Recognition: no theorem link
Morpho-Physiological and Genetic Diversity of Crataegus Taxa (Rosaceae) in Selected Locations of Iraqi Kurdistan-Region
Pith reviewed 2026-05-13 01:17 UTC · model grok-4.3
The pith
Seven Crataegus taxa in Iraqi Kurdistan show high variability in eleven fruit traits among 61 accessions.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The authors establish that the Kurdistan region hosts seven Crataegus taxa, five species and two hybrids, and that analysis of variance demonstrates significant variability (P < 0.01) in physio-morphological fruit data across the 61 accessions, with eleven variables explaining the majority of the variation observed.
What carries the argument
Phenotypic markers combined with measurements of fruit and seed physio-morphological traits, including weight, dimensions, seed count, pH and moisture, to reveal diversity and delimit taxa.
If this is right
- The seven taxa can be observed and distinguished in the studied locations based on plant type, reproductive stage, and fruit morphology.
- Eleven specific fruit variables show significant differences and capture most of the variability among accessions.
- The ecotypes exhibit variation suitable for different production uses.
- Hybrid forms exist between Crataegus azarolus and other species.
- Morphological and molecular systems together prove useful for assessing genetic diversity in hawthorns.
Where Pith is reading between the lines
- Similar diversity surveys could help prioritize conservation of wild fruit trees in other semi-arid regions.
- The eleven key variables might form a practical checklist for characterizing hawthorn accessions elsewhere.
