Recognition: 2 theorem links
· Lean Theoremgemlib.mcmc: composable kernels for Metropolis-within-Gibbs sampling schemes
Pith reviewed 2026-05-12 03:46 UTC · model grok-4.3
The pith
Writer monads enable automatic composition of kernels for Metropolis-within-Gibbs sampling.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
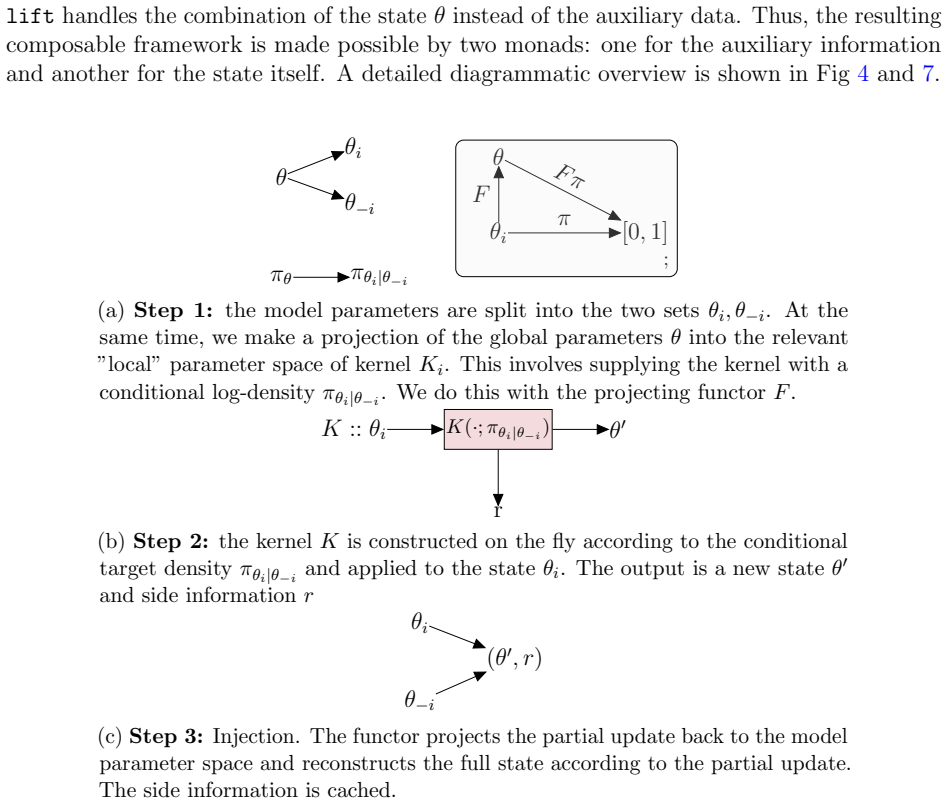
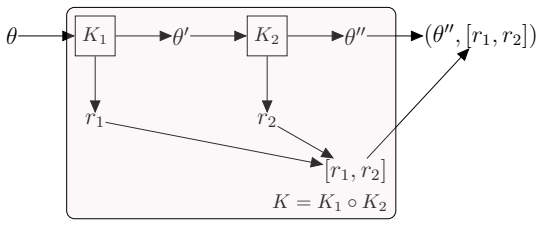
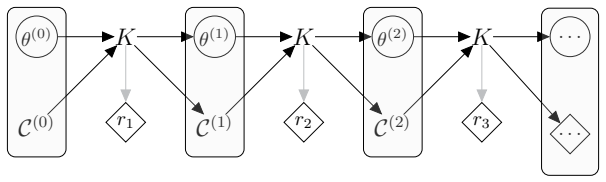
The framework formalizes kernel composition using writer monads from category theory, allowing parameter-estimation kernels and data-augmentation kernels to integrate directly into joint sampling schemes for models with intractable likelihoods and missing data, with the composition logic, state management, and performance handling all automated so that developers define only two methods per new kernel.
What carries the argument
The writer monad, a category-theoretic structure that tracks and propagates state updates across successive kernel applications without explicit user-managed bookkeeping.
