ki affinities. revision: yes
-
Referee: [Methods / Data description] The description of the DEL screens (presumed §2–3) does not report any characterization of sequencing depth, enrichment-score noise, or false-positive rates typical of DEL data; these details are load-bearing for the claim that the read counts constitute a usable training signal rather than pure noise.
Authors: We concur that details on sequencing depth, enrichment-score noise, and false-positive rates are critical for demonstrating that the DEL read counts provide a usable training signal. We have revised the Methods section to incorporate these characterizations, including reported sequencing depths, noise estimates for enrichment scores, and false-positive rate analyses based on the experimental setup. revision: yes
Circularity Check
No significant circularity; benchmark data resource without derivations or predictions
full rationale
The paper is a data-benchmark announcement that describes the assembly of paired DEL read-count screens and independent ChEMBL Ki measurements for three CA isoforms. It asserts only that the paired data exist and can support future Sim-to-Real splits; it presents no models, equations, fitted parameters, or predictions whose validity is claimed to follow from the paper's own construction. The central claim therefore does not reduce to any self-definition, fitted-input renaming, or self-citation chain.
Axiom & Free-Parameter Ledger
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel uncleartraining on noisy DEL screens and testing on high-fidelity biophysical data... Spearman’s rank correlation... Top-N hit rate
-
IndisputableMonolith/Foundation/AlphaCoordinateFixation.leanJ_uniquely_calibrated_via_higher_derivative unclearmulti-modal... 3D protein-ligand conformations... docking... enrichment factors
Reference graph
Works this paper leans on
-
[1]
Vincenzo Alterio, Mika Hilvo, Anna Di Fiore, Claudiu T Supuran, Peiwen Pan, Seppo Parkkila, Andrea Scaloni, Jaromir Pastorek, Silvia Pastorekova, Carlo Pedone, et al . 2009. Crystal structure of the catalytic domain of the tumor- associated human carbonic anhydrase IX.Proceedings of the National Academy of Sciences106, 38 (2009), 16233–16238
work page 2009
-
[2]
Pedro J Ballester and John BO Mitchell. 2010. A machine learning approach to predicting protein–ligand binding affinity with applications to molecular docking. Bioinformatics26, 9 (2010), 1169–1175
work page 2010
-
[3]
G Richard Bickerton, Gaia V Paolini, Jérémy Besnard, Sorel Muresan, and An- drew L Hopkins. 2012. Quantifying the chemical beauty of drugs.Nature chemistry4, 2 (2012), 90–98
work page 2012
-
[4]
Sydney Brenner and Richard A Lerner. 1992. Encoded combinatorial chemistry. Proceedings of the National Academy of Sciences89, 12 (1992), 5381–5383
work page 1992
- [5]
-
[6]
Benson Chen, Tomasz Danel, Gabriel HS Dreiman, Patrick J McEnaney, Nikhil Jain, Kirill Novikov, Spurti Umesh Akki, Joshua L Turnbull, Virja Atul Pandya, Boris P Belotserkovskii, et al. 2024. KinDEL: DNA-encoded library dataset for kinase inhibitors.arXiv preprint arXiv:2410.08938(2024)
-
[7]
Christoph E Dumelin, Jörg Scheuermann, Samu Melkko, and Dario Neri. 2006. Selection of streptavidin binders from a DNA-encoded chemical library.Biocon- jugate chemistry17, 2 (2006), 366–370
work page 2006
-
[8]
John C Faver, Kevin Riehle, David R Lancia Jr, Jared BJ Milbank, Christopher S Kollmann, Nicholas Simmons, Zhifeng Yu, and Martin M Matzuk. 2019. Quanti- tative comparison of enrichment from DNA-encoded chemical library selections. ACS combinatorial science21, 2 (2019), 75–82
work page 2019
-
[9]
