Recognition: 2 theorem links
· Lean TheoremQuantum Injection Pathways for Implicit Graph Neural Networks
Pith reviewed 2026-05-12 02:27 UTC · model grok-4.3
The pith
Independent injection of quantum signals into graph deep equilibrium models yields superior accuracy and efficiency on classification benchmarks.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
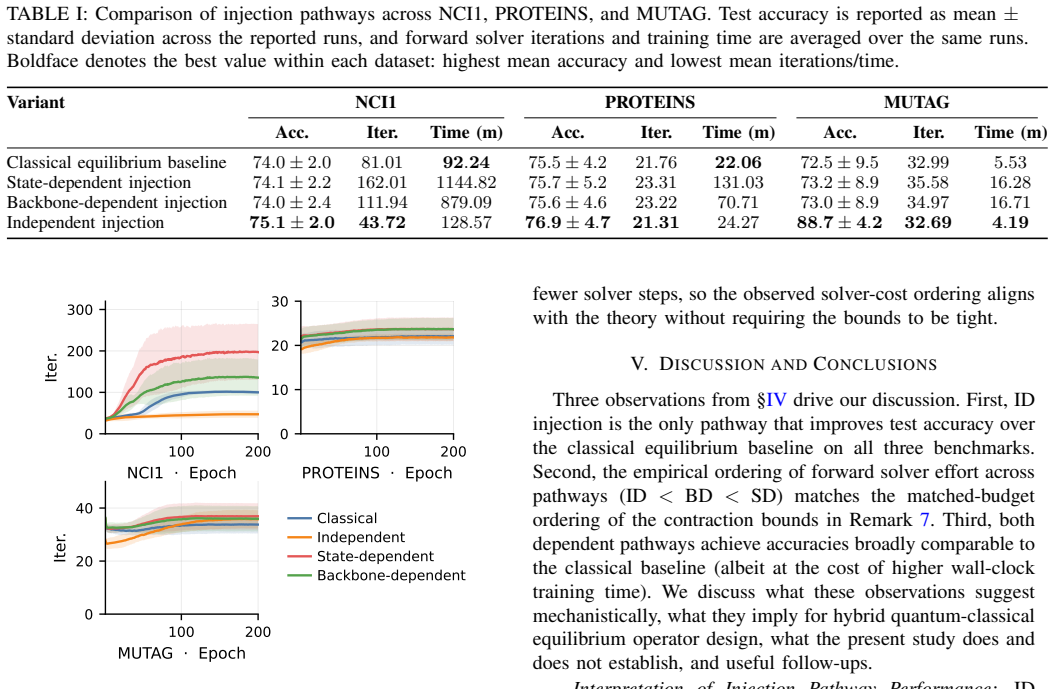
The paper establishes three quantum injection pathways for graph DEQs—independent, state-dependent, and backbone-dependent—and demonstrates through theory and experiment that independent injection, which computes the quantum signal once per graph and holds it fixed during the solve, provides the highest test accuracy on standard benchmarks while requiring fewer forward-solver iterations.
What carries the argument
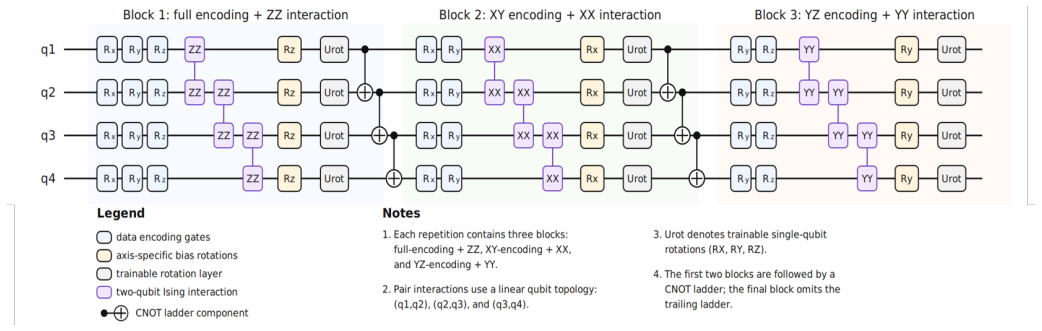
The fixed-point operator of the graph deep equilibrium model, modified by one of three quantum signal injection strategies that differ in the timing and target of the quantum computation within the iterative solve.
