Recognition: 2 theorem links
· Lean TheoremFrom Syntax to Semantics: Unveiling the Emergence of Chirality in SMILES Translation Models
Pith reviewed 2026-05-12 03:58 UTC · model grok-4.3
The pith
Chemical translation models show a sudden jump in chiral accuracy after a long plateau, driven by encoder-side reorganization of representations.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
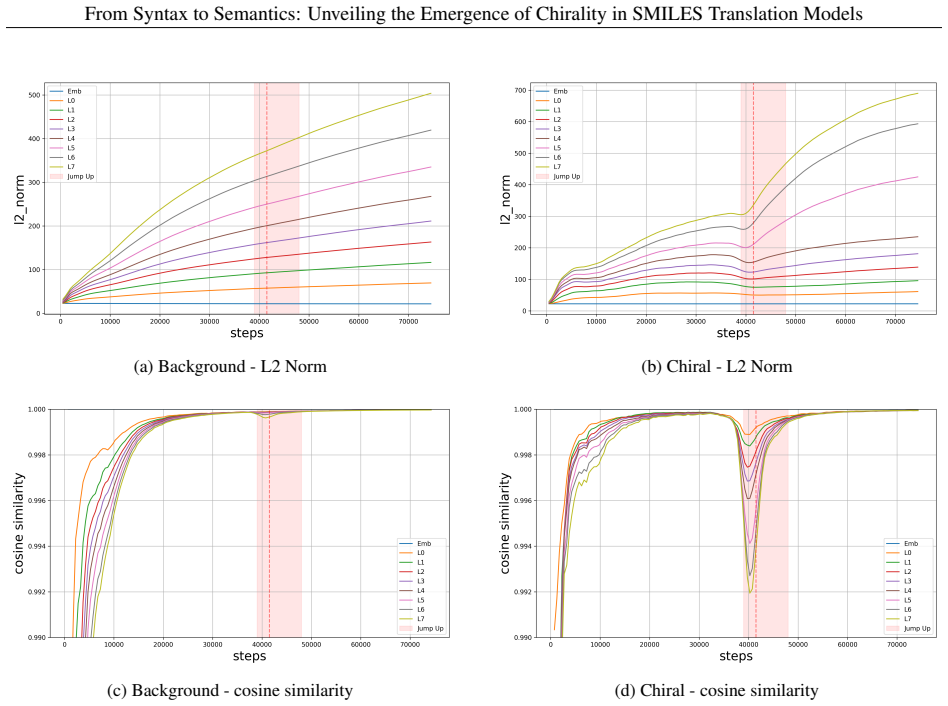
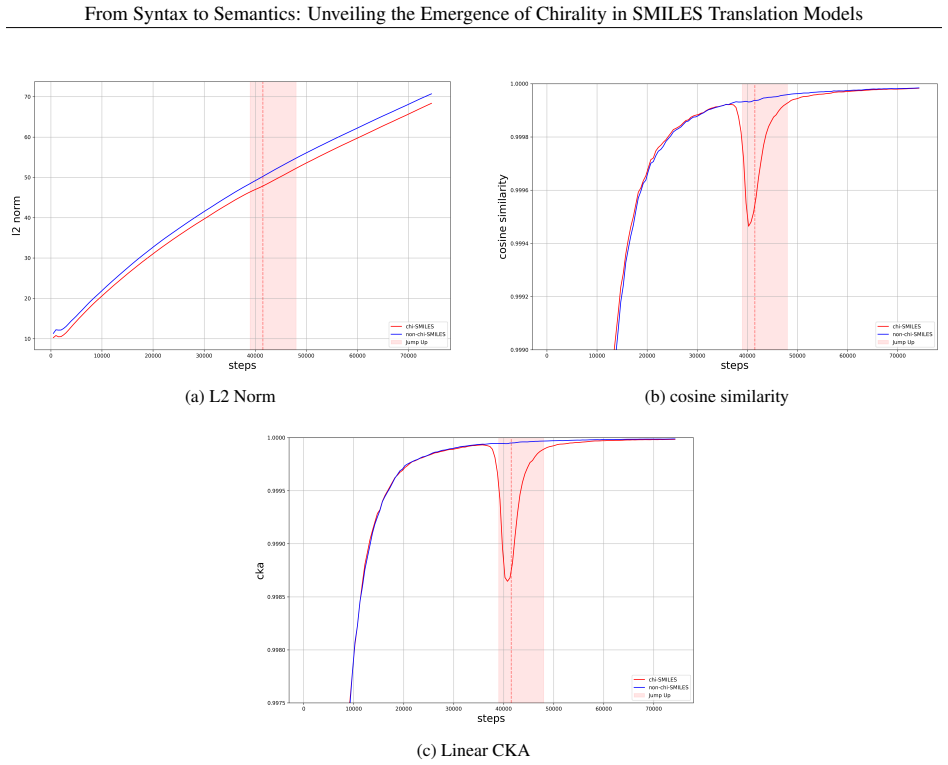
In autoregressive encoder-decoder models for SMILES translation, chiral-token accuracy exhibits a reproducible abrupt increase after an extended plateau phase. This jump coincides with a V-shaped pattern of decreasing then recovering vector norms and directional stability in the residual stream, accompanied by a clear reorganization of chiral molecular representations in latent space. Encoder-decoder cross evaluations and ablation of specific attention heads confirm that the transition is localized to the encoder and involves a small subset of heads that become selectively sensitive to chiral features.