- Recognizing hybrids suggests gene flow occurs and could influence breeding programs for improved varieties.
- Further genetic analysis might clarify the exact relationships among the observed taxa.
Load-bearing premise
That the morphological characters and limited molecular markers used are adequate to reliably identify and separate the seven taxa without misclassifying individuals or overlooking additional variation.
What would settle it
A follow-up study that re-collects samples from the same locations, applies full genome sequencing, and finds that the accessions do not form distinct clusters matching the seven taxa or that the eleven traits do not differ significantly.
Figures














read the original abstract
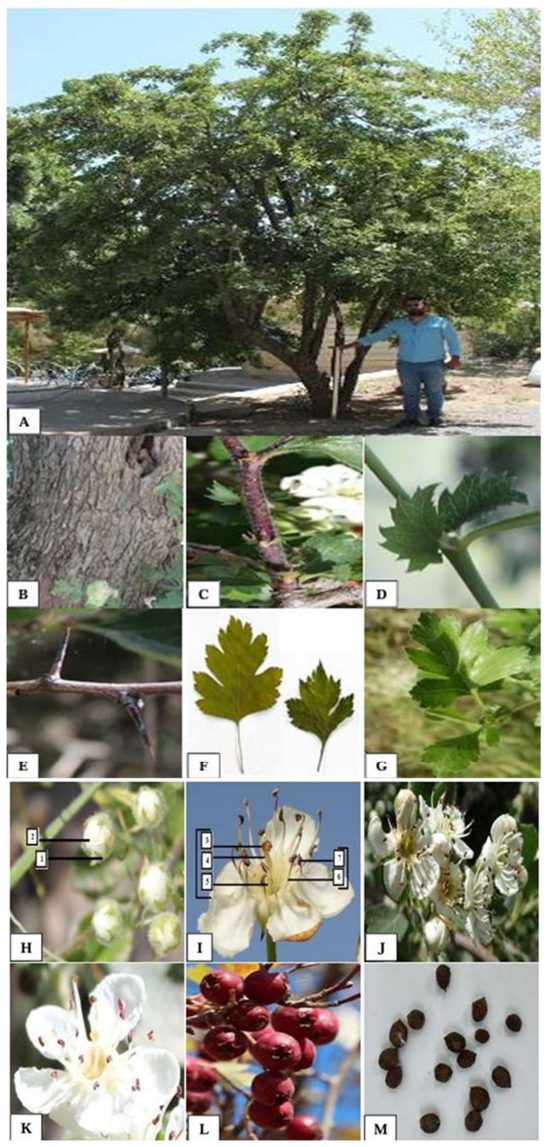
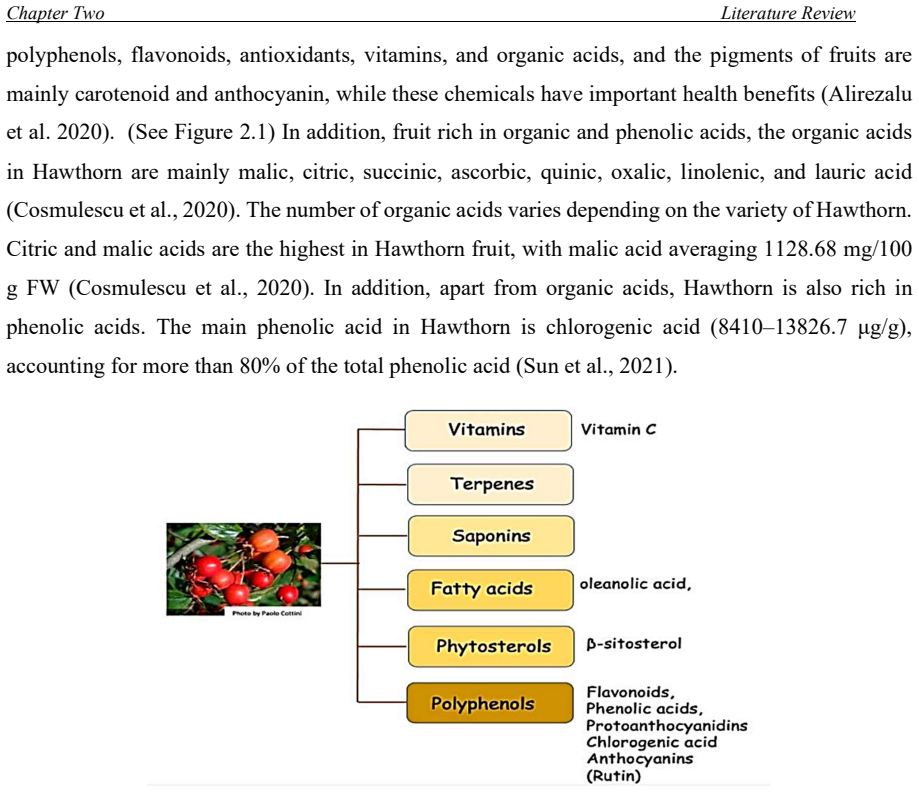
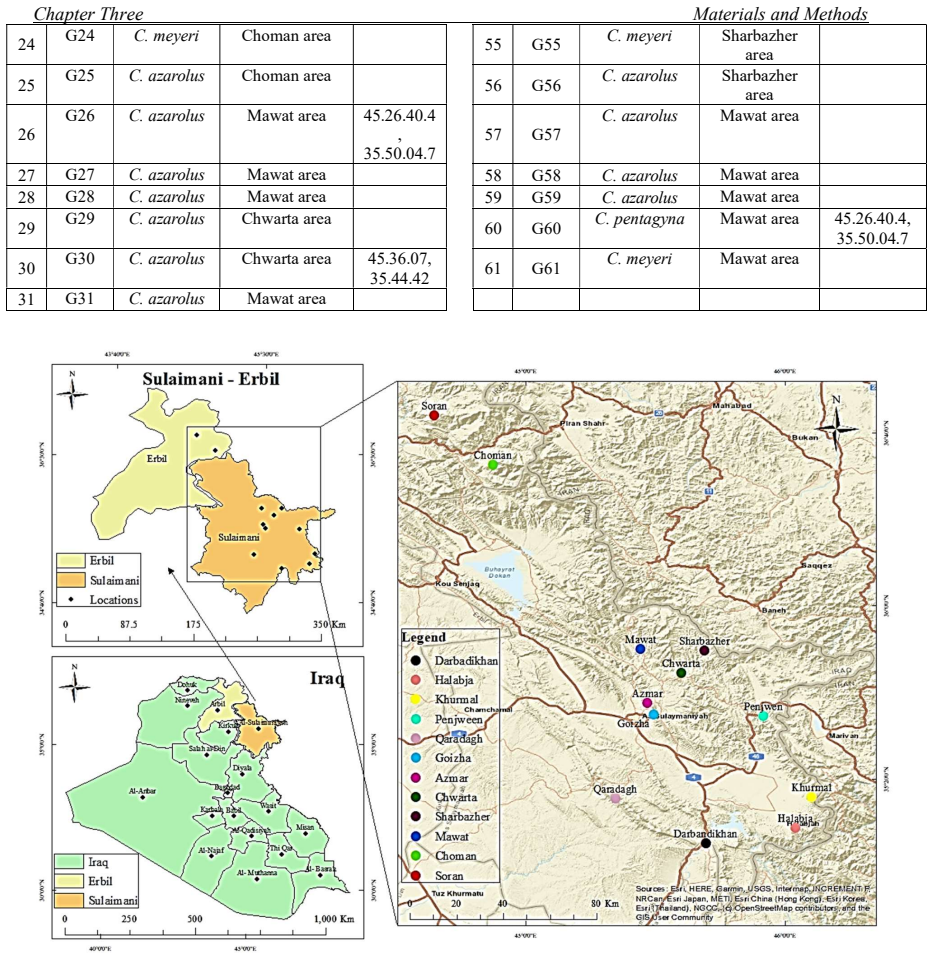
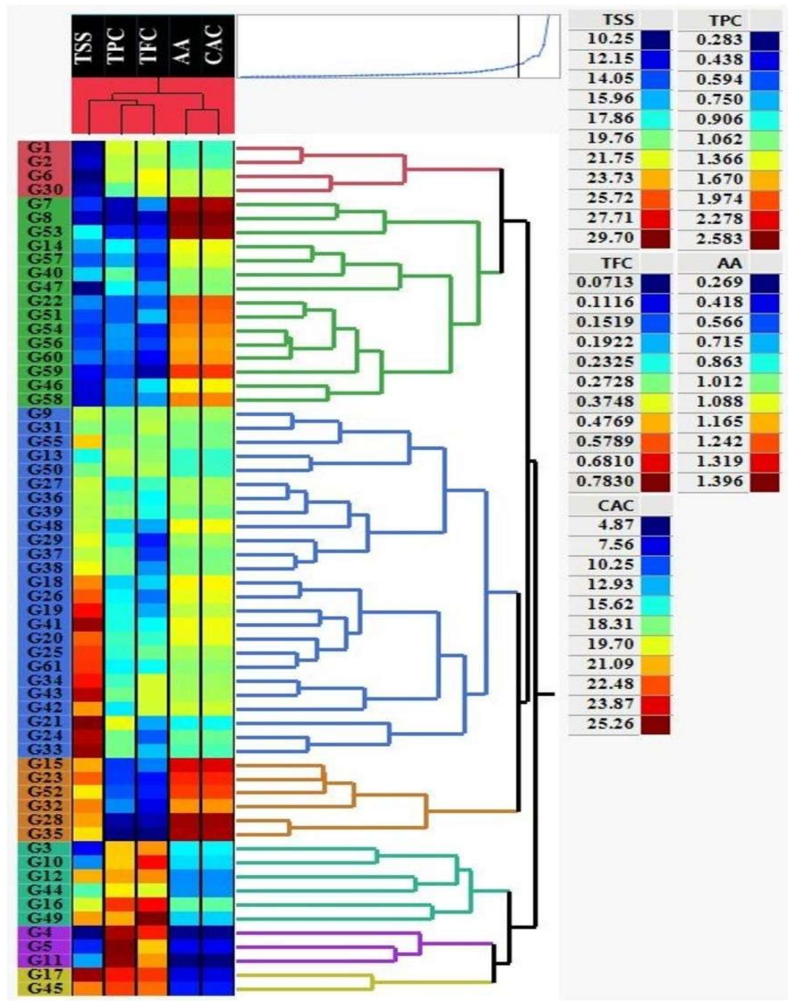
One of the great phytogeography zones of semi-arid lands in the world is the Kurdistan region of Iraq which hosts many important fruit species due to its geographical location and ecology. Mountain Hawthorn (Crataegus spp.) is a vital wild edible deciduous fruit tree of the genus Crataegus for the region, which is highly beneficial for ornamental, economical, industrial and medicinal uses. In the present study, morphological, phytochemical and molecular marker systems were applied on sixty-one Hawthorn accessions from different locations in the Iraqi Kurdistan region during April 2022 to September 2023. Phenotypic markers have proven to be extremely useful in studies of genetic diversity in Hawthorn genotypes, the results of the present morphological study showed that there are seven taxa (five species, two hybrids) were