Recognition: 1 theorem link
· Lean TheoremEstimating Consensus Epidemic Trajectories via a Constrained Power Fr\'echet Mean with Functional Registration
Pith reviewed 2026-05-12 04:37 UTC · model grok-4.3
The pith
Consensus epidemic trajectories are found by solving a constrained power Fréchet mean on exposed-infectious function pairs while enforcing differential-equation and population rules.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The consensus trajectory is defined as the solution of a constrained optimization problem that computes the power Fréchet mean of pairs of exposed and infectious compartment functions. Differential equation constraints and population constraints are incorporated directly into the optimization so that the infectious compartment retains a partial mechanistic interpretation. An efficient block-optimization algorithm based on functional data analysis is developed to obtain the solution, after which the remaining compartments and parameters are recovered to reconstitute the full SEIR dynamics.
What carries the argument
The constrained power Fréchet mean with functional registration applied to pairs of exposed and infectious compartment curves in Hilbert space.
If this is right
- Multiple SEIR solutions can be summarized into a single trajectory that still satisfies the underlying differential equations to a controllable degree.
- The full susceptible and removed compartment curves, together with the model parameters, can be recovered from the estimated exposed-infectious pair.
- The same optimization framework accommodates both mean-type and median-type estimators on the functional space.
- The resulting summaries can be used directly for model averaging or ensemble forecasting in infectious-disease applications.
Where Pith is reading between the lines
- The same constrained-mean construction could be applied to combine outputs from models that differ in structure as well as in parameter values.
- Because the optimization is block-wise, new trajectories can be added incrementally without recomputing the entire mean from scratch.
- The registration step that aligns curves before averaging may also be used to quantify disagreement among individual model runs.
Load-bearing premise
