Recognition: 2 theorem links
· Lean TheoremCellwise and Casewise Robust Multivariate Regression with Inference
Pith reviewed 2026-05-11 02:44 UTC · model grok-4.3
The pith
A new estimator enables robust multivariate regression that handles both whole-observation and single-cell outliers along with missing values.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The cellMR estimator simultaneously accommodates casewise and cellwise outliers, missing data, and high dimensionality in multivariate linear regression by building on a cellwise robust covariance estimator and using ridge regularization. The cellBoot procedure, based on indirect inference, provides asymptotically valid confidence intervals robust to both types of contamination, with derived influence functions supporting this.
What carries the argument
The cellwise multivariate regression (cellMR) estimator, which combines a cellwise robust covariance estimator with ridge regularization to produce regression coefficients that resist mixed outlier patterns and missing entries.
If this is right
- The estimator produces stable coefficients even when the number of variables approaches or exceeds the number of observations.
- cellBoot confidence intervals remain valid under simultaneous casewise and cellwise contamination.
- The procedure works directly on data matrices that contain missing values without requiring separate imputation.
- Influence functions quantify the effect of individual contaminated cells or rows on the fitted coefficients.
- Real-data examples such as genomics applications show competitive finite-sample accuracy compared with classical methods.
Where Pith is reading between the lines
- Similar cellwise-robust covariance building blocks could be inserted into other multivariate techniques such as principal-component analysis or canonical correlation.
- The framework suggests a path toward robust versions of regularized regression that also deliver valid inference without cross-validation tuning.
- In practice this would let analysts keep more observations instead of listwise deletion, potentially increasing power in studies with incomplete records.
- Extensions to time-series or spatial data might follow by adapting the cellwise contamination model to respect dependence structure.
Load-bearing premise