observed including, Crataegus azarolus, Crataegus meyrei, Crataegus monogyna, Crataegus orientalists, Crataegus pentagyna, Crataegus azarolus x Crataegus meyrei and Crataegus azarolus x Crataegus pentagyna. There was significant variation among different ecotypes in terms of plant type, reproductive stage, and fruit morphology and production uses. Fruit Physio-morphological data revealed a high level of significant variability (P 0.01) among accessions based on the analysis of variance. The most important characteristics for explaining fruit morphological variability `were 11 varbales including fruit weight (FW), fruit length (FL), fruit width (FW), seed length (SL), seed width (SW), number of seeds per fruits (NSF), volume solution (VS), fruit fresh weight (WOF), seed weight (WS), Potentional of hydrogen (pH) and mositure content (MC). They all are significantly different for all the traits measured for the studied accessions.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript presents results from a field survey of 61 Crataegus accessions collected from various locations in the Iraqi Kurdistan region. Using morphological, phytochemical, and molecular marker analyses, the authors identify seven taxa, including five species (C. azarolus, C. meyrei, C. monogyna, C. orientalis, C. pentagyna) and two hybrids (C. azarolus × C. meyrei and C. azarolus × C. pentagyna). They report statistically significant variation (P < 0.01) in fruit physio-morphological traits among the accessions, with 11 key variables (fruit weight, length, width, seed length, seed width, number of seeds, volume, fresh weight, seed weight, pH, and moisture content) explaining most of the observed diversity.