Incorporating differential equation constraints and population constraints into the Fréchet mean optimization preserves a partially mechanistic interpretation of the infectious compartment without distorting the consensus trajectory.
What would settle it
Apply the method to a collection of SEIR trajectories generated from known parameters; if the parameters recovered from the consensus trajectory differ substantially from the known inputs after accounting for registration, the claim that the constraints preserve mechanistic content fails.
Figures





read the original abstract
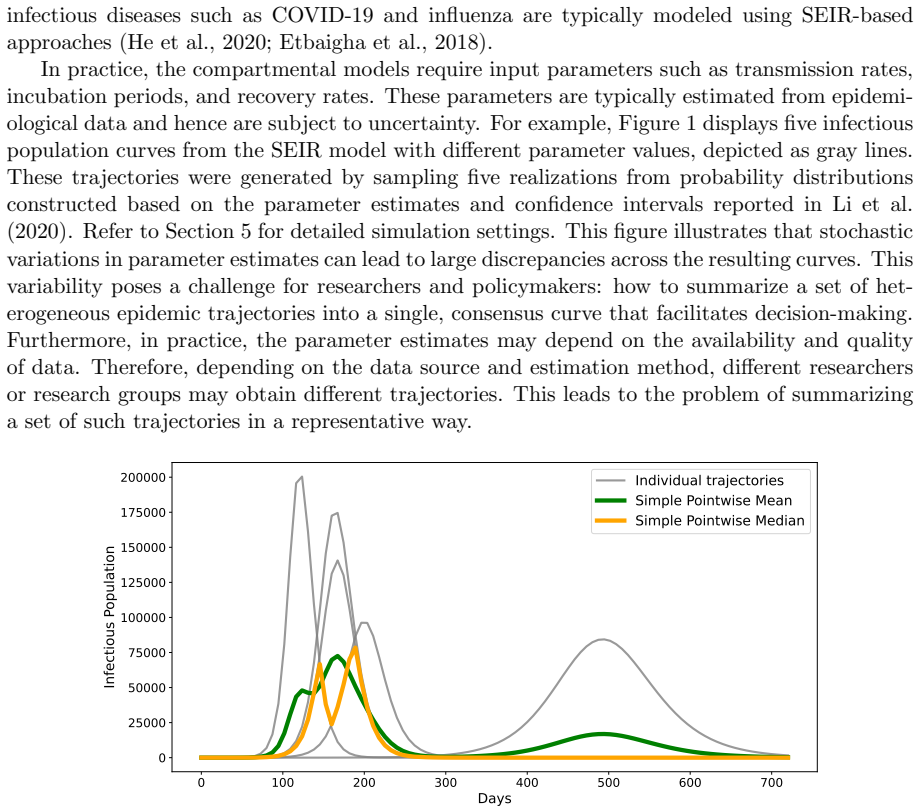
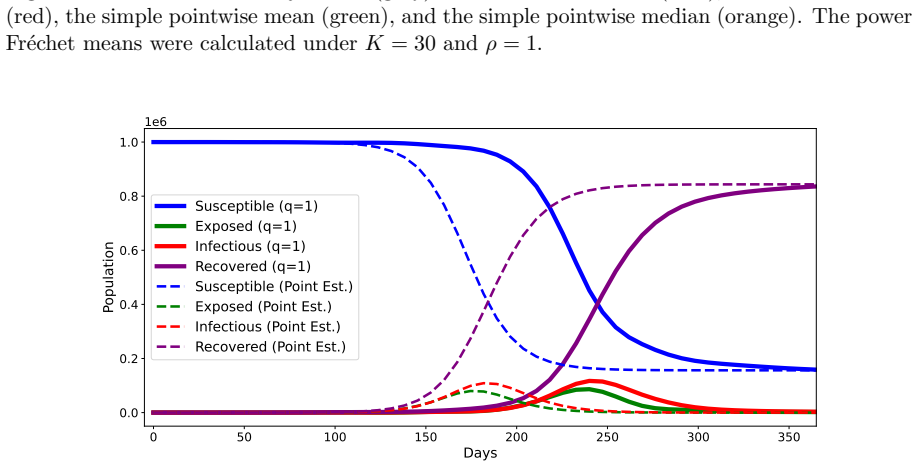
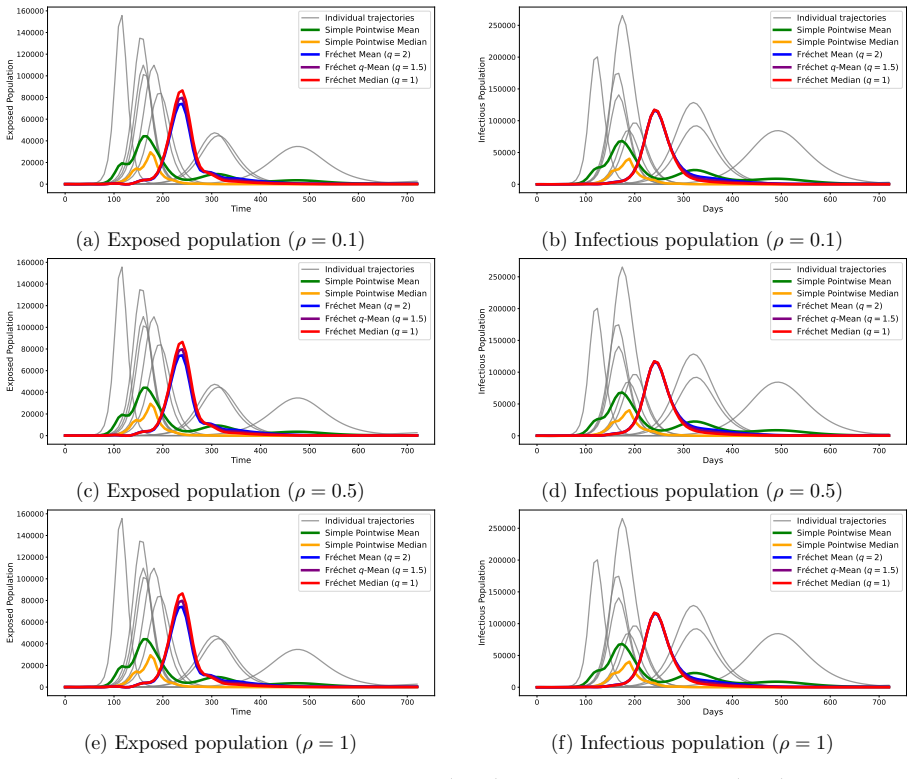
We propose a method for summarizing multiple solutions to SEIR-type compartmental models on a functional space by computing a constrained power Fr\'echet mean with functional registration to obtain consensus epidemic trajectories with partial mechanistic interpretability. In our method, we regard the pairs of exposed and infectious compartments as objects in a Hilbert space, and the consensus trajectory is defined as the solution to a constrained optimization problem. Differential equation constraints and population constraints are incorporated in the optimization to preserve a partially mechanistic interpretation regarding the infectious compartment. The full dynamics with additional susceptible and removed compartments can then be recovered from the estimated trajectories and parameters. We develop an efficient block-optimization algorithm based on functional data analysis and illustrate the method using simulated and literature-derived epidemiological parameters for COVID-19 in the early phase of the pandemic that began in 2020. The proposed approach provides a generalized trajectory-summarization framework that includes mean- and median-type estimators on a functional space and holds potential for model averaging and ensemble forecasting in infectious disease modeling.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proposes a method for summarizing multiple solutions to SEIR-type compartmental models by computing a constrained power Fréchet mean with functional registration on exposed-infectious compartment pairs treated as objects in a Hilbert space. Differential equation and population constraints are incorporated into the optimization to preserve partial mechanistic