If this is right
- Complex Metropolis-within-Gibbs procedures for partially observed models can be expressed as concise chains of kernel operations.
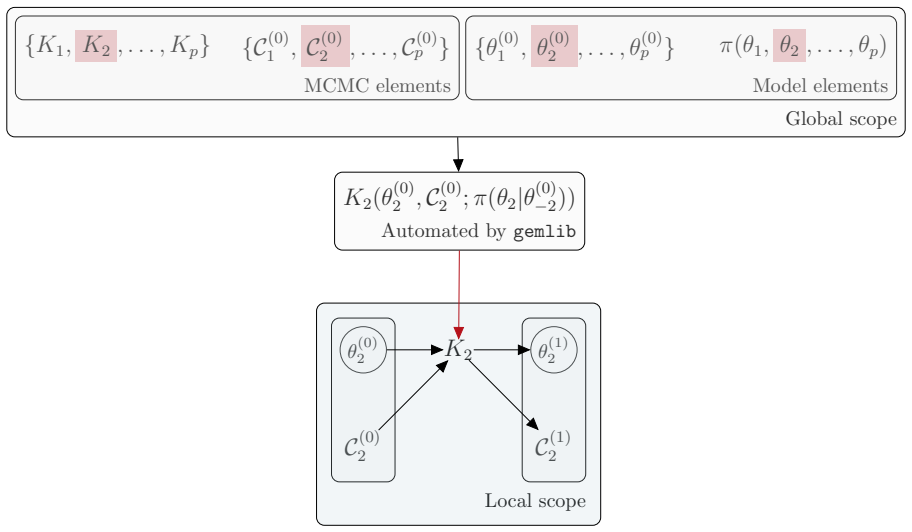
- New kernels require implementation of only two methods, after which composition and reuse become automatic.
- Hardware acceleration of the resulting samplers occurs without additional user code.
- Kernels developed for one epidemic or ecological application can be reused in others without re-deriving the composition logic.
Where Pith is reading between the lines
- The same abstraction pattern could reduce coding effort for other families of iterative statistical algorithms beyond Metropolis-within-Gibbs.
- Applied modelers might shift time from sampler engineering toward domain-specific questions about latent processes.
- Empirical tests on diverse real datasets would reveal whether the reduced code volume also lowers the rate of subtle sampling errors.
Load-bearing premise
The monad-based composition produces a mathematically valid overall transition kernel that preserves the required statistical properties of the individual kernels.
What would settle it
A concrete counterexample in which two individually valid kernels, when composed through the framework, yield a chain whose stationary distribution differs from the target posterior would falsify the claim.
Figures







read the original abstract
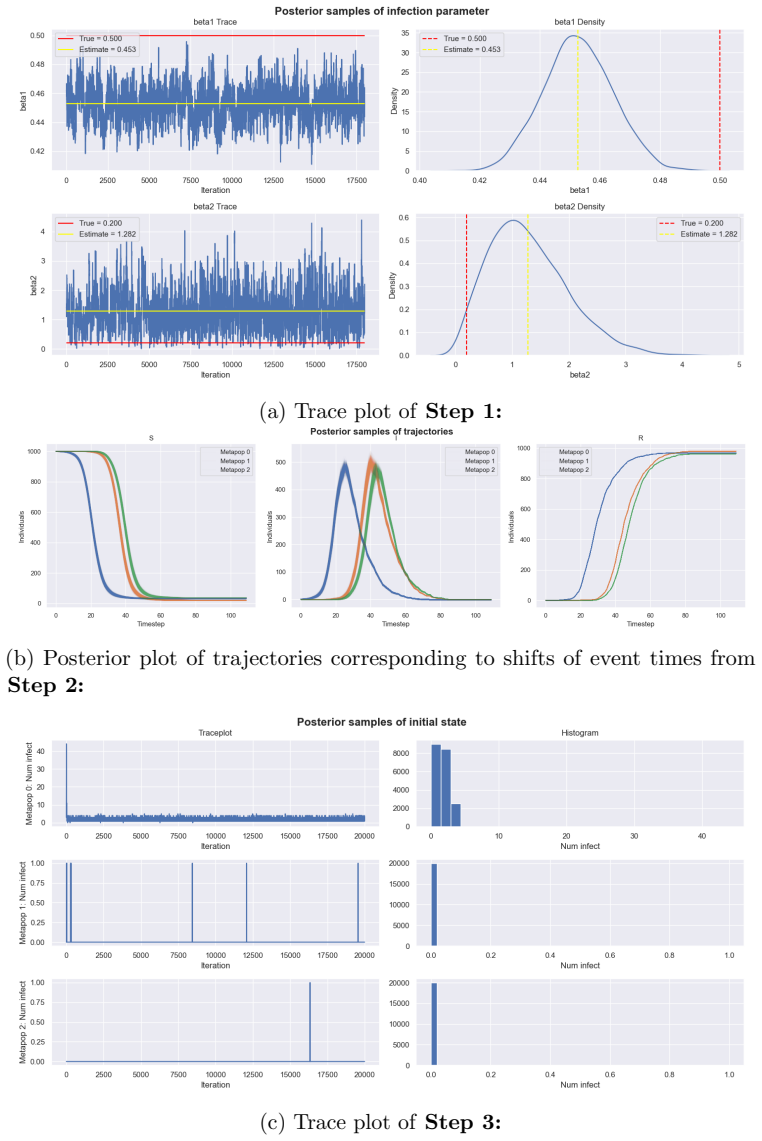
State-transition models are essential across epidemiology and ecology, but statistical inference remains challenging owing to high-dimensional latent state spaces, temporal dependence, and intractable likelihood functions. Bayesian inference via Markov Chain Monte Carlo (MCMC) enables joint estimation of model parameters and missing event times through data augmentation, but Metropolis-within-Gibbs (MWG) schemes that combine multiple specialised kernels are notoriously difficult to implement. Current probabilistic programming frameworks face a trade-off: automation sacrifices extensibility, whilst flexibility demands substantial implementation overhead. This divide has created a software landscape characterised by tightly coupled, model-specific implementations that resist reuse and extension. We introduce gemlib.mcmc, an MCMC module designed to bridge methodological and applied communities through principled, composable kernel abstractions. The framework employs writer monads from category theory to formalise kernel