Significance. If the taxonomic assignments, particularly the hybrid identifications, are adequately supported by the data, the study would provide valuable baseline information on the morpho-physiological and genetic diversity of Crataegus in a phytogeographically important but understudied semi-arid region. This could inform conservation strategies and potential utilization of these wild fruit species for medicinal, ornamental, and edible purposes.
major comments (3)
- [Materials and Methods] Materials and Methods: The description of the molecular marker systems is inadequate. The abstract mentions application of 'molecular marker systems' but provides no details on the specific markers employed (e.g., SSR, ISSR, RAPD), number of loci analyzed, PCR conditions, or the statistical/phylogenetic methods used to delimit taxa and confirm hybrid status. This is critical because the central claim of observing two specific hybrids depends on reliable evidence distinguishing them from morphological variants.
- [Results] Results: No information is given on sample sizes per taxon, replication (e.g., number of trees or fruits measured per accession), or justification for the statistical analyses. The abstract reports P < 0.01 for differences but lacks error bars, standard deviations, or details on ANOVA assumptions and post-hoc tests, undermining the robustness of the variability claims.
- [Abstract and Results] Abstract and Results: The criteria used to assign accessions to species or hybrids are not specified. Given the known morphological plasticity and reticulate evolution in Crataegus, reliance on phenotypic markers alone without explicit molecular evidence (such as additive banding patterns or intermediate clustering) for the hybrids (C. azarolus × C. meyrei and C. azarolus × C. pentagyna) leaves the taxonomic conclusions vulnerable to misclassification.
minor comments (2)
- [Abstract] There are several typographical errors, including 'varbales' (should be 'variables'), 'mositure' (moisture), 'orientalists' (likely 'orientalis'), 'Potentional of hydrogen' (Potential of hydrogen), and 'FW' used for both fruit weight and fruit width.
- [Abstract] The sentence structure is unclear in places, e.g., 'There was significant variation among different ecotypes in terms of plant type, reproductive stage, and fruit morphology and production uses.' It is not clear what 'production uses' refers to.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed comments, which have helped us identify areas where the manuscript can be strengthened. We address each major comment below and will revise the manuscript accordingly to improve clarity, transparency, and support for our findings.
read point-by-point responses
-
Referee: [Materials and Methods] The description of the molecular marker systems is inadequate. The abstract mentions application of 'molecular marker systems' but provides no details on the specific markers employed (e.g., SSR, ISSR, RAPD), number of loci analyzed, PCR conditions, or the statistical/phylogenetic methods used to delimit taxa and confirm hybrid status. This is critical because the central claim of observing two specific hybrids depends on reliable evidence distinguishing them from morphological variants.
Authors: We agree that the Materials and Methods section requires expansion to fully document the molecular component of the study. In the revised manuscript, we will add detailed descriptions of the molecular marker systems employed (specifically ISSR and RAPD), including the number of primers and loci scored, PCR reaction conditions and thermal cycling parameters, electrophoresis and visualization protocols, and the analytical approaches (UPGMA clustering, principal coordinate analysis, and assessment of additive banding patterns) used to support taxon delimitation and hybrid identification. These additions will explicitly link the molecular data to the hybrid assignments. revision: yes
-
Referee: [Results] No information is given on sample sizes per taxon, replication (e.g., number of trees or fruits measured per accession), or justification for the statistical analyses. The abstract reports P < 0.01 for differences but lacks error bars, standard deviations, or details on ANOVA assumptions and post-hoc tests, undermining the robustness of the variability claims.