interpretability regarding the infectious compartment, after which the full dynamics (including susceptible and removed compartments) are recovered from the estimated trajectories and parameters. An efficient block-optimization algorithm based on functional data analysis is developed, and the approach is illustrated using simulated data and literature-derived COVID-19 parameters from early 2020. The method is positioned as a generalized trajectory-summarization framework encompassing mean- and median-type estimators with potential applications in model averaging and ensemble forecasting.
Significance. If the constraints ensure that recovered full SEIR trajectories remain consistent with the original ODE system, the work would provide a principled extension of Fréchet means and functional registration to constrained mechanistic models. This could support ensemble methods in epidemiology by enabling robust summarization of multiple epidemic trajectories while retaining some interpretability, which is a strength over purely data-driven averaging approaches.
major comments (2)
- [Abstract] Abstract: The central claim that 'Differential equation constraints and population constraints are incorporated in the optimization to preserve a partially mechanistic interpretation regarding the infectious compartment' and that 'The full dynamics with additional susceptible and removed compartments can then be recovered from the estimated trajectories and parameters' lacks any derivation, error analysis, or validation showing that the optimized (E,I) pair, after registration and parameter inference, satisfies the full four-equation SEIR system to within discretization error. This is load-bearing for the interpretability assertion.
- [Abstract] The optimization is performed only on the exposed-infectious subspace; it is not shown how the functional registration warping or power-mean weighting preserves the implicit balance between compartments required for the back-recovered S and R trajectories to remain consistent with the original SEIR ODEs. A concrete numerical check or proof of consistency is needed.
minor comments (1)
- The abstract would benefit from a brief statement of the specific form of the power Fréchet mean (e.g., the value of the power parameter) and the precise Hilbert space metric employed.