Philippe Ferrara, Holger Gohlke, Daniel J Price, Gerhard Klebe, and Charles L Brooks. 2004. Assessing scoring functions for protein- ligand interactions.Journal of medicinal chemistry47, 12 (2004), 3032–3047
work page 2004
-
[10]
Anna Gaulton, Louisa J Bellis, A Patricia Bento, Jon Chambers, Mark Davies, Anne Hersey, Yvonne Light, Shaun McGlinchey, David Michalovich, Bissan Al-Lazikani, et al. 2012. ChEMBL: a large-scale bioactivity database for drug discovery.Nucleic acids research40, D1 (2012), D1100–D1107
work page 2012
-
[11]
Caroli Genis, Katherine H Sippel, Nicolette Case, Wengang Cao, Balendu Sankara Avvaru, Lawrence J Tartaglia, Lakshmanan Govindasamy, Chingkuang Tu, Mavis Agbandje-McKenna, David N Silverman, et al. 2009. Design of a carbonic anhy- drase IX active-site mimic to screen inhibitors for possible anticancer properties. Biochemistry48, 6 (2009), 1322–1331
work page 2009
-
[12]
Christopher J Gerry, Mathias J Wawer, Paul A Clemons, and Stuart L Schreiber
-
[13]
Journal of the American Chemical Society141, 26 (2019), 10225–10235
DNA barcoding a complete matrix of stereoisomeric small molecules. Journal of the American Chemical Society141, 26 (2019), 10225–10235
work page 2019
- [14]
-
[15]
Rui Hou, Chao Xie, Yuhan Gui, Gang Li, and Xiaoyu Li. 2023. Machine-learning- based data analysis method for cell-based selection of DNA-encoded libraries. ACS omega8, 21 (2023), 19057–19071
work page 2023
-
[16]
Sumaiya Iqbal, Wei Jiang, Eric Hansen, Tonia Aristotelous, Shuang Liu, Andrew Reidenbach, Cerise Raffier, Alison Leed, Chengkuan Chen, Lawrence Chung, et al. 2025. Evaluation of DNA encoded library and machine learning model combinations for hit discovery.npj Drug Discovery2, 1 (2025), 5
work page 2025
-
[17]
David Ryan Koes, Matthew P Baumgartner, and Carlos J Camacho. 2013. Lessons learned in empirical scoring with smina from the CSAR 2011 benchmarking exercise.Journal of chemical information and modeling53, 8 (2013), 1893–1904
work page 2013
-
[18]
Péter Kómár and Marko Kalinic. 2020. Denoising DNA encoded library screens with sparse learning.ACS Combinatorial Science22, 8 (2020), 410–421
work page 2020
-
[19]
Letian Kuai, Thomas O’Keeffe, and Christopher Arico-Muendel. 2018. Random- ness in DNA encoded library selection data can be modeled for more reliable enrichment calculation.SLAS DISCOVERY: Advancing the Science of Drug Discov- ery23, 5 (2018), 405–416
work page 2018
-
[20]
H Li, CW Yap, CY Ung, Y Xue, ZR Li, LY Han, HH Lin, and Yu Zong Chen
-
[21]
Machine learning approaches for predicting compounds that interact with therapeutic and ADMET related proteins.Journal of pharmaceutical sciences96, 11 (2007), 2838–2860
work page 2007
-
[22]
Katherine S Lim, Andrew G Reidenbach, Bruce K Hua, Jeremy W Mason, Christo- pher J Gerry, Paul A Clemons, and Connor W Coley. 2022. Machine learning on DNA-encoded library count data using an uncertainty-aware probabilistic loss function.Journal of chemical information and modeling62, 10 (2022), 2316–2331
work page 2022
- [23]
-
[24]
Ralph Ma, Gabriel HS Dreiman, Fiorella Ruggiu, Adam Joseph Riesselman, Bowen Liu, Keith James, Mohammad Sultan, and Daphne Koller. 2021. Regression modeling on DNA encoded libraries. InNeurIPS 2021 AI for Science Workshop
work page 2021
-
[25]
Mam Y Mboge, Zhijuan Chen, Alyssa Wolff, John V Mathias, Chingkuang Tu, Kevin D Brown, Murat Bozdag, Fabrizio Carta, Claudiu T Supuran, Robert McKenna, et al. 2018. Selective inhibition of carbonic anhydrase IX over carbonic anhydrase XII in breast cancer cells using benzene sulfonamides: Disconnect between activity and growth inhibition.PloS one13, 11 (2...