Authors: We acknowledge this omission and will correct it in the revision. Each of the 61 accessions was represented by a single tree, from which 15 fruits were randomly collected for morphological and physiological measurements. We will insert this sampling detail into the Materials and Methods and add a supplementary table summarizing sample sizes per taxon. In the Results, we will include standard deviations in tables, error bars on figures, and explicit statements that one-way ANOVA was performed after verifying normality (Shapiro-Wilk) and homogeneity of variances (Levene’s test), followed by Tukey’s HSD post-hoc tests. These changes will substantiate the reported significance levels. revision: yes
-
Referee: [Abstract and Results] The criteria used to assign accessions to species or hybrids are not specified. Given the known morphological plasticity and reticulate evolution in Crataegus, reliance on phenotypic markers alone without explicit molecular evidence (such as additive banding patterns or intermediate clustering) for the hybrids (C. azarolus × C. meyrei and C. azarolus × C. pentagyna) leaves the taxonomic conclusions vulnerable to misclassification.
Authors: Taxonomic assignments integrated morphological characters (using standard Crataegus keys), phytochemical profiles, and molecular marker results. Hybrid status for the two reported taxa was supported by intermediate phenotypes plus molecular evidence of additive banding patterns and intermediate clustering positions. We recognize that these criteria were not stated explicitly. In the revised manuscript, we will add a dedicated paragraph in the Results (or a new subsection in Materials and Methods) that lists the specific morphological descriptors, phytochemical thresholds, and molecular criteria (including references to parental band profiles) used for each taxon and hybrid. This will address concerns about morphological plasticity and strengthen the hybrid identifications. revision: yes
Circularity Check
No circularity: empirical survey of observed traits and standard statistics
full rationale
The paper reports a field collection of 61 accessions, applies ANOVA to fruit physio-morphological variables, and assigns taxa (including two putative hybrids) on the basis of observed morphological, phytochemical and molecular variation. No equations, fitted parameters, predictions derived from prior fits, or self-citation chains appear in the derivation of any result. All claims reduce directly to the collected data and conventional statistical tests rather than to any self-referential construction.
Axiom & Free-Parameter Ledger
axioms (1)
- standard math Standard assumptions underlying analysis of variance (ANOVA) for detecting significant differences
Reference graph
Works this paper leans on
-
[1]
Ahmad, N. S. (2024). Assessment of genetic relation for Aegilops and Triticum possessing different genome-bearing species using start codon targeted marker. Genetic Resources and Crop Evolution, 71(4), 1465-1479. Ahmad, N. S., Abid, M. A., & Al-Assie, A. A. (2022). Profiling of foxtail millet (Setaria italica L.) germplasm using in vitro drought screening...
work page 2024
-
[2]
Alirezalu, A., Salehi, P., Ahmadi, N., Sonboli, A., Aceto, S., Hatami Maleki, H., & Ayyari, M. (2018). Flavonoids profile and antioxidant activity in flowers and leaves of Hawthorn species (Crataegus spp.) from different regions of Iran. International journal of food properties, 21(1), 452-470. Amiteye, S. (2021). Basic concepts and methodologies of DNA m...
work page 2018
-
[3]
Journal of Agricultural Science, 15(3), pp.1-58
Molecular Markers in Plant Breeding. Journal of Agricultural Science, 15(3), pp.1-58. Betancourt-Olvera, M., Nieto-Ángel, R., Urbano, B., & González-Andrés, F. (2018). Analysis of the biodiversity of Hawthorn (Crataegus spp.) from the morphological, molecular, and ethnobotanical approaches, and implications for genetic resource conservation in scenery of ...
work page 2018
-
[4]
Cole, C. T. (2003). Genetic variation in rare and common plants. Annual Review of Ecology, Evolution, and Systematics, 34(1), 213-237. Collard, B. C., and Mackill, D. J. (2009a) 'Conserved DNA-derived polymorphism (CDDP): A simple and novel method for generating DNA markers in plants', Plant Molecular Biology Reporter, 27(4): pp. 558-562. Collard, B. C., ...