If this is right
- Independent injection requires fewer iterations to reach the fixed point compared to both the classical DEQ and the state- and backbone-dependent quantum injections.
- All variants admit unique fixed points when the Lipschitz constants satisfy the contraction conditions derived in the paper.
- The quantum DEQ framework achieves the expressive power of deep networks at the cost of a single layer's memory.
- Quantum signals can be coupled to implicit graph models without requiring repeated quantum evaluations at every solver step.
Where Pith is reading between the lines
- The independent injection method could be particularly advantageous on quantum hardware where each quantum computation is costly, as it minimizes the number of quantum calls.
- The contraction mapping results provide a theoretical foundation that may extend to other implicit models incorporating quantum components.
- Testable extensions include applying these injection pathways to larger graph datasets or different quantum signal encodings.
Load-bearing premise
The contraction guarantees and thus the well-posedness of the fixed-point equations depend on the classical backbone and the quantum signal having sufficiently small Lipschitz constants.
What would settle it
Observing that independent injection fails to achieve the highest accuracy or uses more iterations than the baselines on the NCI1, PROTEINS, or MUTAG benchmarks would directly challenge the main empirical claim.
Figures

read the original abstract
Deep Equilibrium Models (DEQs) replace a stack of explicit layers with a single operator whose fixed point defines the output, giving the expressive power of an arbitrarily deep network at the memory cost of a single layer. Quantum Deep Equilibrium Models (QDEQs) bring this idea to quantum machine learning, offering an alternative to Parameterized Quantum Circuits (PQCs), whose depth is limited by hardware coherence and trainability. Here, we introduce, formulate, and compare three ways of coupling a quantum signal to graph DEQs, differing in where the signal enters the fixed-point operator. \textit{Independent} injection computes the quantum signal once per graph and forward fixed-point solve, and holds it fixed throughout the solve. \textit{State-dependent} injection instead recomputes the signal at every solver step and applies it to the current iterate. \textit{Backbone-dependent} injection likewise recomputes at every iteration but applies the signal to the classical backbone's output evaluated at the current iterate. We establish contraction guarantees for each variant under explicit assumptions on the Lipschitz constants of the classical backbone and the quantum signal. On the TU Dortmund graph-classification benchmarks NCI1, PROTEINS, and MUTAG, independent injection achieves the best test accuracy while using fewer forward-solver iterations than both the classical equilibrium baseline and the two dependent variants.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces three quantum injection pathways—independent, state-dependent, and backbone-dependent—for coupling quantum signals to graph Deep Equilibrium Models (DEQs). It derives contraction guarantees for the fixed-point iterations of each variant under assumptions on the Lipschitz constants of the classical backbone and quantum signal. Empirically, on the NCI1, PROTEINS, and MUTAG graph classification tasks, the independent injection variant is reported to achieve the highest test accuracy while requiring fewer solver iterations than the classical DEQ baseline and the dependent injection methods.
Significance. If the contraction guarantees hold under the stated assumptions and the empirical superiority is robust, this work offers a memory-efficient hybrid quantum-classical approach to graph neural networks that sidesteps the depth limitations of parameterized quantum circuits. The explicit derivation of contraction conditions for each injection variant and the head-to-head comparison on standard TU Dortmund benchmarks constitute clear strengths that could guide future design of implicit QML models.
major comments (1)
- [Theoretical analysis of contraction guarantees] The contraction guarantees (stated in the abstract as holding 'under explicit assumptions' on the Lipschitz constants of the classical backbone and quantum signal) are load-bearing for the central claim that independent injection uses fewer forward-solver iterations. The manuscript provides no evidence that these Lipschitz bounds were measured, enforced, or satisfied by the trained models evaluated on NCI1, PROTEINS, and MUTAG. If the quantum-signal Lipschitz constant exceeds the threshold required for a contraction factor <1, the reported iteration advantage disappears and the comparison to the classical DEQ baseline is no longer valid.
minor comments (2)
- [Experimental evaluation] The experimental section should report standard deviations or error bars on test accuracies and specify the precise train/validation/test splits and random seeds used for the NCI1, PROTEINS, and MUTAG benchmarks.