What carries the argument
The encoder-centered mechanism of chiral emergence, marked by transient destabilization and reconstruction of residual-stream vectors together with reorganization of chiral representations in latent space.
If this is right
- Chiral learning plateaus arise from the intrinsic complexity of chiral constraints rather than insufficient model capacity.
- A small number of attention heads can be isolated whose removal selectively lowers chiral-token accuracy even after full training.
- SMILES translation supplies a controlled experimental system for studying how semantic features emerge in chemical language models.
- High-temporal-resolution checkpoint tracking exposes dynamic representation changes that endpoint evaluations alone would miss.
Where Pith is reading between the lines
- Vector-norm monitoring during training could flag the moment when complex semantic features such as chirality are being internalized.
- The encoder-focused transition pattern may appear for other stereochemical or conformational properties beyond chirality.
- Architecture choices that strengthen encoder capacity or stabilize residual streams might shorten the plateau phase for difficult chemical properties.
Load-bearing premise
The abrupt accuracy jump, V-shaped norm changes, and latent-space reorganization reflect genuine acquisition of chiral semantics rather than training artifacts, data imbalances, or choices in which checkpoints and heads are examined.
What would settle it
Training the same models on SMILES strings where all chiral center labels have been randomly reassigned or removed entirely would produce the same jump and norm trajectory if the observed dynamics are unrelated to actual chirality.
Figures

read the original abstract
Understanding how chemical language models (CLMs) learn chemical meaning from molecular string representations, rather than only surface-level string patterns, is an important question in chemical representation learning and machine learning for chemistry. Chirality provides a demanding test case: enantiomers can differ greatly in pharmacological activity and toxicity, yet CLMs often struggle to distinguish chiral configurations reliably. Here we present Pan-CORE (Pan-Chemical Omniscale Representation Engine), a family of autoregressive Transformer-based encoder-decoder models for SMILES translation, and use high-temporal-resolution checkpoint analysis to investigate how chiral information is learned during training. Across all tested Pan-CORE variants, we observe a reproducible jump-up in which chiral-token accuracy rises abruptly after a long plateau, suggesting that chiral learning stagnation is not explained by model capacity alone and instead reflects the complexity of chiral constraints. Analyses of attention dynamics, residual-stream trajectories, and latent-space geometry support an encoder-centered mechanism in which chiral-token representations undergo transient destabilization and reconstruction, seen as a V-shaped drop and recovery in vector norm and directional stability, together with a clear reorganization of chiral molecular representations in the latent space. Encoder-decoder cross-evaluation further supports the encoder-centered nature of the transition, and targeted attention-head ablation identifies a small set of chiral-sensitive heads whose removal selectively reduces chiral-token accuracy even in the fully trained model. These findings show that SMILES translation can serve as a useful experimental system for mechanistic analysis of semantic emergence in CLMs, with implications for interpretable chemical representation learning.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces the Pan-CORE family of autoregressive Transformer encoder-decoder models for SMILES translation and uses high-temporal-resolution checkpoint analysis to demonstrate that chiral-token accuracy exhibits a reproducible abrupt jump after a long plateau across model variants. It interprets this as evidence that chiral learning stagnation reflects the complexity of chiral constraints rather than insufficient model capacity, supported by analyses of attention dynamics, V-shaped residual norm trajectories, latent-space reorganization, encoder-decoder cross-evaluations, and targeted attention-head ablations that identify a small set of chiral-sensitive heads.
Significance. If the transition and supporting dynamics are shown to be specific to chirality, the work would provide a useful mechanistic case study for semantic emergence in chemical language models, with the high-resolution checkpointing, residual-stream analysis, and head-ablation results offering a concrete template for interpretability research in CLMs. The reproducibility across Pan-CORE variants and the identification of encoder-centered mechanisms are particular strengths that could inform more reliable handling of stereochemistry in downstream applications.
major comments (3)
- [Results on training dynamics and ablation studies] The central claim that the abrupt jump in chiral-token accuracy demonstrates chirality-specific constraint complexity (rather than a generic training artifact) is load-bearing for the interpretation of semantic emergence. However, the training-dynamics and ablation sections do not report accuracy trajectories for non-chiral tokens or include control experiments on datasets with randomized chirality labels, so the plateau-jump pattern, V-shaped norm drop, and latent reorganization cannot yet be distinguished from general autoregressive SMILES phase transitions.