work page 2003
-
[5]
Dickinson, T. A., Yan, B. X., Han, S., & Zarrei, M. (2021). Niche shifts, hybridization, polyploidy and geographic parthenogenesis in western North American Hawthorns (Crataegus subg. Sanguineae, Rosaceae). Agronomy, 11(11),
work page 2021
-
[6]
Doenmez, A. A. (2007). Taxonomic notes on the genus Crataegus (Rosaceae) in Turkey. Botanical Journal of the Linnean Society, 155(2), 231-240. Dönmez, A. A., & Özderin, S. (2019). Additional contributions to taxonomy, nomenclature and biogeography of the Turkish Crataegus (Rosaceae) taxa. PhytoKeys, 122,
work page 2007
-
[7]
V., de Almeida Vieira, F., dos Santos Ferrari, C., Félix, F
dos Santos Araújo, F., Pacheco, M. V., de Almeida Vieira, F., dos Santos Ferrari, C., Félix, F. C., & das Chagas, K. P. T. (2016). ISSR molecular markers for the study of the genetic diversity of Mimosa caesalpiniaefolia Benth. Idesia, 34(3), 47-52. Dmeria S. (2001). Biodiversity study of wild relatives of Amygdales and Crataegus in the North- West Region...
work page 2016
-
[8]
Dönmez, A. A. (2014). Nomenclatural, taxonomic and biogeographic novelties in the Turkish Crataegus L. (Rosaceae-Maleae) taxa. Adansonia, 36(2), 245-253. Donmez, A. A. (2004). The genus Crataegus L. (Rosaceae) with special reference to hybridisation and biodiversity in Turkey. Turkish Journal of Botany, 28(1), 29-37. Donno, D., M. G. Mellano, Ž. Prgomet, ...
work page 2014
-
[9]
Du, X., Zhang, X., Bu, H., Zhang, T., Lao, Y., & Dong, W. (2019). Molecular analysis of evolution and origins of cultivated Hawthorn (Crataegus spp.) and related species in China. Frontiers in Plant Science, 10,
work page 2019
-
[10]
80 de Carvalho, F. A. L., Lorenzo, J. M., Pateiro, M., Bermúdez, R., Purriños, L., & Trindade, M. A. (2019). Effect of guarana (Paullinia cupana) seed and pitanga (Eugenia uniflora L.) leaf extracts on lamb burgers with fat replacement by chia oil emulsion during shelf-life storage at 2 C. Food Research International, 125, 108554. Dhutmal, R. R., Mundhe, ...
work page 2019
-
[11]
Durazzo, A., Lucarini, M., Kiefer, J., and Mahesar, S. A. (2020). State-of-the-Art Infrared Applications in Drugs, Dietary Supplements, and Nutraceuticals. Journal of Spectroscopy,, NA-NA El-Mansy, A. B., El-Moneim, D. A., Alshamrani, S. M., Alsafhi, F. A., Abdein, M. A. and Ibrahim, A. A. (2021) 'Genetic diversity analysis of tomato (Solanum lycopersicum...
work page 2020
-
[12]
Eghlima, G., Aghamir, F., Mohammadi, M., SeyedHajizadeh, H. and Kaya, O., (2025). Bioactive Compounds and Antimicrobial Activities in Iranian Crataegus persica Ecotypes for Potential Food and Medicinal Uses. Food Science and Nutrition, 13 (1), 4748-4760 Ellegren, H., & Galtier, N. (2016). Determinants of genetic diversity. Nature Reviews Genetics, 17(7), ...
work page 2025
-
[13]
Fineschi, S., Salvini, D., Turchini, D., Pastorelli, R., & Vendramin, G. G. (2005). Crataegus monogyna Jacq. and C. laevigata (Poir.) DC.(Rosaceae, Maloideae) display low level of genetic diversity assessed by chloroplast markers. Plant Systematics and Evolution, 250, 187-196. García-Martínez, S., Andreani, L., Garcia-Gusano, M., Geuna, F. and Ruiz, J. J....
-
[14]
Halshoy, H. S., Rasul, K. S., Ahmed, H. M., Mohammed, H. A., Mohammed, A. A., Ibrahim, A. S., & Braim, S. A. (2024). Effect of nano titanium and organic fertilizer on broccoli growth, production, and biochemical profiles. Journal of plant Nutrition, 1-20. Hinsley, S. A., & Bellamy, P. E. (2000). The influence of hedge structure, management and landscape c...
work page 2024
-
[15]
Hou, W., Wang, Y., Wang, W., Zhi, M., Gou, X., Qian, C., and Zhang, F. (2020). Determination and evaluation of flavonoids in Hawthorn in China. Journal of Biobased Materials and Bioenergy, 14(5), 664-669 Huang, X., Bian, Y., Liu, T., Xu, Z., Song, Z., Wang, F., ... and Li, S. (2022). Antioxidant potential and in vitro inhibition of starch digestion of fla...
work page 2020
-
[16]
Hsiao, C., Chatterton, N., Asay, K. and Jensen, K. (1995). Molecular phylogeny of the Pooideae (Poaceae) based on nuclear rDNA (ITS) sequences. Theoretical and Applied Genetics, 90, pp. 389-398. Ibrahimov, A. M., Matsyura, A. V., and Jankowski, K. (2020). Taxonomy of the wild species of genus Crataegus (Rosaceae): An updated review for the flora of Nakhch...