The cellwise robust covariance estimator must perform reliably under the paper's contamination model and the indirect-inference bootstrap must be correctly calibrated for the asymptotic validity proofs to go through.
What would settle it
Repeated simulations in which the cellBoot intervals achieve coverage well below the nominal level when 5-10 percent of cells are contaminated and some entries are missing would falsify the asymptotic-validity claim.
Figures










read the original abstract
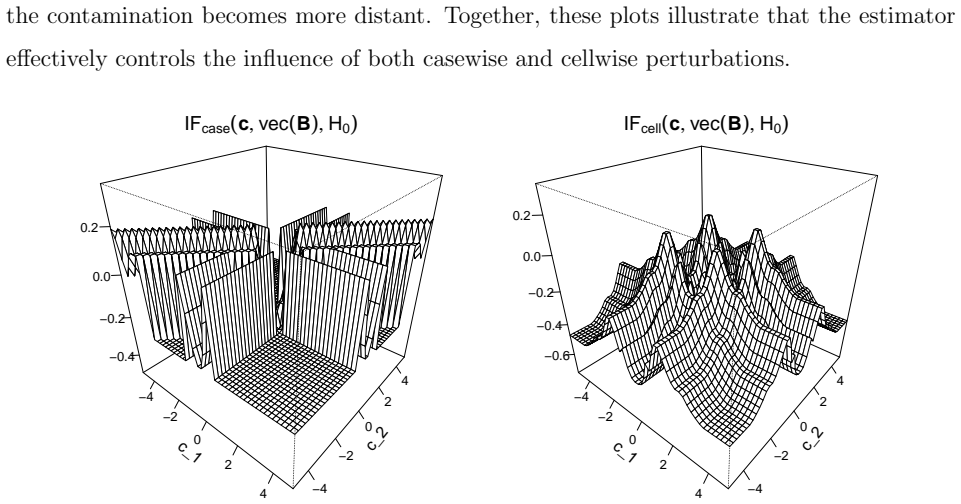
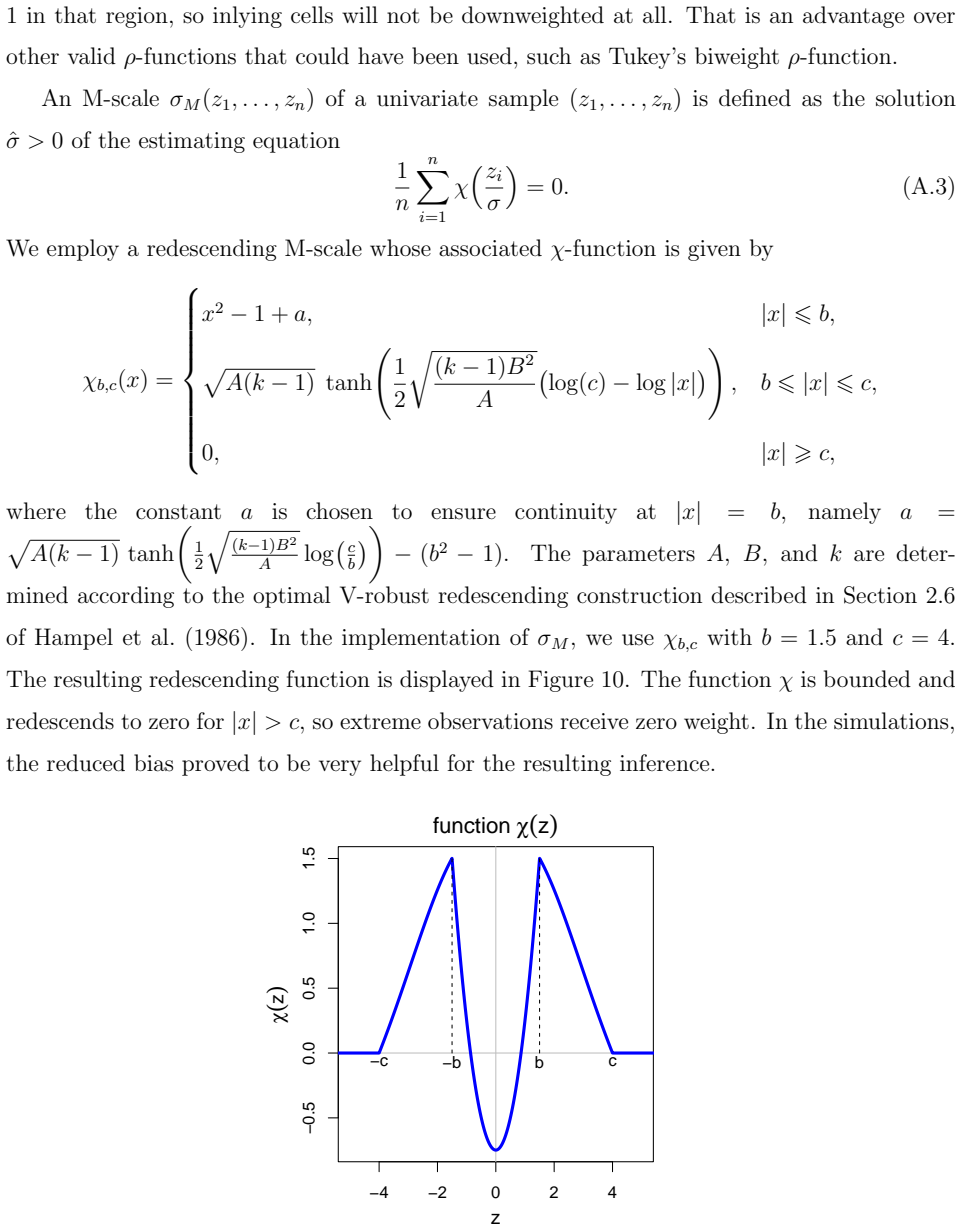
Multivariate linear regression is a fundamental statistical task, but classical estimators such as ordinary least squares are highly sensitive to outliers. These may occur as casewise outliers that affect entire observations, or as outlying cells, that are individual contaminated entries in the predictor and/or response matrix. Moreover, modern datasets frequently contain missing values and are high-dimensional. To address these challenges we propose the cellwise multivariate regression (cellMR) estimator, a robust regression method that simultaneously accommodates casewise and cellwise outliers, missing data, and high dimensionality. The approach builds on a cellwise robust covariance estimator and uses ridge regularization for numerical stability. We further introduce cellBoot, a novel bootstrap-based inference procedure tailored to the cellMR framework. Relying on indirect inference, cellBoot provides asymptotically valid confidence intervals that are robust to casewise and cellwise contamination. We derive influence functions of the regression estimator and prove the asymptotic validity of the cellBoot confidence intervals. Simulations and a real genomics application illustrate the strong finite-sample performance of the proposed methods.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes the cellMR estimator for robust multivariate linear regression that simultaneously handles casewise and cellwise outliers, missing data, and high dimensionality by building on a cellwise robust covariance estimator with ridge regularization. It introduces cellBoot, an indirect-inference bootstrap procedure for asymptotically valid confidence intervals robust to contamination. The authors derive influence functions of the regression estimator and prove the asymptotic validity of the cellBoot CIs, with supporting simulations and a real genomics application.