composition, enabling seamless integration of parameter-estimation and data-augmentation kernels without manual state management. Built on JAX and TensorFlow Probability for high-performance computation, gemlib.mcmc provides an ergonomic interface -- leveraging Python's right-shift operator for intuitive kernel chaining -- whilst maintaining statistical rigour and transparency. Developers can extend the library by implementing only two methods; composition and hardware acceleration are automated. We demonstrate the framework through parameter inference on partially observed epidemic models, showing how complex inference algorithms can be expressed concisely and reused across applications. By reducing implementation burden we provide access to sophisticated MCMC methods and enable applied researchers to employ state-of-the-art algorithms without reimplementation overhead.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces gemlib.mcmc, a Python library for Metropolis-within-Gibbs MCMC that uses writer monads to formalize composable kernels. It claims to enable seamless integration of parameter-estimation and data-augmentation kernels via an ergonomic interface (Python right-shift operator), with users needing to implement only two methods for extension; composition, state management, and JAX/TFP hardware acceleration are automated while preserving statistical rigour. The framework is demonstrated through parameter inference on partially observed epidemic models, with the goal of reducing implementation burden for complex MWG schemes in epidemiology and ecology.
Significance. If the monad-based composition is shown to preserve detailed balance and stationary distributions, the library could meaningfully lower barriers for applied researchers implementing data-augmentation MCMC in high-dimensional state-transition models, promoting reuse across applications and reducing manual state-management errors.
major comments (1)
- Abstract: the central claim that the two-method extension interface 'maintains statistical rigour and transparency' while automating composition is unsupported; the manuscript provides no proofs of correctness, numerical validation against standard MWG implementations, benchmarks, or error analysis to confirm that kernel chaining preserves the target distribution.
minor comments (2)
- The manuscript would benefit from explicit code examples showing the right-shift operator for kernel chaining and the two required extension methods.