- Implementation details for the quantum signal (circuit architecture, embedding into the graph DEQ, and how the signal is computed in each injection variant) should be expanded to enable reproduction.
Simulated Author's Rebuttal
We thank the referee for their careful reading and constructive feedback on our manuscript. We address the major comment regarding the contraction guarantees below.
read point-by-point responses
-
Referee: The contraction guarantees (stated in the abstract as holding 'under explicit assumptions' on the Lipschitz constants of the classical backbone and quantum signal) are load-bearing for the central claim that independent injection uses fewer forward-solver iterations. The manuscript provides no evidence that these Lipschitz bounds were measured, enforced, or satisfied by the trained models evaluated on NCI1, PROTEINS, and MUTAG. If the quantum-signal Lipschitz constant exceeds the threshold required for a contraction factor <1, the reported iteration advantage disappears and the comparison to the classical DEQ baseline is no longer valid.
Authors: We agree that explicit verification of the Lipschitz assumptions in the trained models would strengthen the connection between the theoretical guarantees and the empirical iteration counts. The contraction analysis provides sufficient conditions under which each injection variant is guaranteed to converge, and the independent variant is predicted to do so under milder conditions on the quantum-signal Lipschitz constant. The reported iteration numbers, however, are direct empirical measurements obtained from the converged solver runs on the trained models for NCI1, PROTEINS, and MUTAG; all methods reached a fixed point within the stated iteration budgets. In the revised manuscript we will add a supplementary analysis that estimates the Lipschitz constants of the quantum signal and classical backbone for the final trained models on each dataset. This will allow us to check whether the contraction thresholds are met in practice and to discuss any implications for the observed iteration advantages. We view this as a partial revision that directly addresses the referee's concern while preserving the manuscript's core empirical and theoretical contributions. revision: partial
Circularity Check
No circularity: definitions and guarantees are independent of reported outcomes
full rationale
The three injection variants are introduced via explicit procedural distinctions (independent vs. state-dependent vs. backbone-dependent recomputation of the quantum signal) that do not presuppose the accuracy or iteration-count results. Contraction guarantees are derived from standard fixed-point arguments under explicit Lipschitz-constant assumptions on the backbone and quantum signal; these assumptions are not fitted to the NCI1/PROTEINS/MUTAG benchmarks and do not reduce the empirical claims to tautologies. No self-citations appear as load-bearing premises, and the benchmark numbers are presented as direct observations rather than predictions forced by the model definition.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Lipschitz constants of the classical backbone and quantum signal are bounded such that each injection variant contracts
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearWe establish contraction guarantees for each variant under explicit assumptions on the Lipschitz constants of the classical backbone and the quantum signal (Theorems III.1–III.3)
-
IndisputableMonolith/Foundation/BranchSelection.leanbranch_selection unclearLip_F(Φ_ID_Θ) ≤ κ; Lip_F(Φ_SD_Θ) ≤ κ + α L_q; Lip_F(Φ_BD_Θ) ≤ κ(1 + α L_q)
Reference graph
Works this paper leans on
-
[1]
Graph neural networks: A review of methods and applications,
J. Zhou, G. Cui, S. Hu, Z. Zhang, C. Yang, Z. Liu, L. Wang, C. Li, and M. Sun, “Graph neural networks: A review of methods and applications,”AI Open, vol. 1, pp. 57–81, 2020. [Online]. Available: https://doi.org/10.1016/j.aiopen.2021.01.001
-
[2]
G. Corso, H. Stark, S. Jegelka, T. Jaakkola, and R. Barzilay, “Graph neural networks,”Nature Reviews Methods Primers, vol. 4, no. 1, p. 17, mar 2024. [Online]. Available: https://doi.org/10.1038/ s43586-024-00294-7
work page 2024
-
[3]
Semi-Supervised Classification with Graph Convolutional Networks
T. N. Kipf and M. Welling, “Semi-supervised classification with graph convolutional networks,” inInternational Conference on Learning Representations (ICLR), 2017. [Online]. Available: https: //doi.org/10.48550/arXiv.1609.02907
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1609.02907 2017
-
[4]
Inductive Representation Learning on Large Graphs,
W. L. Hamilton, R. Ying, and J. Leskovec, “Inductive representation learning on large graphs,” 2018. [Online]. Available: https://doi.org/10. 48550/arXiv.1706.02216
-
[5]
How Powerful are Graph Neural Networks?