Significance. If the theoretical claims hold, this would be a useful contribution to robust multivariate methods by extending cellwise robust covariance ideas to regression with mixed contamination, missingness, and high dimensions while providing inference. The simulations and genomics application provide concrete evidence of finite-sample behavior and practical utility.
major comments (1)
- [theoretical results on asymptotic validity and influence functions] The central claim of deriving influence functions and proving asymptotic validity of cellBoot (as stated in the abstract) requires the underlying cellwise robust covariance estimator to satisfy uniform consistency, bounded eigenvalues, and appropriate rates for contamination fraction and p/n even after ridge stabilization and missing-data handling. The manuscript does not explicitly state or verify these regularity conditions in the theoretical development, which is load-bearing for the bootstrap calibration to remain valid in the p ≫ n regime.
minor comments (1)
- The abstract refers to 'strong finite-sample performance' without specifying the exact performance metrics (e.g., bias, coverage rates) or contamination levels used in the simulations; adding this detail would improve clarity.
Simulated Author's Rebuttal
We appreciate the referee's thorough review and valuable feedback on our manuscript. We address the major comment regarding the theoretical results below. We believe the revisions will clarify the assumptions and strengthen the presentation of our theoretical contributions.
read point-by-point responses
-
Referee: [theoretical results on asymptotic validity and influence functions] The central claim of deriving influence functions and proving asymptotic validity of cellBoot (as stated in the abstract) requires the underlying cellwise robust covariance estimator to satisfy uniform consistency, bounded eigenvalues, and appropriate rates for contamination fraction and p/n even after ridge stabilization and missing-data handling. The manuscript does not explicitly state or verify these regularity conditions in the theoretical development, which is load-bearing for the bootstrap calibration to remain valid in the p ≫ n regime.
Authors: We thank the referee for highlighting this important point. The regularity conditions on the cellwise robust covariance estimator are indeed crucial for the validity of the influence function derivations and the asymptotic results for cellBoot, particularly in high-dimensional settings with ridge regularization and missing data. While some of these conditions are implicitly assumed through the properties of the base estimator (as referenced in the literature on cellwise robust methods), we agree that they should be stated explicitly to ensure the theoretical development is self-contained and transparent. In the revised manuscript, we will introduce a new subsection that explicitly lists the required regularity conditions, including uniform consistency of the covariance estimator, bounded eigenvalues (accounting for ridge stabilization), and the allowable rates for the contamination fraction and p/n ratio. We will also discuss how these are maintained under the missing-data handling procedure. This addition will not change the main results but will provide the necessary foundation for the claims. revision: yes
Circularity Check
No significant circularity in derivation chain
full rationale
The paper introduces a new cellMR estimator built on an existing cellwise robust covariance approach plus ridge regularization, along with a new cellBoot procedure using indirect inference. It derives influence functions and proves asymptotic validity of the confidence intervals. No equations or steps in the abstract or described structure reduce the central claims to fitted parameters by construction, self-definitional loops, or load-bearing self-citations that collapse the result. The proofs rely on regularity conditions for the covariance estimator under contamination, but these are external assumptions rather than internal reductions. The derivation remains self-contained with independent content from the new methods and proofs.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Standard regularity conditions for consistency and asymptotic normality of robust M-estimators
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearWe derive influence functions of the regression estimator and prove the asymptotic validity of the cellBoot confidence intervals.