Simulated Author's Rebuttal
We thank the referee for their thoughtful and constructive comments on the manuscript. We appreciate the focus on rigorously establishing the mechanistic consistency of the recovered SEIR trajectories. We agree that the abstract claims require additional supporting derivations, error analysis, and numerical validation, which were not fully detailed. We will revise the manuscript accordingly to strengthen these aspects while preserving the core contribution of the constrained power Fréchet mean framework.
read point-by-point responses
-
Referee: [Abstract] Abstract: The central claim that 'Differential equation constraints and population constraints are incorporated in the optimization to preserve a partially mechanistic interpretation regarding the infectious compartment' and that 'The full dynamics with additional susceptible and removed compartments can then be recovered from the estimated trajectories and parameters' lacks any derivation, error analysis, or validation showing that the optimized (E,I) pair, after registration and parameter inference, satisfies the full four-equation SEIR system to within discretization error. This is load-bearing for the interpretability assertion.
Authors: We thank the referee for identifying this gap. The differential equation constraints (enforcing the relationship between E and I via the SEIR ODEs) and population constraints (conserving total population N) are incorporated directly into the constrained optimization problem defined in Section 3 of the manuscript. These ensure that the estimated (E,I) pair respects the infectious compartment dynamics, allowing recovery of S and R via the remaining equations and parameters. However, we acknowledge that a formal derivation of consistency for the full four-equation system (including error bounds relative to discretization) and explicit validation are not provided. In the revision, we will add a dedicated subsection deriving the recovery procedure and proving that the constrained solution satisfies the original ODEs up to the functional approximation error, along with numerical error analysis on the simulated data. revision: yes
-
Referee: [Abstract] The optimization is performed only on the exposed-infectious subspace; it is not shown how the functional registration warping or power-mean weighting preserves the implicit balance between compartments required for the back-recovered S and R trajectories to remain consistent with the original SEIR ODEs. A concrete numerical check or proof of consistency is needed.
Authors: The optimization operates on the (E,I) subspace in the Hilbert space, with functional registration applied to align phase variations across trajectories before computing the power Fréchet mean. The DE and population constraints are enforced within the optimization to preserve compartment balances for the infectious dynamics. The power-mean weighting occurs on the registered functions, and back-recovery of S and R uses the inferred parameters to complete the system. We agree that an explicit demonstration of how warping and weighting preserve the full ODE balances is absent. In the revised manuscript, we will include a proof sketch showing that if input trajectories satisfy the SEIR ODEs, the constrained mean does so within the Hilbert space projection, and add a concrete numerical check in the Results section (using both simulated and COVID-19 parameter examples) comparing recovered trajectories against the original ensemble for consistency within discretization error. revision: yes
Circularity Check
No significant circularity; method defined directly as constrained optimization without reduction to inputs.
full rationale
The paper defines the consensus epidemic trajectory explicitly as the solution to a constrained optimization problem on (E,I) pairs treated as Hilbert-space objects, with DE and population constraints incorporated by construction to support partial mechanistic recovery of S and R compartments. No derived quantity is obtained by fitting a parameter to a subset of data and then relabeling it as a prediction, nor does any central claim reduce to a self-citation chain or an ansatz smuggled from prior work. The approach is a self-contained definition of a summarization procedure on functional data; external benchmarks or independent verification of the recovered trajectories are not required for the definitional step itself to be non-circular.
Axiom & Free-Parameter Ledger
Lean theorems connected to this paper
-
IndisputableMonolith/Foundation/AbsoluteFloorClosure.leanreality_from_one_distinction unclearWe regard the pairs of exposed and infectious compartments as objects in a Hilbert space, and the consensus trajectory is defined as the solution to a constrained optimization problem. Differential equation constraints and population constraints are incorporated...