K. Xu, W. Hu, J. Leskovec, and S. Jegelka, “How powerful are graph neural networks?” inInternational Conference on Learning Representations (ICLR), 2019. [Online]. Available: https://doi.org/10. 48550/arXiv.1810.00826
work page internal anchor Pith review arXiv 2019
-
[6]
Graph neural networks exponentially lose expressive power for node classification,
K. Oono and T. Suzuki, “Graph neural networks exponentially lose expressive power for node classification,” inInternational Conference on Learning Representations (ICLR), 2020. [Online]. Available: https://openreview.net/forum?id=S1ldO2EFPr
work page 2020
-
[7]
Zico Kolter, and Vladlen Koltun
S. Bai, J. Z. Kolter, and V . Koltun, “Deep equilibrium models,” in Advances in Neural Information Processing Systems, 2019. [Online]. Available: https://doi.org/10.48550/arXiv.1909.01377
-
[8]
Implicit graph neural networks,
F. Gu, H. Chang, W. Zhu, S. Sojoudi, and L. El Ghaoui, “Implicit graph neural networks,” inAdvances in Neural Information Processing Systems, H. Larochelle, M. Ranzato, R. Hadsell, M. Balcan, and H. Lin, Eds., vol. 33. Curran Associates, Inc., 2020, pp. 11 984– 11 995. [Online]. Available: https://proceedings.neurips.cc/paper files/ paper/2020/file/8b5c84...
work page 2020
-
[9]
Torchdeq: A library for deep equilibrium models,
Z. Geng and J. Z. Kolter, “Torchdeq: A library for deep equilibrium models,” 2023. [Online]. Available: https://doi.org/10.48550/arXiv.2310. 18605
-
[10]
Monotone operator equilibrium networks,
E. Winston and J. Z. Kolter, “Monotone operator equilibrium networks,” inAdvances in Neural Information Process- ing Systems, vol. 33, 2020, pp. 10 718–10 728. [Online]. Available: https://proceedings.neurips.cc/paper files/paper/2020/hash/ 798d1c2813cbdf8bcdb388db0e32d496-Abstract.html
work page 2020
-
[11]
Nature Reviews Physics3(9), 625–644 (2021) https: //doi.org/10.1038/s42254-021-00348-9
M. Cerezo, A. Arrasmith, R. Babbush, S. C. Benjamin, S. Endo, K. Fujii, J. R. McClean, K. Mitarai, X. Yuan, L. Cincio, and P. J. Coles, “Variational quantum algorithms,”Nature Reviews Physics, vol. 3, no. 9, pp. 625–644, Aug. 2021. [Online]. Available: https://doi.org/10.1038/s42254-021-00348-9
-
[12]
Quantum Science and Technology4(4), 043001 (2019) https://doi.org/10.1088/2058-9565/ab4eb5
M. Benedetti, E. Lloyd, S. Sack, and M. Fiorentini, “Parameterized quantum circuits as machine learning models,”Quantum Science and Technology, vol. 4, no. 4, p. 043001, Nov. 2019. [Online]. Available: https://doi.org/10.1088/2058-9565/ab4eb5
-
[13]
Embedding learning in hybrid quantum-classical neural networks,
M. Liu, J. Liu, R. Liu, H. Makhanov, D. Lykov, A. Apte, and Y . Alexeev, “Embedding learning in hybrid quantum-classical neural networks,” in2022 IEEE International Conference on Quantum Computing and Engineering (QCE). IEEE, Sep. 2022, pp. 79–86. [Online]. Available: https://doi.org/10.1109/QCE53715.2022.00026
-
[14]
GraphQNTK: Quantum neural tangent kernel for graph data,
Y . Tang and J. Yan, “GraphQNTK: Quantum neural tangent kernel for graph data,” inAdvances in Neural Information Processing Systems, S. Koyejo, S. Mohamed, A. Agarwal, D. Belgrave, K. Cho, and A. Oh, Eds., vol. 35. Curran Associates, Inc., 2022, pp. 6104–6118. [Online]. Available: https://doi.org/10.5555/3600270.3600712
-
[15]
A. Ray, D. Madan, S. Patil, P. Pati, M. Rapsomaniki, A. Kohlakala, T. R. Dlamini, S. J. Muller, K. Rhrissorrakrai, F. Utro, and L. Parida, “Hybrid quantum-classical graph neural networks for tumor classification in digital pathology,” in2024 IEEE International Conference on Quantum Computing and Engineering (QCE), vol. 01, 2024, pp. 1611–1616. [Online]. A...