-
IndisputableMonolith/Foundation/RealityFromDistinction.leanreality_from_one_distinction unclearcellMR builds on a cellwise robust covariance estimator and uses ridge regularization
Reference graph
Works this paper leans on
- [1]
-
[2]
Alfons, A., C. Croux, and S. Gelper (2013). Sparse least trimmed squares regression for analyzing high-dimensional large data sets . The Annals of Applied Statistics\/ 7\/ (1), 226 -- 248
work page 2013
-
[3]
Alqallaf, F., S. Van Aelst, V. J. Yohai, and R. H. Zamar (2009). Propagation of outliers in multivariate data. The Annals of Statistics\/ 37 , 311--331
work page 2009
-
[4]
Amado, C. and A. M. Pires (2004). Robust bootstrap with non random weights based on the influence function. Communications in Statistics-Simulation and Computation\/ 33\/ (2), 377--396
work page 2004
-
[5]
Bickel, P. J. and D. A. Freedman (1981). Some asymptotic theory for the bootstrap. The Annals of Statistics\/ 9\/ (6), 1196--1217
work page 1981
-
[6]
Bottmer, L., C. Croux, and I. Wilms (2022). Sparse regression for large data sets with outliers. European Journal of Operational Research\/ 297\/ (2), 782--794
work page 2022
-
[7]
Centofanti, F., M. Hubert, and P. J. Rousseeuw (2025). Cellwise and Casewise Robust Covariance in High Dimensions, arXiv preprint arXiv:2505.19925
-
[8]
Centofanti, F., M. Hubert, and P. J. Rousseeuw (2026). Robust Principal Components by Casewise and Cellwise Weighting . Technometrics, to appear \/ \!\! , \; https://doi.org/10.1080/00401706.2026.2643216\,
-
[9]
Cohen Freue, G. V., D. Kepplinger, M. Salibi \'a n-Barrera, and E. Smucler (2019). Robust elastic net estimators for variable selection and identification of proteomic biomarkers. The Annals of Applied Statistics\/ 13\/ (4), 2065--2090
work page 2019
-
[10]
Efron, B. and R. J. Tibshirani (1994). An Introduction to the Bootstrap . CRC press
work page 1994
-
[11]
Filzmoser, P., S. H \"o ppner, I. Ortner, S. Serneels, and T. Verdonck (2020). Cellwise robust M regression. Computational Statistics & Data Analysis\/ 147 , 106944
work page 2020
-
[12]
Filzmoser, P. and K. Nordhausen (2021). Robust linear regression for high-dimensional data: An overview. Wiley Interdisciplinary Reviews: Computational Statistics\/ 13\/ (4), e1524
work page 2021
-
[13]
Gourieroux, C., A. Monfort, and E. Renault (1993). Indirect inference. Journal of Applied Econometrics\/ 8\/ (S1), S85--S118
work page 1993
-
[14]
Guerrier, S., E. Dupuis-Lozeron , Y. Ma, and M.-P. Victoria-Feser (2019). Simulation-based bias correction methods for complex models. Journal of the American Statistical Association\/ 114 , 146--157
work page 2019
-
[15]
Hampel, F. R., E. M. Ronchetti, and P. J. Rousseeuw (1981). The Change-of-Variance Curve and Optimal Redescending M-Estimators . Journal of the American Statistical Association\/ 76 , 643--648
work page 1981
-
[16]
Hampel, F. R., E. M. Ronchetti, P. J. Rousseeuw, and W. A. Stahel (1986). Robust Statistics: the Approach based on Influence Functions . Wiley
work page 1986
-
[17]
Hastie, T., R. Tibshirani, and J. Friedman (2009). The Elements of Statistical Learning: Data Mining, Inference, and Prediction . Springer Series in Statistics. Springer
work page 2009
-
[18]
Huber, P. J. (1964). Robust estimation of a location parameter. The Annals of Mathematical Statistics\/ 35\/ (1), 73--101