- [Analyses of residual-stream trajectories and latent-space geometry] The encoder-centered mechanism is supported by cross-evaluation and head-ablation results, but the manuscript provides no quantitative baseline (e.g., norm dynamics or latent geometry) for non-chiral molecular representations or shuffled-chirality controls. This omission leaves the V-shaped vector-norm behavior and latent-space reorganization open to alternative explanations as post-hoc observations of general training phenomena.
- [Methods and experimental setup] The experimental protocol lacks sufficient detail on data splits, exact definition and computation of chiral-token accuracy, checkpoint sampling intervals, and statistical controls for the reported reproducibility of the jump. These omissions directly affect the ability to assess whether the observed transition is robust or sensitive to unexamined confounds.
minor comments (2)
- [Abstract and §1] The abstract and introduction would benefit from an explicit definition of 'chiral-token accuracy' and 'chiral-sensitive heads' before the results are presented.
- [Figures] Figure captions should specify the exact Pan-CORE variants, number of runs, and checkpoint resolution used for each panel to improve reproducibility.
Simulated Author's Rebuttal
We thank the referee for their positive assessment of the work's potential significance and for the constructive major comments, which help clarify the evidence needed to support chirality-specific emergence. We address each point below and have prepared revisions that add the requested controls, baselines, and methodological details.
read point-by-point responses
-
Referee: [Results on training dynamics and ablation studies] The central claim that the abrupt jump in chiral-token accuracy demonstrates chirality-specific constraint complexity (rather than a generic training artifact) is load-bearing for the interpretation of semantic emergence. However, the training-dynamics and ablation sections do not report accuracy trajectories for non-chiral tokens or include control experiments on datasets with randomized chirality labels, so the plateau-jump pattern, V-shaped norm drop, and latent reorganization cannot yet be distinguished from general autoregressive SMILES phase transitions.
Authors: We agree that direct comparisons to non-chiral tokens and randomized-chirality controls are necessary to strengthen the claim of specificity. In the revised manuscript we have added accuracy trajectories for non-chiral tokens (e.g., atom and bond tokens without stereodescriptors), which improve monotonically without an abrupt jump. We have also included results from shuffled-chirality control datasets in a new supplementary figure; these show the absence of both the plateau-jump and the V-shaped residual-norm signature, supporting that the observed transition reflects the complexity of chiral constraints rather than a generic autoregressive phase change. revision: yes
-
Referee: [Analyses of residual-stream trajectories and latent-space geometry] The encoder-centered mechanism is supported by cross-evaluation and head-ablation results, but the manuscript provides no quantitative baseline (e.g., norm dynamics or latent geometry) for non-chiral molecular representations or shuffled-chirality controls. This omission leaves the V-shaped vector-norm behavior and latent-space reorganization open to alternative explanations as post-hoc observations of general training phenomena.
Authors: We acknowledge the value of explicit quantitative baselines. The revised manuscript now reports residual-norm trajectories and latent-space geometry metrics for non-chiral molecular representations, which lack the V-shaped drop and directional reorganization. Parallel analyses on shuffled-chirality controls are included and demonstrate that the encoder-centered reorganization is absent when chirality labels are randomized. These additions, placed alongside the original cross-evaluation and ablation results, reduce the plausibility of purely generic training explanations. revision: yes
-
Referee: [Methods and experimental setup] The experimental protocol lacks sufficient detail on data splits, exact definition and computation of chiral-token accuracy, checkpoint sampling intervals, and statistical controls for the reported reproducibility of the jump. These omissions directly affect the ability to assess whether the observed transition is robust or sensitive to unexamined confounds.
Authors: We have expanded the Methods section with the requested information. Data splits are now described as an 80/10/10 train/validation/test partition on a curated set of 500k SMILES strings containing explicit stereochemistry, generated with RDKit canonicalization. Chiral-token accuracy is defined as the fraction of correctly predicted '@' and '@@' tokens across all chiral centers in the decoded SMILES, computed token-wise on the test set. Checkpoints were sampled every 500 steps during the plateau and every 50 steps in the transition window; reproducibility was assessed across five independent runs with distinct random seeds, confirming the jump occurs within a consistent 200-step window. These details have been added to the main text and supplementary material. revision: yes
Circularity Check
No circularity: purely empirical training dynamics analysis
full rationale
The paper presents no derivation chain, equations, or theoretical claims that reduce to inputs by construction. All load-bearing results (chiral-token accuracy jumps, V-shaped norm dynamics, latent reorganization, head ablations) are direct measurements from model training runs and checkpoint analyses on Pan-CORE variants. These are falsifiable experimental observations rather than self-definitional, fitted-input predictions, or self-citation-dependent uniqueness arguments. The study is self-contained against external benchmarks via ablation controls and cross-evaluations.