Reference graph
Works this paper leans on
-
[1]
Anderson, R. M. and May, R. M. , title =
-
[2]
Aron, J. L. and Schwartz, I. B. , title =. Journal of Theoretical Biology , volume =. 1984 , publisher =
work page 1984
- [3]
-
[4]
Bj. The. Nature Methods , volume =. 2020 , publisher =
work page 2020
-
[5]
Boschi, T. and Di Iorio, J. and Testa, L. and Cremona, M. A. and Chiaromonte, F. , title =. Scientific Reports , volume =. 2021 , publisher =
work page 2021
-
[6]
Chowell, G. and Luo, R. and Sun, K. and Roosa, K. and Tariq, A. and Viboud, C. , title =. Epidemics , volume =. 2020 , publisher =
work page 2020
-
[7]
Etbaigha, F. and Willms, A. R. and Poljak, Z. , title =. PLoS One , volume =. 2018 , publisher =
work page 2018
-
[8]
Fox, S. J. and Kim, M. and Meyers, L. A. and Reich, N. G. and Ray, E. L. , title =. Emerging Infectious Diseases , volume =
-
[9]
He, S. and Peng, Y. and Sun, K. , title =. Nonlinear Dynamics , volume =. 2020 , publisher =
work page 2020
-
[10]
He, X. and Lau, E. H. and Wu, P. and Deng, X. and Wang, J. and Hao, X. and Lau, Y. C. and Wong, J. Y. and Guan, Y. and Tan, X. and Mo, X. and Chen, Y. and Liao, B. and Chen, W. and Hu, F. and Zhang, Q. and Zhong, M. and Wu, Y. and Zhao, L. and Zhang, F. and Cowling, B. J. and Li, F. and Leung, G. M. , title =. Nature Medicine , volume =. 2020 , publisher =
work page 2020
-
[11]
Kermack, W. O. and McKendrick, A. G. , title =. Proceedings of the Royal Society of London. Series A, Containing Papers of a Mathematical and Physical Character , volume =. 1927 , publisher =
work page 1927
-
[12]
Kurisu, D. and Zhou, Y. and Otsu, T. and M. Geodesic causal inference , journal =
-
[13]
Kurisu, D. and Otsu, T. , title =. Journal of Multivariate Analysis , volume =. 2025 , publisher =
work page 2025
-
[14]
Li, Q. and Guan, X. and Wu, P. and Wang, X. and Zhou, L. and Tong, Y. and Ren, R. and Leung, K. S. and Lau, E. H. and Wong, J. Y. and Xing, X. and Xiang, N. and Wu, Y. and Li, C. and Chen, Q. and Li, D. and Liu, T. and Zhao, J. and Liu, M. and Tu, W. and Chen, C. and Jin, L. and Yang, R. and Wang, Q. and Zhou, S. and Wang, R. and Liu, H. and Luo, Y. and L...
work page 2020
-
[15]
McAndrew, T. and Reich, N. G. , title =. Statistics in Medicine , volume =. 2021 , publisher =
work page 2021
-
[16]
McGowan, C. J. and Biggerstaff, M. and Johansson, M. and Apfeldorf, K. M. and Ben-Nun, M. and Brooks, L. and Convertino, M. and Erraguntla, M. and Farrow, D. C. and Freeze, J. and Ghosh, S. and Hyun, S. and Kandula, S. and Lega, J. and Liu, Y. and Michaud, N. and Morita, H. and Niemi, J. and Ramakrishnan, N. and Ray, E. L. and Reich, N. G. and Riley, P. a...