work page 2018
-
[26]
Kevin McCloskey, Eric A Sigel, Steven Kearnes, Ling Xue, Xia Tian, Dennis Moccia, Diana Gikunju, Sana Bazzaz, Betty Chan, Matthew A Clark, et al. 2020. Machine learning on DNA-encoded libraries: a new paradigm for hit finding. Journal of Medicinal Chemistry63, 16 (2020), 8857–8866
work page 2020
-
[27]
Alba L Montoya, Adam S Hogendorf, Steven Tingey, Aadarsh Kuberan, Lik Hang Yuen, Herwig Schüler, and Raphael M Franzini. 2025. Widespread false negatives in DNA-encoded library data: how linker effects impair machine learning-based lead prediction.Chemical Science(2025)
work page 2025
-
[28]
Sudipto Mukherjee, Trent E Balius, and Robert C Rizzo. 2010. Docking validation resources: protein family and ligand flexibility experiments.Journal of chemical information and modeling50, 11 (2010), 1986–2000
work page 2010
-
[29]
Michael C Needels, David G Jones, Emily H Tate, Gregory L Heinkel, Lynn M Kochersperger, William J Dower, Ronald W Barrett, and Mark A Gallop. 1993. Generation and screening of an oligonucleotide-encoded synthetic peptide library. Proceedings of the National Academy of Sciences90, 22 (1993), 10700–10704
work page 1993
-
[30]
Ian K Quigley, Andrew Blevins, Brayden J Halverson, and Nate Wilkinson. 2024. Belka: The big encoded library for chemical assessment. InNeurIPS 2024 Compe- tition Track
work page 2024
-
[31]
Alexander L Satz. 2016. Simulated screens of DNA encoded libraries: the potential influence of chemical synthesis fidelity on interpretation of structure–activity relationships.ACS combinatorial science18, 7 (2016), 415–424
work page 2016
-
[32]
Valeria Scardino, Mariela Bollini, and Claudio N Cavasotto. 2021. Combination of pose and rank consensus in docking-based virtual screening: the best of both worlds.RSC advances11, 56 (2021), 35383–35391
work page 2021
-
[33]
Sara Shamsian, Babak Sokouti, and Siavoush Dastmalchi. 2023. Benchmarking different docking protocols for predicting the binding poses of ligands complexed with cyclooxygenase enzymes and screening chemical libraries.BioImpacts: BI 14, 2 (2023), 29955
work page 2023
-
[34]
Yu Shi, Wei Xu, and Pingzhao Hu. 2025. Out of distribution learning in bioin- formatics: advancements and challenges.Briefings in Bioinformatics26, 3 (2025), bbaf294
work page 2025
-
[35]
Kirill Shmilovich, Benson Chen, Theofanis Karaletsos, and Mohammad M Sultan
-
[36]
Journal of Chemical Information and Modeling63, 9 (2023), 2719–2727
Del-dock: Molecular docking-enabled modeling of dna-encoded libraries. Journal of Chemical Information and Modeling63, 9 (2023), 2719–2727
work page 2023
-
[37]
Michael D Shultz. 2018. Two decades under the influence of the rule of five and the changing properties of approved oral drugs: miniperspective.Journal of Medicinal Chemistry62, 4 (2018), 1701–1714
work page 2018
-
[38]
Prudencio Tossou, Cas Wognum, Michael Craig, Hadrien Mary, and Emmanuel Noutahi. 2024. Real-world molecular out-of-distribution: Specification and inves- tigation.Journal of Chemical Information and Modeling64, 3 (2024), 697–711
work page 2024
-
[39]
Lingle Wang, Yujie Wu, Yuqing Deng, Byungchan Kim, Levi Pierce, Goran Krilov, Dmitry Lupyan, Shaughnessy Robinson, Markus K Dahlgren, Jeremy Greenwood, et al. 2015. Accurate and reliable prediction of relative ligand binding potency in prospective drug discovery by way of a modern free-energy calculation protocol and force field.Journal of the American Ch...
work page 2015
-
[40]
Renxiao Wang, Yipin Lu, and Shaomeng Wang. 2003. Comparative evaluation of 11 scoring functions for molecular docking.Journal of medicinal chemistry46, 12 (2003), 2287–2303
work page 2003
-
[41]
Moreno Wichert, Laura Guasch, and Raphael M Franzini. 2024. Challenges and Prospects of DNA-Encoded Library Data Interpretation.Chemical Reviews124, 22 (2024), 12551–12572. CA-DEL: An Open Multi-Target, Multi-Modal Benchmark for Learning from DNA-Encoded Library Screens Conference acronym ’XX, June 03–05, 2018, Woodstock, NY A Appendix A.1 Appendix figure...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.