Axiom & Free-Parameter Ledger
axioms (1)
- standard math Standard properties of autoregressive transformer attention and residual streams hold during training
Lean theorems connected to this paper
-
IndisputableMonolith/Foundation/RealityFromDistinctionreality_from_one_distinction unclearAcross all tested Pan-CORE variants, we observe a reproducible jump-up in which chiral-token accuracy rises abruptly after a long plateau... Analyses of attention dynamics, residual-stream trajectories, and latent-space geometry support an encoder-centered mechanism in which chiral-token representations undergo transient destabilization and reconstruction, seen as a V-shaped drop and recovery in vector norm
-
IndisputableMonolith/Cost/FunctionalEquationwashburn_uniqueness_aczel unclearWe identify the jump-up as the abrupt learning of SMILES-level chiral constraints, characterized by a sudden rise in chiral-token accuracy and a sharp improvement in model confidence over R/S configurations.
Reference graph
Works this paper leans on
-
[1]
A comprehensive overview of large language models.ACM Trans
Humza Naveed, Asad Ullah Khan, Shi Qiu, Muhammad Saqib, Saeed Anwar, Muhammad Usman, Naveed Akhtar, Nick Barnes, and Ajmal Mian. A comprehensive overview of large language models.ACM Trans. Intell. Syst. Technol., 16(5):1–72, 2025. ISSN 2157-6904,2157-6912. doi:10.1145/3744746
-
[2]
A systematic review of deep learning chemical language models in recent era.J
Hector Flores-Hernandez and Emmanuel Martinez-Ledesma. A systematic review of deep learning chemical language models in recent era.J. Cheminform., 16(1):129, 2024. ISSN 1758-2946,1758-2946. doi:10.1186/s13321- 024-00916-y
-
[3]
Protein large language models: A comprehensive survey
Yijia Xiao, Wanjia Zhao, Junkai Zhang, Yiqiao Jin, Han Zhang, Zhicheng Ren, Renliang Sun, Haixin Wang, Guancheng Wan, Pan Lu, Xiao Luo, Yu Zhang, James Zou, Yizhou Sun, and Wei Wang. Protein large language models: A comprehensive survey. InFindings of the Association for Computational Linguistics: EMNLP 2025, pages 23080–23103, Stroudsburg, PA, USA, 2025....
-
[4]
A comprehensive survey of genome language models in bioinformatics.Brief
Liyuan Shu, Jiao Tang, Xiaoyu Guan, and Daoqiang Zhang. A comprehensive survey of genome language models in bioinformatics.Brief. Bioinform., 27(1):bbaf724, 2026. ISSN 1467-5463,1477-4054. doi:10.1093/bib/bbaf724
-
[5]
Smiles, a chemical language and information system
David Weininger. SMILES, a chemical language and information system. 1. introduction to methodol- ogy and encoding rules.J. Chem. Inf. Comput. Sci., 28(1):31–36, 1988. ISSN 0095-2338,1520-5142. doi:10.1021/ci00057a005
-
[6]
Automatic chemical design using a data-driven continuous representation of molecules.ACS Cent
Rafael Gómez-Bombarelli, Jennifer N Wei, David Duvenaud, José Miguel Hernández-Lobato, Benjamín Sánchez- Lengeling, Dennis Sheberla, Jorge Aguilera-Iparraguirre, Timothy D Hirzel, Ryan P Adams, and Alán Aspuru- Guzik. Automatic chemical design using a data-driven continuous representation of molecules.ACS Cent. Sci., 4 (2):268–276, 2018. ISSN 2374-7943,23...