-
[16]
M. Cui, L. Chang, A. Chau, H. Mekuria, L. Adwankar, S. Pendyala, and L. McMahan, “Efficient and optimized small organic molecular graph generation pathway using a quantum generative adversarial network,” in2024 IEEE International Conference on Quantum Computing and Engineering (QCE), vol. 01, 2024, pp. 1565–1570. [Online]. Available: https://doi.org/10.11...
-
[18]
Nature Communications9(1) (2018) https:// doi.org/10.1038/s41467-018-07090-4
J. R. McClean, S. Boixo, V . N. Smelyanskiy, R. Babbush, and H. Neven, “Barren plateaus in quantum neural network training landscapes,”Nature Communications, vol. 9, no. 1, Nov. 2018. [Online]. Available: https://doi.org/10.1038/s41467-018-07090-4
-
[19]
A lie algebraic theory of barren plateaus for deep parameterized quantum circuits,
M. Ragone, B. N. Bakalov, F. Sauvage, A. F. Kemper, C. Ortiz Marrero, M. Larocca, and M. Cerezo, “A lie algebraic theory of barren plateaus for deep parameterized quantum circuits,”Nature Communications, vol. 15, no. 1, Aug. 2024. [Online]. Available: https://doi.org/10.1038/ s41467-024-49909-3
work page 2024
-
[20]
Characterizing barren plateaus in quantum ans ¨atze with the adjoint representation,
E. Fontana, D. Herman, S. Chakrabarti, N. Kumar, R. Yalovetzky, J. Heredge, S. H. Sureshbabu, and M. Pistoia, “Characterizing barren plateaus in quantum ans ¨atze with the adjoint representation,”Nature Communications, vol. 15, no. 1, Aug. 2024. [Online]. Available: https://doi.org/10.1038/s41467-024-49910-w
-
[21]
Evaluating analytic gradients on quantum hardware,
M. Schuld, V . Bergholm, C. Gogolin, J. Izaac, and N. Killoran, “Evaluating analytic gradients on quantum hardware,”Physical Review A, vol. 99, no. 3, p. 032331, 2019. [Online]. Available: https://doi.org/10.1103/PhysRevA.99.032331
-
[22]
The power of quantum neural networks,
A. Abbas, D. Sutter, C. Zoufal, A. Lucchi, A. Figalli, and S. Woerner, “The power of quantum neural networks,”Nature Computational Science, vol. 1, no. 6, pp. 403–409, Jun. 2021. [Online]. Available: https://doi.org/10.1038/s43588-021-00084-1
-
[23]
Quantum deep equilibrium models,
P. Schleich, M. Skreta, L. B. Kristensen, R. Vargas-Hernandez, and A. Aspuru-Guzik, “Quantum deep equilibrium models,” inThe Thirty-eighth Annual Conference on Neural Information Processing Systems, 2024. [Online]. Available: https://openreview.net/forum?id= CWhwKb0Q4k
work page 2024
-
[24]
L. E. Ghaoui, F. Gu, B. Travacca, A. Askari, and A. Y . Tsai, “Implicit deep learning,” 2020. [Online]. Available: https: //doi.org/10.48550/arXiv.1908.06315
-
[25]
CerDEQ: Certifiable deep equilibrium model,
M. Li, Y . Wang, and Z. Lin, “CerDEQ: Certifiable deep equilibrium model,” inProceedings of the 39th International Conference on Machine Learning, ser. Proceedings of Machine Learning Research, K. Chaudhuri, S. Jegelka, L. Song, C. Szepesvari, G. Niu, and S. Sabato, Eds., vol. 162. PMLR, 17–23 Jul 2022, pp. 12 998–13 013. [Online]. Available: https://proc...