- Clarify the precise interface contract (method signatures and expected return types) for the two extension methods to aid reproducibility.
Simulated Author's Rebuttal
We thank the referee for their constructive review and for identifying a key area where the manuscript's claims require stronger support. We address the major comment below and will incorporate the necessary revisions to strengthen the presentation of the framework's statistical properties.
read point-by-point responses
-
Referee: [—] Abstract: the central claim that the two-method extension interface 'maintains statistical rigour and transparency' while automating composition is unsupported; the manuscript provides no proofs of correctness, numerical validation against standard MWG implementations, benchmarks, or error analysis to confirm that kernel chaining preserves the target distribution.
Authors: We agree that the manuscript as submitted does not contain explicit proofs of correctness for the monad-based composition or numerical validation against reference MWG implementations. The abstract's phrasing regarding maintenance of statistical rigour is therefore not fully substantiated in the current text. In the revised manuscript we will add a new section (likely Section 3 or an appendix) that formally shows preservation of detailed balance: if each constituent kernel satisfies detailed balance with respect to its target conditional distribution, then the writer-monad composition (via the right-shift operator) yields a kernel that satisfies detailed balance with respect to the joint target. The proof will rely on the standard properties of the writer monad and the fact that state updates are performed only through the monadic bind. We will also include a short numerical validation study on a low-dimensional Gaussian target and on a simple SIR epidemic model, comparing posterior samples and acceptance rates obtained from gemlib.mcmc kernels against hand-written Metropolis-within-Gibbs code. Benchmarks measuring wall-clock time and effective sample size per second, together with a brief error analysis (e.g., total-variation distance to a long-run reference chain), will be added to the results section. revision: yes
Circularity Check
No significant circularity
full rationale
The paper introduces a software library (gemlib.mcmc) that uses writer monads to compose MCMC kernels, with users implementing only two methods for extension. No equations, fitted parameters, predictions, or first-principles derivations are present that could reduce to inputs by construction. The presentation is an implementation and interface description rather than a mathematical derivation chain; category theory concepts are adopted as external tools without self-referential reduction. No self-citation load-bearing steps, ansatz smuggling, or renaming of known results occur in the provided text.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Writer monads from category theory can formalise kernel composition in MCMC without manual state management
Lean theorems connected to this paper
-
IndisputableMonolith/Foundation/ArithmeticFromLogic.leanLogicNat recovery and embed_injective unclearThe framework employs writer monads from category theory to formalise kernel composition... Developers can extend the library by implementing only two methods
-
IndisputableMonolith/Foundation/AbsoluteFloorClosure.leanabsolute_floor_iff_bare_distinguishability uncleargemlib.mcmc provides an ergonomic interface... for parameter inference on partially observed epidemic models
Reference graph
Works this paper leans on
-
[1]
Sampling-Based Approaches to Calculating Marginal Densities
Alan E. Gelfand and Adrian F. M. Smith. “Sampling-Based Approaches to Calculating Marginal Densities”. In:Journal of the American Statistical Association85.410 (1990), pp. 398–409.issn: 01621459, 1537274X.url: http : / / www . jstor . org / stable / 2289776(visited on 02/27/2026)