-
[7]
Beyond performance: how design choices shape chemical language models.J
Inken Fender, Jannik Adrian Gut, and Thomas Lemmin. Beyond performance: how design choices shape chemical language models.J. Cheminform., 17(1):173, 2025. ISSN 1758-2946,1758-2946. doi:10.1186/s13321-025-01099- w
-
[8]
Measuring chemical LLM robustness to molecular representations: a SMILES variation-based framework.J
Veronika Ganeeva, Kuzma Khrabrov, Artur Kadurin, and Elena Tutubalina. Measuring chemical LLM robustness to molecular representations: a SMILES variation-based framework.J. Cheminform., 17(1):164, 2025. ISSN 1758-2946,1758-2946. doi:10.1186/s13321-025-01079-0
-
[9]
Graph neural networks in particle physics
Mario Krenn, Florian Häse, Akshatkumar Nigam, Pascal Friederich, and Alán Aspuru-Guzik. Self-referencing em- bedded strings (SELFIES): A 100% robust molecular string representation.arXiv [cs.LG], 2019. doi:10.1088/2632- 2153/aba947
-
[10]
Ashish Vaswani, Noam Shazeer, Niki Parmar, Jakob Uszkoreit, Llion Jones, Aidan N Gomez, Lukasz Kaiser, and Illia Polosukhin. Attention is all you need.arXiv [cs.CL], 2017. doi:10.48550/arXiv.1706.03762
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1706.03762 2017
-
[11]
Chemberta: Large-scale self- supervised pretraining for molecular property prediction, 2020
Seyone Chithrananda, Gabe Grand, and Bharath Ramsundar. ChemBERTa: Large-scale self-supervised pretraining for molecular property prediction.arXiv [cs.LG], 2020. doi:10.48550/arXiv.2010.09885. 10 From Syntax to Semantics: Unveiling the Emergence of Chirality in SMILES Translation Models
-
[12]
Large-scale chemical language representations capture molecular structure and properties.Nat
Jerret Ross, Brian Belgodere, Vijil Chenthamarakshan, Inkit Padhi, Youssef Mroueh, and Payel Das. Large-scale chemical language representations capture molecular structure and properties.Nat. Mach. Intell., 4(12):1256–1264,
-
[13]
doi:10.1038/s42256-022-00580-7
ISSN 2522-5839,2522-5839. doi:10.1038/s42256-022-00580-7
-
[14]
An open-source family of large encoder-decoder foundation models for chemistry.Commun
Eduardo Soares, Emilio Vital Brazil, Victor Shirasuna, Dmitry Zubarev, Renato Cerqueira, and Kristin Schmidt. An open-source family of large encoder-decoder foundation models for chemistry.Commun. Chem., 8(1):193,
-
[15]
doi:10.1038/s42004-025-01585-0
ISSN 2399-3669,2399-3669. doi:10.1038/s42004-025-01585-0
-
[16]
MolFM: A multimodal molecular foundation model.arXiv [q-bio.BM], 2023
Yizhen Luo, Kai Yang, Massimo Hong, Xing Yi Liu, and Zaiqing Nie. MolFM: A multimodal molecular foundation model.arXiv [q-bio.BM], 2023. doi:10.48550/arXiv.2307.09484
-
[17]
ChemBERTa-3: An open source training framework for chemical foundation models.ChemRxiv, 2025
Riya Singh, Aryan Amit Barsainyan, Rida Irfan, Connor Joseph Amorin, Stewart He, Tony Davis, Arun Thia- garajan, Shiva Sankaran, Seyone Chithrananda, Walid Ahmad, Derek Jones, Kevin McLoughlin, Hyojin Kim, Anoushka Bhutani, Shreyas Vinaya Sathyanarayana, Venkat Viswanathan, Jonathan E Allen, and Bharath Ram- sundar. ChemBERTa-3: An open source training fr...
-
[18]
Eduardo Soares, Victor Yukio Shirasuna, Emilio Vital Brazil, Indra Priyadarsini, and Seiji Takeda. Multi-view mixture-of-experts for predicting molecular properties using SMILES, SELFIES, and graph-based representations. Mach. Learn. Sci. Technol., 6(2):025070, 2025. ISSN 2632-2153. doi:10.1088/2632-2153/ade4ef
-
[19]
What does bert look at? an analysis of bert’s attention
Kevin Clark, Urvashi Khandelwal, Omer Levy, and Christopher D Manning. What does BERT look at? an analysis of BERT’s attention.arXiv [cs.CL], 2019. doi:10.48550/arXiv.1906.04341
-
[20]
A primer in BERTology: What we know about how BERT works.arXiv [cs.CL], 2020
Anna Rogers, Olga Kovaleva, and Anna Rumshisky. A primer in BERTology: What we know about how BERT works.arXiv [cs.CL], 2020. doi:10.48550/arXiv.2002.12327
-
[21]
A mathematical framework for transformer circuits
Nelson Elhage, Neel Nanda, Catherine Olsson, Tom Henighan, Nicholas Joseph, Ben Mann, Amanda Askell, Yuntao Bai, Anna Chen, Tom Conerly, Nova DasSarma, Dawn Drain, Deep Ganguli, Zac Hatfield-Dodds, Danny Hernandez, Andy Jones, Jackson Kernion, Liane Lovitt, Kamal Ndousse, Dario Amodei, Tom Brown, Jack Clark, Jared Kaplan, Sam McCandlish, Chris Olah. A mat...