-
[17]
Khdir, S. A., Ahmad, N. S., Hama-Ali, E. O., & Abdullah, S. M. (2023). Genetic diversity and population structure of common bean genotypes using morphological traits and SSR. Iraqi Journal of Agricultural Sciences, 54(3), 792-805. Khiari, S., Boussaid, M., & Messaoud, C. (2015). Genetic diversity and population structure in natural populations of Tunisian...
-
[18]
Mahmud, S. A., Al-Habib, O. A., Bugoni, S., Clericuzio, M., & Vidari, G. (2016). A new ursane- type triterpenoid and other constituents from the leaves of Crataegus azarolus var. aronia. Natural product communications, 11(11), 1934578X1601101103. Majeed, H. O., Faraj, J. M., Rasul, K. S., Lateef, D. D., & Tahir, N. A. R. (2024). Evaluation of the genetic ...
work page 2016
-
[19]
Moustafa, A., Zaghloul, M., Mansour, S., & Alotaibi, M. (2019). Conservation Strategy for protecting Crataegus x sinaica against climate change and anthropologic activities in South Sinai Mountains, Egypt. Catrina: The International Journal of Environmental Sciences, 18(1), 1-6.. Mata-Nicolás, E., Montero-Pau, J., Gimeno-Paez, E., Garcia-Carpintero, V., Z...
work page 2019
-
[20]
Miao, J., Li, X., Fan, Y.Y., Zhao, C.C., Mao, X.H., Chen, X.T., Huang, H.H., and Gao, W.Y. (2016). Effect of different solvents on the chemical composition, antioxidant activity and α-glucosidase inhibitory activity of Hawthorn extracts. International Journal of Food Science & Technology 51(5): 1244–1251. Mullis, K., Faloona, F., Scharf, S., Saiki, R., Ho...
work page 2016
-
[21]
Nadeem, M. A., Nawaz, M. A., Shahid, M. Q., Doğan, Y., Comertpay, G., Yıldız, M., ... & Baloch, F. S. (2018). DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnology & Biotechnological Equipment, 32(2), 261-285. Nadeem, M. A. (2021). Deciphering the genetic diversity and popula...
work page 2018
-
[22]
Nguyen, Q. T. N., Fang, M., Zhang, M., Do, N. Q., Kim, M., Zheng, S.D., Hwang, E., and Yi, T. H. (2021). Crataegus laevigata suppresses LPS-induced oxidative stress during inflammatory response in human ker-atinocytes by regulating the MAPKs/AP-1, NFκB, and NFAT signalingpathways. Molecules (Basel, Switzerland), 26(4),
work page 2021
-
[23]
Ng, W. L., & Tan, S. G. (2015). Inter-simple sequence repeat (ISSR) markers: are we doing it right. ASM Sci J, 9(1), 30-39. Obideen, R., Alhasan, A. and Draie, R.,
work page 2015
-
[24]
African Journal of Biological Sciences
Morphological Characterization of Hawthorn Types (Crataegus spp.) Widespread in Northwestern Syria. African Journal of Biological Sciences.. 6(8) 1058-1073 87 Orhan, I. E. (2018). Phytochemical and pharmacological activity profile of Crataegus oxyacantha L.(Hawthorn)-a cardiotonic herb. Current medicinal chemistry, 25(37), 4854-4865. Orhan, I. E. (2018). ...
work page 2018
-
[25]
K., Stephens, M., & Donnelly, P
Pritchard, J. K., Stephens, M., & Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics, 155(2), 945-959. Priyanka, V., Kumar, R., Dhaliwal, I., & Kaushik, P. (2021). Germplasm conservation: instrumental in agricultural biodiversity—a review. Sustainability, 13(12),
work page 2000
-
[26]
Kurdistan Journal of Applied Research, 10:116-125, https://doi.org/10.24017/science.2025.1.8
Qalatobzany HSA, Muhammad KA, Lateef DD, Rasul KS, Ibrahim AS, Parreira MC, Weisany W (2025) Analysis of the Metabolic Profile and Biological Activity of Hawthorn Species twigs: Crataegus azarolus and Crataegus monogyna. Kurdistan Journal of Applied Research, 10:116-125, https://doi.org/10.24017/science.2025.1.8. Radha, K. O., & Khwarahm, N. R. (2022). An...