work page 1990
-
[2]
Christian P Robert and George Casella.Monte Carlo statistical methods. Vol. 2. Springer, 2004
work page 2004
-
[3]
Brooks et al.Handbook of Markov Chain Monte Carlo
S. Brooks et al.Handbook of Markov Chain Monte Carlo. Chapman & Hall/CRC Handbooks of Modern Statistical Methods. CRC Press, 2011.isbn: 9781420079425. url:https://books.google.co.uk/books?id=qfRsAIKZ4rIC
work page 2011
-
[4]
The No-U-Turn sampler: adaptively setting path lengths in Hamiltonian Monte Carlo
Matthew D Hoffman, Andrew Gelman, et al. “The No-U-Turn sampler: adaptively setting path lengths in Hamiltonian Monte Carlo.” In:J. Mach. Learn. Res.15.1 (2014), pp. 1593–1623
work page 2014
-
[5]
H. I. Freedman.Deterministic Mathematical Models in Population Ecology. Vol. 57. Pure and Applied Mathematics: A Series of Monographs and Textbooks. New York: Marcel Dekker, 1980, p. 254.isbn: 0-8247-6653-9
work page 1980
-
[6]
A contribution to the mathematical theory of epidemics
WO Kermack and AG McKendrick. “A contribution to the mathematical theory of epidemics”. In:Proc. R. Soc. Lond. A115 (1927), pp. 700–721
work page 1927
-
[7]
JM Read et al. “Novel coronavirus 2019-nCoV (COVID-19): early estimation of epi- demicological parameters and epidemic size estimates”. In:Philos Trans R Soc Lond B Biol Sci376 (1829 2021), p. 20200265
work page 2019
-
[8]
Philip D O’Neill. “A tutorial introduction to Bayesian inference for stochastic epidemic models using Markov chain Monte Carlo methods”. en. In:Math. Biosci.180.1-2 (Nov. 2002), pp. 103–114
work page 2002
-
[9]
PLOS Com- putational Biology18(9), 1010492 (2022) https://doi.org/10.1371/journal.pcbi
Sam Moore et al. “Modelling optimal vaccination strategy for SARS-CoV-2 in the UK”. In:PLOS Computational Biology17.5 (2021), e1008849.doi: 10.1371/journal.pcbi. 1008849.url:https://doi.org/10.1371/journal.pcbi.1008849
-
[10]
Cambridge University Press, 2024.url:https://arxiv.org/pdf/2407.12751
Paul Fearnhead et al.Scalable Monte Carlo for Bayesian Learning. Cambridge University Press, 2024.url:https://arxiv.org/pdf/2407.12751
-
[11]
F. W. Lawvere.The category of probabilistic mappings - With applications to stocahstic process, statistics, and pattern recognition. 1962.url: https://ncatlab.org/nlab/ files/lawvereprobability1962.pdf
work page 1962
-
[12]
Monad transformers and modular inter- preters
Sheng Liang, Paul Hudak, and Mark Jones. “Monad transformers and modular inter- preters”. In:Proceedings of the 22nd ACM SIGPLAN-SIGACT Symposium on Principles of Programming Languages. POPL ’95. San Francisco, California, USA: Association for Computing Machinery, 1995, 333–343.isbn: 0897916921.doi: 10.1145/199448.199528. url:https://doi.org/10.1145/199448.199528
- [13]
- [14]
-
[15]
Apache Software Foundation.OpenXLA Project. Feb. 19, 2010.url: https://github. com/openxla/xla
work page 2010
-
[16]
School of Haskell, FP Complete
Bartosz Milewski.Basics of Haskell. School of Haskell, FP Complete. Archived from the original on 2016-10-27. Retrieved 2018-07-13. 2013
work page 2016
-
[17]
Functional Programming with Overloading and Higher-Order Polymor- phism
Mark P. Jones. “Functional Programming with Overloading and Higher-Order Polymor- phism”. In:Advanced Functional Programming. Ed. by Johan Jeuring and Erik Meijer. Vol. 925. Lecture Notes in Computer Science. Berlin, Heidelberg: Springer-Verlag, May 1995
work page 1995
-
[18]
gemlib - probabilistic programming for epidemic models
Alin Morariu, Jess Bridgen, and C.P. Jewell. “gemlib - probabilistic programming for epidemic models”. In:arXiv preprint arXiv(2025).url: https://arxiv.org/abs/ 2511.08124
-
[19]