work page 2022
-
[26]
Lyapunov-stable deep equilibrium models,
H. Chu, S. Wei, T. Liu, Y . Zhao, and Y . Miyatake, “Lyapunov-stable deep equilibrium models,” inProceedings of the AAAI Conference on Artificial Intelligence, vol. 38, no. 10, 2024, pp. 11 615–11 623. [Online]. Available: https://doi.org/10.1609/aaai.v38i10.29044
-
[27]
Implicit graph neural networks: A monotone operator viewpoint,
J. Baker, Q. Wang, C. D. Hauck, and B. Wang, “Implicit graph neural networks: A monotone operator viewpoint,” inProceedings of the 40th International Conference on Machine Learning, ser. Proceedings of Machine Learning Research, vol. 202. PMLR, 2023, pp. 1521–1548. [Online]. Available: https://proceedings.mlr.press/v202/baker23a.html
work page 2023
-
[28]
Eignn: Efficient infinite-depth graph neural networks,
J. Liu, K. Kawaguchi, B. Hooi, Y . Wang, and X. Xiao, “Eignn: Efficient infinite-depth graph neural networks,” inAdvances in Neural Information Processing Systems, vol. 34, 2021, pp. 18 762– 18 773. [Online]. Available: https://proceedings.neurips.cc/paper/2021/ hash/9bd5ee6fe55aaeb673025dbcb8f939c1-Abstract.html
work page 2021
-
[29]
Mgnni: Multiscale graph neural networks with implicit layers,
J. Liu, B. Hooi, K. Kawaguchi, and X. Xiao, “Mgnni: Multiscale graph neural networks with implicit layers,” 2022. [Online]. Available: https://doi.org/10.48550/arXiv.2210.08353
-
[30]
Optimization- induced graph implicit nonlinear diffusion,
Q. Chen, Y . Wang, Y . Wang, J. Yang, and Z. Lin, “Optimization- induced graph implicit nonlinear diffusion,” 2022. [Online]. Available: https://doi.org/10.48550/arXiv.2206.14418
-
[31]
URLhttps://doi.org/10.48550/arXiv
J. Zheng, Q. Gao, and Y . Lv, “Quantum graph convolutional neural networks,” 2021. [Online]. Available: https://doi.org/10.48550/arXiv. 2107.03257
work page internal anchor Pith review doi:10.48550/arxiv 2021
-
[32]
On the design of quantum graph convolutional neural network in the nisq-era and beyond,
Z. Hu, J. Li, Z. Pan, S. Zhou, L. Yang, C. Ding, O. Khan, T. Geng, and W. Jiang, “On the design of quantum graph convolutional neural network in the nisq-era and beyond,” in Proceedings - 2022 IEEE 40th International Conference on Computer Design, ICCD 2022, 2022, pp. 290–297. [Online]. Available: https://doi.org/10.1109/ICCD56317.2022.00050
-
[33]
Quantum graph neural networks,
G. Verdon, T. McCourt, E. Luzhnica, V . Singh, S. Leichenauer, and J. Hidary, “Quantum graph neural networks,” 2019, arXiv preprint. [Online]. Available: https://doi.org/10.48550/arXiv.1909.12264 11
-
[34]
X. Ye, H. Xiong, J. Huang, Z. Chen, J. Wang, and J. Yan, “On designing general and expressive quantum graph neural networks with applications to MILP instance representation,” inThe Thirteenth International Conference on Learning Representations, 2025. [Online]. Available: https://openreview.net/forum?id=IQi8JOqLuv
work page 2025
-
[35]
Inductive graph representation learning with quantum graph neural networks,
A. M. Faria, I. F. Gra ˜na, and S. Varsamopoulos, “Inductive graph representation learning with quantum graph neural networks,” 2026. [Online]. Available: https://arxiv.org/abs/2503.24111
-
[36]
From graphs to qubits: A critical review of quantum graph neural networks,
A. Ceschini, F. Mauro, F. D. Falco, A. Sebastianelli, A. Verdone, A. Rosato, B. L. Saux, M. Panella, P. Gamba, and S. L. Ullo, “From graphs to qubits: A critical review of quantum graph neural networks,”
-
[37]
Available: https://doi.org/10.48550/arXiv.2408.06524
[Online]. Available: https://doi.org/10.48550/arXiv.2408.06524
-
[38]
Hardware-Aware Quantum Kernel Design Based on Graph Neural Networks