work page 2021
-
[22]
Shawn Im, Changdae Oh, Zhen Fang, and Sharon Li. How do transformers learn to associate tokens: Gradient leading terms bring mechanistic interpretability. InThe F ourteenth International Conference on Learning Representations, 2025
work page 2025
-
[23]
Evolution of concepts in language model pre-training.arXiv [cs.CL], 2026
Xuyang Ge, Wentao Shu, Jiaxing Wu, Yunhua Zhou, Zhengfu He, and Xipeng Qiu. Evolution of concepts in language model pre-training.arXiv [cs.CL], 2026. doi:10.48550/arXiv.2509.17196
-
[24]
Investigation of chemical structure recognition by encoder-decoder models in learning progress.J
Shumpei Nemoto, Tadahaya Mizuno, and Hiroyuki Kusuhara. Investigation of chemical structure recognition by encoder-decoder models in learning progress.J. Cheminform., 15(1):45, 2023. ISSN 1758-2946,1758-2946. doi:10.1186/s13321-023-00713-z
-
[25]
Circuits, features, and heuristics in molecular transformers.arXiv [cs.LG], 2025
Kristof Varadi, Mark Marosi, and Peter Antal. Circuits, features, and heuristics in molecular transformers.arXiv [cs.LG], 2025. doi:10.48550/arXiv.2512.09757
-
[26]
A short history of thalidomide embryopathy.Teratology, 38(3):203–215, 1988
W Lenz. A short history of thalidomide embryopathy.Teratology, 38(3):203–215, 1988. ISSN 1096-9926,0040-
work page 1988
-
[27]
doi:10.1002/tera.1420380303
-
[28]
G Blaschke, H P Kraft, K Fickentscher, and F Köhler. Chromatographic separation of racemic thalidomide and teratogenic activity of its enantiomers (author’s transl).Arzneimittelforschung, 29(10):1640–1642, 1979. ISSN 0004-4172,1616-7066
work page 1979
-
[29]
Sayyed Jalil Mahdizadeh and Leif A Eriksson. MolAI: A deep learning framework for data-driven molecular descriptor generation and advanced drug discovery applications.J. Chem. Inf. Model., 2025. ISSN 1549- 9596,1549-960X. doi:10.1021/acs.jcim.5c00491
-
[30]
Yasuhiro Yoshikai, Tadahaya Mizuno, Shumpei Nemoto, and Hiroyuki Kusuhara. Difficulty in chirality recognition for transformer architectures learning chemical structures from string representations.Nat. Commun., 15(1):1197,
-
[31]
doi:10.1038/s41467-024-45102-8
ISSN 2041-1723,2041-1723. doi:10.1038/s41467-024-45102-8
-
[32]
ZINC20—A Free Ultralarge-Scale Chemical Database for Ligand Discovery
John J Irwin, Khanh G Tang, Jennifer Young, Chinzorig Dandarchuluun, Benjamin R Wong, Munkhzul Khurelbaatar, Yurii S Moroz, John Mayfield, and Roger A Sayle. ZINC20-a free ultralarge-scale chemical database for ligand discovery.J. Chem. Inf. Model., 60(12):6065–6073, 2020. ISSN 1549-9596,1549-960X. doi:10.1021/acs.jcim.0c00675
-
[33]
PubChem 2025 update.Nucleic Acids Res., 53(D1):D1516–D1525, 2025
Sunghwan Kim, Jie Chen, Tiejun Cheng, Asta Gindulyte, Jia He, Siqian He, Qingliang Li, Benjamin A Shoemaker, Paul A Thiessen, Bo Yu, Leonid Zaslavsky, Jian Zhang, and Evan E Bolton. PubChem 2025 update.Nucleic Acids Res., 53(D1):D1516–D1525, 2025. ISSN 0305-1048,1362-4962. doi:10.1093/nar/gkae1059. 11 From Syntax to Semantics: Unveiling the Emergence of C...
-
[34]
Neural Machine Translation of Rare Words with Subword Units
Rico Sennrich, Barry Haddow, and Alexandra Birch. Neural machine translation of rare words with subword units. arXiv [cs.CL], 2015. doi:10.48550/arXiv.1508.07909
work page internal anchor Pith review doi:10.48550/arxiv.1508.07909 2015
-
[35]
Google's Neural Machine Translation System: Bridging the Gap between Human and Machine Translation
Yonghui Wu, Mike Schuster, Zhifeng Chen, Quoc V Le, Mohammad Norouzi, Wolfgang Macherey, Maxim Krikun, Yuan Cao, Qin Gao, Klaus Macherey, Jeff Klingner, Apurva Shah, Melvin Johnson, Xiaobing Liu, Łukasz Kaiser, Stephan Gouws, Yoshikiyo Kato, Taku Kudo, Hideto Kazawa, Keith Stevens, George Kurian, Nishant Patil, Wei Wang, Cliff Young, Jason Smith, Jason Ri...