work page 1964
-
[19]
Huber, P. J. (1981). Robust Statistics . John Wiley & Sons
work page 1981
-
[20]
Hubert, M., P. J. Rousseeuw, and T. Verdonck (2012). A deterministic algorithm for robust location and scatter. Journal of Computational and Graphical Statistics\/ 21\/ (3), 618--637
work page 2012
-
[21]
Kosorok, M. R. (2008). Introduction to Empirical Processes and Semiparametric Inference . Springer
work page 2008
-
[22]
Leung, A., H. Zhang, and R. Zamar (2016). Robust regression estimation and inference in the presence of cellwise and casewise contamination. Computational Statistics & Data Analysis\/ 99 , 1--11
work page 2016
-
[23]
Little, R. J. (1992). Regression with missing x's: a review. Journal of the American Statistical Association\/ 87\/ (420), 1227--1237
work page 1992
-
[24]
Maronna, R. A. (2011). Robust ridge regression for high-dimensional data. Technometrics\/ 53\/ (1), 44--53
work page 2011
-
[25]
Maronna, R. A., R. D. Martin, V. J. Yohai, and M. Salibi \'a n-Barrera (2019). Robust Statistics: T heory and Methods (with R) . John Wiley & Sons
work page 2019
-
[26]
Newey, W. K. and D. McFadden (1994). Large sample estimation and hypothesis testing. Handbook of Econometrics\/ 4 , 2111--2245
work page 1994
-
[27]
\"O llerer, V. and C. Croux (2015). Robust high-dimensional precision matrix estimation. In Modern Nonparametric, Robust and Multivariate Methods , pp.\ 325--350. Springer
work page 2015
-
[28]
Raymaekers, J. and P. J. Rousseeuw (2021). Fast robust correlation for high-dimensional data. Technometrics\/ 63 , 184--198
work page 2021
-
[29]
Raymaekers, J. and P. J. Rousseeuw (2026). Challenges of cellwise outliers. Econometrics and Statistics\/ 38 , 6--25, DOI https://doi.org/10.1016/j.ecosta.2024.02.002
-
[30]
Rousseeuw, P. J. (1984). Least median of squares regression. Journal of the American Statistical Association\/ 79\/ (388), 871--880
work page 1984
-
[31]
Rousseeuw, P. J. and A. Leroy (1987). Robust R egression and O utlier D etection . Wiley
work page 1987
-
[32]
Rousseeuw, P. J. and W. Van den Bossche (2018). Detecting deviating data cells. Technometrics\/ 60\/ (2), 135--145
work page 2018
-
[33]
Salibi \'a n-Barrera, M., S. Van Aelst, and G. Willems (2008). Fast and robust bootstrap. Statistical Methods and Applications\/ 17\/ (1), 41--71
work page 2008
-
[34]
Salibian-Barrera, M. and R. H. Zamar (2002). Bootstrapping robust estimates of regression. The Annals of Statistics\/ 30 , 556--582
work page 2002
-
[35]
Shankavaram, U. T., W. C. Reinhold, S. Nishizuka, S. Major, D. Morita, K. K. Chary, M. A. Reimers, U. Scherf, A. Kahn, D. Dolginow, et al. (2007). Transcript and protein expression profiles of the NCI-60 cancer cell panel: an integromic microarray study. Molecular Cancer Therapeutics\/ 6\/ (3), 820--832
work page 2007
-
[36]
Su, P., G. Tarr, S. Muller, and S. Wang (2024). CR-Lasso: Robust cellwise regularized sparse regression . Computational Statistics & Data Analysis\/ 197\/ (107971), 1--14
work page 2024
-
[37]
Van Aelst, S. and G. Willems (2005). Multivariate regression S -estimators for robust estimation and inference. Statistica Sinica\/ 15 , 981--1001
work page 2005
-
[38]
Van der Vaart, A. W. (2000). Asymptotic Statistics . Cambridge University Press
work page 2000
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.