Gareth O. Roberts and Jeffrey S. Rosenthal. “Examples of Adaptive MCMC”. In: Journal of Computational and Graphical Statistics18.2 (2009), pp. 349–367.doi: 10.1198/jcgs.2009.06134 . eprint: https://doi.org/10.1198/jcgs.2009.06134 . url:https://doi.org/10.1198/jcgs.2009.06134
-
[20]
Bayesian Analysis for Emerging Infectious Diseases
Chris Jewell et al. “Bayesian Analysis for Emerging Infectious Diseases”. In:Bayesian Analysis4 (Sept. 2009), pp. 465–496.doi:10.1214/09-BA417
-
[21]
A case study in non-centering for data augmentation: Stochastic epidemics
Peter Neal and Gareth Roberts. “A case study in non-centering for data augmentation: Stochastic epidemics”. In:Statistics and Computing15 (2005), pp. 315–327
work page 2005
-
[22]
PyMC: A Modern and Comprehensive Probabilistic Programming Framework in Python
Oriol Abril-Pla et al. “PyMC: A Modern and Comprehensive Probabilistic Programming Framework in Python”. In:PeerJ Computer Science9.e1516 (2023).doi: 10.7717/ peerj-cs.1516
work page 2023
-
[23]
Programming With Models: Writing Statistical Algorithms for General Model Structures With NIMBLE
Perry de Valpine et al. “Programming With Models: Writing Statistical Algorithms for General Model Structures With NIMBLE”. In:Journal of Computational and Graphical Statistics26.2 (Apr. 2017), 403–413.issn: 1537-2715.doi: 10.1080/10618600.2016. 1172487.url:http://dx.doi.org/10.1080/10618600.2016.1172487
-
[24]
Real-time decision-making during emergency disease outbreaks
William J. M. Probert et al. “Real-time decision-making during emergency disease outbreaks”. In:PLOS Computational Biology14.7 (July 2018), pp. 1–18.doi: 10.1371/ journal.pcbi.1006202.url:https://doi.org/10.1371/journal.pcbi.1006202
-
[25]
Jessica R. E. Bridgen et al. “A Bayesian approach to identifying the role of hospital structure and staff interactions in nosocomial transmission of SARS-CoV-2”. In:Journal of The Royal Society Interface21.212 (Mar. 2024), p. 20230525.issn: 1742-5689.doi: 10.1098/rsif.2023.0525 . eprint: https://royalsocietypublishing.org/rsif/ article-pdf/doi/10.1098/rsi...
-
[26]
Christopher N Davis et al. “A modelling assessment for the impact of control measures on highly pathogenic avian influenza transmission in poultry in Great Britain”. In: bioRxiv(2025).doi: 10.1101/2025.04.24.650264 . eprint: https://www.biorxiv. org / content / early / 2025 / 04 / 25 / 2025 . 04 . 24 . 650264 . full . pdf.url: https : //www.biorxiv.org/co...
-
[27]
The Journal of Chemical Physics21(6), 1087–1092 (1953) https://doi.org/10.1063/1.1699114
Nicholas Metropolis et al. “Equation of State Calculations by Fast Computing Machines”. In:The Journal of Chemical Physics21.6 (June 1953), pp. 1087–1092.issn: 0021- 9606.doi: 10.1063/1.1699114. eprint: https://pubs.aip.org/aip/jcp/article- pdf/21/6/1087/18802390/1087\_1\_online.pdf.url: https://doi.org/10.1063/ 1.1699114
-
[28]
Monte Carlo Sampling Methods Using Markov Chains and Their Applications
W. K. Hastings. “Monte Carlo Sampling Methods Using Markov Chains and Their Applications”. In:Biometrika57.1 (1970), pp. 97–109.issn: 00063444, 14643510.url: http://www.jstor.org/stable/2334940(visited on 07/01/2025)
-
[29]
Coupling and Ergodicity of adaptive Markov chain Monte Carlo algorithms
Gareth Roberts and Jeffrey Rosenthal. “Coupling and Ergodicity of adaptive Markov chain Monte Carlo algorithms”. In:Journal of Applied Probability44 (2007), pp. 458– 475
work page 2007
-
[30]
An adaptive Metropolis algorithm
H. Haario, E. Saksman, and J. Tamminen. “An adaptive Metropolis algorithm”. In: Bernoulli7.2 (2001), pp. 223–242
work page 2001
-
[31]
SGMCMCJax: a lightweight JAX library for stochastic gradient Markov chain Monte Carlo algorithms
Jeremie Coullon and Christopher Nemeth. “SGMCMCJax: a lightweight JAX library for stochastic gradient Markov chain Monte Carlo algorithms”. In:Journal of Open Source Software7.72 (2022), p. 4113. 38
work page 2022
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.