work page internal anchor Pith review doi:10.48550/arxiv.1609.08144 2016
-
[36]
Taku Kudo and John Richardson. SentencePiece: A simple and language independent subword tokenizer and detokenizer for neural text processing.arXiv [cs.CL], 2018. doi:10.48550/arXiv.1808.06226
work page internal anchor Pith review doi:10.48550/arxiv.1808.06226 2018
-
[37]
RDKit: Open-source cheminformatics
-
[38]
Esben Jannik Bjerrum. SMILES enumeration as data augmentation for neural network modeling of molecules. arXiv [cs.LG], 2017. doi:10.48550/arXiv.1703.07076
-
[39]
Randomized SMILES strings improve the quality of molecular generative models.J
Josep Arús-Pous, Simon Viet Johansson, Oleksii Prykhodko, Esben Jannik Bjerrum, Christian Tyrchan, Jean-Louis Reymond, Hongming Chen, and Ola Engkvist. Randomized SMILES strings improve the quality of molecular generative models.J. Cheminform., 11(1):71, 2019. ISSN 1758-2946,1758-2946. doi:10.1186/s13321-019-0393-0
-
[40]
A novel molecule generative model of V AE combined with transformer.arXiv [q-bio.BM], 2024
Yasuhiro Yoshikai, Tadahaya Mizuno, Shumpei Nemoto, and Hiroyuki Kusuhara. A novel molecule generative model of V AE combined with transformer.arXiv [q-bio.BM], 2024. doi:10.48550/arXiv.2402.11950
-
[41]
RoFormer: Enhanced Transformer with Rotary Position Embedding
Jianlin Su, Yu Lu, Shengfeng Pan, Ahmed Murtadha, Bo Wen, and Yunfeng Liu. RoFormer: Enhanced transformer with rotary position embedding.arXiv [cs.CL], 2021. doi:10.48550/arXiv.2104.09864
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2104.09864 2021
-
[42]
Aaron Grattafiori, Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Alex Vaughan, Amy Yang, Angela Fan, Anirudh Goyal, Anthony Hartshorn, Aobo Yang, Archi Mitra, Archie Sravankumar, Artem Korenev, Arthur Hinsvark, Arun Rao, Aston Zhang, Aurelien Rodriguez, Austen Gregerson, Ava S...
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2407.21783 2024
-
[43]
gpt-oss-120b & gpt-oss-20b Model Card
OpenAI, :, Sandhini Agarwal, Lama Ahmad, Jason Ai, Sam Altman, Andy Applebaum, Edwin Arbus, Rahul K Arora, Yu Bai, Bowen Baker, Haiming Bao, Boaz Barak, Ally Bennett, Tyler Bertao, Nivedita Brett, Eugene Brevdo, Greg Brockman, Sebastien Bubeck, Che Chang, Kai Chen, Mark Chen, Enoch Cheung, Aidan Clark, Dan Cook, Marat Dukhan, Casey Dvorak, Kevin Fives, Vl...
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2508.10925 2025
-
[44]
An Yang, Anfeng Li, Baosong Yang, Beichen Zhang, Binyuan Hui, Bo Zheng, Bowen Yu, Chang Gao, Chengen Huang, Chenxu Lv, Chujie Zheng, Dayiheng Liu, Fan Zhou, Fei Huang, Feng Hu, Hao Ge, Haoran Wei, Huan Lin, Jialong Tang, Jian Yang, Jianhong Tu, Jianwei Zhang, Jianxin Yang, Jiaxi Yang, Jing Zhou, Jingren Zhou, Junyang Lin, Kai Dang, Keqin Bao, Kexin Yang, ...
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2505.09388 2025
-
[45]
Gemma: Open Models Based on Gemini Research and Technology
Gemma Team, Thomas Mesnard, Cassidy Hardin, Robert Dadashi, Surya Bhupatiraju, Shreya Pathak, Laurent Sifre, Morgane Rivière, Mihir Sanjay Kale, Juliette Love, Pouya Tafti, Léonard Hussenot, Pier Giuseppe Sessa, Aakanksha Chowdhery, Adam Roberts, Aditya Barua, Alex Botev, Alex Castro-Ros, Ambrose Slone, Amélie Héliou, Andrea Tacchetti, Anna Bulanova, Anto...