work page 2019
-
[17]
Petersen, A. and M. Fr. The Annals of Statistics , volume =. 2019 , publisher =
work page 2019
-
[18]
Ray, E. L. and Reich, N. G. , title =. PLoS Computational Biology , volume =. 2018 , publisher =
work page 2018
-
[19]
Ray, E. L. and Brooks, L. C. and Bien, J. and Biggerstaff, M. and Bosse, N. I. and Bracher, J. and Cramer, E. Y. and Funk, S. and Gerding, A. and Johansson, M. A. and Rumack, A. and Wang, Y. and Zorn, M. and Tibshirani, R. J. and Reich, N. G. , title =. International Journal of Forecasting , volume =. 2023 , publisher =
work page 2023
-
[20]
Reich, N. G. and McGowan, C. J. and Yamana, T. K. and Tushar, A. and Ray, E. L. and Osthus, D. and Kandula, S. and Brooks, L. C. and Crawford-Crudell, W. and Gibson, G. C. and Moore, E. and Silva, R. and Biggerstaff, M. and Johansson, M. A. and Rosenfeld, R. and Shaman, J. , title =. PLoS Computational Biology , volume =. 2019 , publisher =
work page 2019
-
[21]
Taylor, K. S. and Taylor, J. W. , title =. PLoS One , volume =. 2022 , publisher =
work page 2022
-
[22]
Taylor, J. W. and Taylor, K. S. , title =. European Journal of Operational Research , volume =. 2023 , publisher =
work page 2023
-
[23]
Wang, J.-L. and Chiou, J.-M. and M. Functional data analysis , journal =. 2016 , publisher =
work page 2016
-
[24]
White, P. J. and Enright, M. C. , title =. Infectious Diseases (Third Edition) , pages =. 2010 , publisher =
work page 2010
-
[25]
Yamana, T. K. and Kandula, S. and Shaman, J. , title =. Journal of the Royal Society Interface , volume =. 2016 , publisher =
work page 2016
-
[26]
Yamana, T. K. and Kandula, S. and Shaman, J. , title =. PLoS Computational Biology , volume =. 2017 , publisher =
work page 2017
-
[27]
Functional Analysis, Sobolev Spaces and Partial Differential Equations , publisher =
Br. Functional Analysis, Sobolev Spaces and Partial Differential Equations , publisher =
-
[28]
Griette, Q. and Demongeot, J. and Magal, P. , title =. Mathematical Biosciences and Engineering , volume =. 2022 , issn =. doi:10.3934/mbe.2022025 , url =
-
[29]
Weitz, J. S. and Dushoff, J. , title =. Scientific Reports , volume =. 2015 , publisher =
work page 2015
-
[30]
Gerberry, D. J. and Milner, F. A. , title =. Journal of Mathematical Biology , volume =. 2009 , publisher =
work page 2009
-
[31]
Dubey, P. and Chen, Y. and M. Metric statistics: Exploration and inference for random objects with distance profiles , journal =. 2024 , publisher =
work page 2024
-
[32]
Peter hall, functional data analysis and random objects , journal =
M. Peter hall, functional data analysis and random objects , journal =. 2016 , publisher =
work page 2016
-
[33]
Ramsay, J. O. and Silverman, B. W. , title =. 2005 , edition =. doi:10.1007/b98888 , isbn =
- [34]
-
[35]
Strong laws of large numbers for generalizations of Fr
Sch. Strong laws of large numbers for generalizations of Fr. Statistics , volume =. 2022 , publisher =
work page 2022
-
[36]
Allen, K. and Parry, A. E. and Glass, K. , title =. Western Pacific Surveillance and Response Journal: WPSAR , volume =
-
[37]
Wan, K. and Chen, J. and Lu, C. and Dong, L. and Wu, Z. and Zhang, L. , title =. Journal of Global Health , volume =
-
[38]
Dagpunar, J. S. , title =. Infectious Disease Modelling , volume =. 2020 , publisher =
work page 2020
-
[39]
Carcione, J. M. and Santos, J. E. and Bagaini, C. and Ba, J. , title =. Frontiers in Public Health , volume =. 2020 , publisher =
work page 2020
-
[40]
Peirlinck, M. and Linka, K. and Sahli Costabal, F. and Kuhl, E. , title =. Biomechanics and Modeling in Mechanobiology , volume =. 2020 , publisher =
work page 2020
-
[41]
Wu, J. T. and Leung, K. and Leung, G. M. , title =. The Lancet , volume =. 2020 , publisher =
work page 2020
-
[42]
Tang, B. and Wang, X. and Li, Q. and Bragazzi, N. L. and Tang, S. and Xiao, Y. and Wu, J. , title =. Journal of Clinical Medicine , volume =. 2020 , publisher =
work page 2020
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.