Y . Liu, F. Meng, L. Wang, Y . Hu, S. Li, X. Yu, and Z. Zhang, “Haqgnn: Hardware-aware quantum kernel design based on graph neural networks,” 2025. [Online]. Available: https://doi.org/10.48550/ arXiv.2506.21161
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[39]
FiLM : Visual reasoning with a general conditioning layer
E. Perez, F. Strub, H. de Vries, V . Dumoulin, and A. Courville, “Film: Visual reasoning with a general conditioning layer,”Proceedings of the AAAI Conference on Artificial Intelligence, vol. 32, no. 1, Apr. 2018. [Online]. Available: https://doi.org/10.1609/aaai.v32i1.11671
-
[40]
A hybrid quantum-classical neural network with deep residual learning,
Y . Liang, W. Peng, Z.-J. Zheng, O. Silv ´en, and G. Zhao, “A hybrid quantum-classical neural network with deep residual learning,” 2021. [Online]. Available: https://doi.org/10.48550/arXiv.2012.07772
-
[41]
Resqnets: A residual approach for mitigating barren plateaus in quantum neural networks,
M. Kashif and S. Al-kuwari, “Resqnets: A residual approach for mitigating barren plateaus in quantum neural networks,”EPJ Quantum Technology, vol. 11, p. 4, 2024. [Online]. Available: https://doi.org/10.1140/epjqt/s40507-023-00216-8
-
[42]
On optimizing hyperparameters for quantum neural networks,
S. Herbst, V . De Maio, and I. Brandic, “On optimizing hyperparameters for quantum neural networks,” in2024 IEEE International Conference on Quantum Computing and Engineering (QCE). IEEE, Sep. 2024, pp. 1478–1489. [Online]. Available: https://doi.org/10.1109/QCE60285. 2024.00174
-
[43]
Data re-uploading for a universal quantum classifier,
A. P ´erez-Salinas, A. Cervera-Lierta, E. Gil-Fuster, and J. I. Latorre, “Data re-uploading for a universal quantum classifier,” Quantum, vol. 4, p. 226, Feb. 2020. [Online]. Available: https: //doi.org/10.22331/q-2020-02-06-226
-
[44]
Rudin,Principles of mathematical analysis
W. Rudin,Principles of mathematical analysis. McGraw-Hill, 1976, vol. 3
work page 1976
-
[45]
R. A. Horn and C. R. Johnson,Matrix Analysis, 2nd ed. Cambridge University Press, 2012. [Online]. Available: https://doi.org/10.1017/ 9781139020411
work page 2012
-
[46]
A. Vaswani, N. Shazeer, N. Parmar, J. Uszkoreit, L. Jones, A. N. Gomez, L. Kaiser, and I. Polosukhin, “Attention is all you need,” inAdvances in Neural Information Processing Systems (NeurIPS), vol. 30, 2017, pp. 5998–6008. [Online]. Available: https://doi.org/10.48550/arXiv.1706.03762
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1706.03762 2017
-
[47]
C. Morris, N. M. Kriege, F. Bause, K. Kersting, P. Mutzel, and M. Neumann, “Tudataset: A collection of benchmark datasets for learning with graphs,” inICML 2020 Workshop on Graph Representation Learning and Beyond (GRL+ 2020), 2020. [Online]. Available: https://doi.org/10.48550/arXiv.2007.08663
-
[48]
A fair comparison of graph neural networks for graph classification,
F. Errica, M. Podda, D. Bacciu, and A. Micheli, “A fair comparison of graph neural networks for graph classification,” inInternational Conference on Learning Representations (ICLR), 2020. [Online]. Available: https://openreview.net/forum?id=HygDF6NFPB
work page 2020
-
[49]
Benchmarking graph neural networks,
V . P. Dwivedi, C. K. Joshi, A. T. Luu, T. Laurent, Y . Bengio, and X. Bresson, “Benchmarking graph neural networks,” 2020. [Online]. Available: https://doi.org/10.48550/arXiv.2003.00982
-
[50]
Decoupled weight decay regularization,
I. Loshchilov and F. Hutter, “Decoupled weight decay regularization,”
-
[51]
Decoupled Weight Decay Regularization
[Online]. Available: https://doi.org/10.48550/arXiv.1711.05101 12
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1711.05101
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.