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2403.08295 2024
-
[46]
Language models are unsupervised multitask learners
Radford, A., Wu, J., Child, R., Luan, D., Amodei, D., and Sutskever, I. Language models are unsupervised multitask learners. Technical report, OpenAI, 2019
work page 2019
-
[47]
GLU Variants Improve Transformer
Noam Shazeer. GLU variants improve transformer.arXiv [cs.LG], 2020. doi:10.48550/arXiv.2002.05202
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2002.05202 2020
-
[48]
BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding
Jacob Devlin, Ming-Wei Chang, Kenton Lee, and Kristina Toutanova. BERT: Pre-training of deep bidirectional transformers for language understanding.arXiv [cs.CL], 2018. doi:10.48550/arXiv.1810.04805
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1810.04805 2018
-
[49]
Scalable Diffusion Models with Transformers
William Peebles and Saining Xie. Scalable diffusion models with transformers.arXiv [cs.CV], 2022. doi:10.48550/arXiv.2212.09748
work page internal anchor Pith review doi:10.48550/arxiv.2212.09748 2022
-
[50]
Robin Winter, Floriane Montanari, Frank Noé, and Djork-Arné Clevert. Learning continuous and data-driven molecular descriptors by translating equivalent chemical representations.Chem. Sci., 10(6):1692–1701, 2019. ISSN 2041-6520,2041-6539. doi:10.1039/c8sc04175j
-
[51]
Yoshua Bengio, Jérôme Louradour, Ronan Collobert, and Jason Weston. Curriculum learning. InProceedings of the 26th Annual International Conference on Machine Learning, New York, NY , USA, 2009. ACM. ISBN 9781605585161. doi:10.1145/1553374.1553380
-
[52]
The road less scheduled.arXiv [cs.LG], 2024
Aaron Defazio, Xingyu Alice Yang, Harsh Mehta, Konstantin Mishchenko, Ahmed Khaled, and Ashok Cutkosky. The road less scheduled.arXiv [cs.LG], 2024. doi:10.48550/arXiv.2405.15682
-
[53]
Zclip: Adaptive spike mitigation for llm pre-training,
Abhay Kumar, Louis Owen, Nilabhra Roy Chowdhury, and Fabian Güra. ZClip: Adaptive spike mitigation for LLM pre-training.arXiv [cs.LG], 2025. doi:10.48550/arXiv.2504.02507
-
[54]
Simon Kornblith, Mohammad Norouzi, Honglak Lee, and Geoffrey Hinton. Similarity of neural network representations revisited.arXiv [cs.LG], 2019. doi:10.48550/arXiv.1905.00414
-
[55]
What happens during the loss plateau? understanding abrupt learning in transformers
Pulkit Gopalani and Wei Hu. What happens during the loss plateau? understanding abrupt learning in transformers. arXiv [cs.LG], 2025. doi:10.48550/arXiv.2506.13688
-
[56]
The geometric anatomy of capability acquisition in transformers.arXiv [cs.LG], 2026
Jayadev Billa. The geometric anatomy of capability acquisition in transformers.arXiv [cs.LG], 2026. doi:10.48550/arXiv.2602.15997
-
[57]
arXiv preprint arXiv:2309.07311 (2023)
Angelica Chen, Ravid Shwartz-Ziv, Kyunghyun Cho, Matthew L Leavitt, and Naomi Saphra. Sudden drops in the loss: Syntax acquisition, phase transitions, and simplicity bias in MLMs.arXiv [cs.CL], 2023. doi:10.48550/arXiv.2309.07311. 14 From Syntax to Semantics: Unveiling the Emergence of Chirality in SMILES Translation Models Tables and Figures Table 1: V o...
-
[58]
(a, b) Encoder; (c, d) Decoder
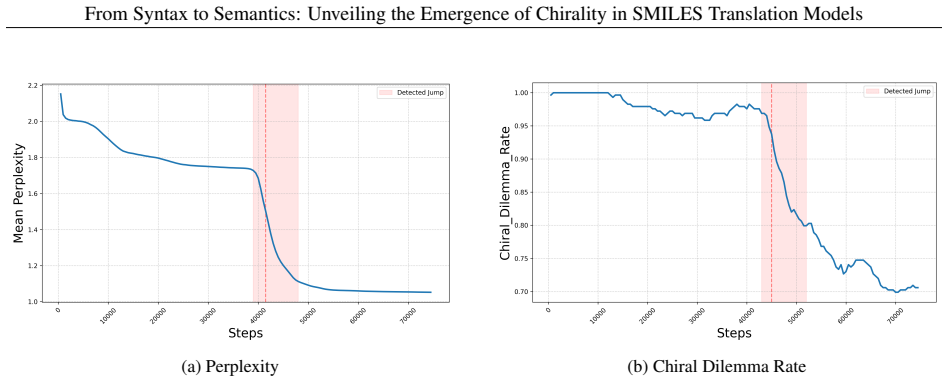
to L7H7, forpancore-addonceon ZINC20 ( ∼100)). (a, b) Encoder; (c, d) Decoder. (a, c) The region bounded by the two vertical dashed lines indicates the jump-up interval defined by perplexity. (b, d) Changes in the three heads with the largest entropy decrease within the jump-up interval. The red line and the light red shaded region indicate the perplexity...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.