-
[27]
Z., Bencheikh, N., Anarghou, H., Bouhrim, M., Alqahtani, A
Radi, F. Z., Bencheikh, N., Anarghou, H., Bouhrim, M., Alqahtani, A. S., Hawwal, M. F., ... & Zair, T. (2023). Quality control, phytochemical profile, and biological activities of Crataegus monogyna Jacq. and Crataegus laciniata Ucria fruits aqueous extracts. Saudi Pharmaceutical Journal, 31(10), 101753. Rahmani, M. S., Shabanian, N., Khadivi-Khub, A., Wo...
work page 2023
-
[28]
Rasul, K. S., Grundler, F. M., & Abdul-razzak Tahir, N. (2022). Genetic diversity and population structure assessment of Iraqi tomato accessions using fruit characteristics and molecular markers. Horticulture, Environment, and Biotechnology, 63(4), 523-538. Rasul, K. S., Majeed, H. O., Faraj, J. M., Lateef, D. D., & Tahir, N. A. R. (2024). Genetic diversi...
work page 2022
-
[29]
Rasul, K. S., Grundler, F. M., & Abdul-razzak Tahir, N. (2022). Genetic diversity and population structure assessment of Iraqi tomato accessions using fruit characteristics and molecular markers. Horticulture, Environment, and Biotechnology, 63(4), 523-538. Rasul, K. S., Majeed, H. O., Faraj, J. M., Lateef, D. D., & Tahir, N. A. R. (2024). Genetic diversi...
work page 2022
-
[30]
Sevindik, E., Efe, F., & Murathan, Z. T. (2023). Molecular genetic diversity and phylogenetic investigation of Pyrus communis L.(Rosaceae) genotypes using cpDNA sequences with RAPD and ISSR analyses. Erwerbs-Obstbau, 65(2), 231-240. Sharef, A.A., Faraj, J.M., Rasul, K.S. et al. Morphological and Molecular Marker Analysis of F1 Hybrid Petunia Obtained from...
-
[31]
Tahir, N. A. R., Rasul, K. S., & Lateef, D. D. (2023). Effect of mixing oak leaf biomass with soil on cadmium toxicity and translocation in tomato genotypes. Heliyon, 9(8). Tahir, N., Lateef, D. D., Rasul, K. S., Rahim, D., Mustafa, K. M., Sleman, S., ... & Aziz, R. (2023). Assessment of genetic variation and population structure in Iraqi barley accession...
work page 2023
-
[32]
Tahir, N. A.-r., Lateef, D. D., Mustafa, K. M., Rasul, K. S., & Khurshid, F. F. (2023). Determination of Physiochemical Characteristics Associated with Various Degrees of Cadmium Tolerance in Barley Accessions. Agronomy, 13(6),
work page 2023
-
[33]
https://doi.org/10.3390/agronomy13061502 Townsend, C., & Guest, G. (1985). Flora of Iraq, vol. 8:
-
[34]
Ministry of Agriculture & Agrarian Reform, Baghdad, 137-177. Vaughn, B. (2015). Hawthorn: The tree that has nourished, healed, and inspired through the ages. Yale University Press. Victoria, F. C., da Maia, L. C., & de Oliveira, A. C. (2011). In silico comparative analysis of SSR markers in plants. BMC plant biology, 11, 1-15. Wróblewska, A., Brzosko, E.,...
work page 2015
-
[35]
Sagheb Talebi, K., Sajedi, T., and Pourhashemi, M. (2014). Forests of Iran: A Treasure from the Past, a Hope for the Future (No. 15325). Springer Netherlands. Sakar, E., Ercisli, S., Durul, M. S., Singh, M., Anjum, M. A., Orhan, E., & Kan, T. (2024). Sensory, morphological, biochemical, and antioxidant characteristics of the fruits of different Cactus 91 ...
work page 2014
-
[36]
Shahat AA, Cos P, De Bruyne T, Apers S, Hammouda FM, Ismail SI, Azzam S, Clayes M, Goovaerts E, Pieters L, Van den Berghe D, Vlietinck AJ. (2002). Antiviral and antioxidant activity of flavonoids and proanthocyanidins from Crataegus sinaica. Planta Medica 68: 539–541 Shahidi, F. (2004). Functional foods: Their role in health promotion and disease preventi...
work page 2002
-
[37]
Phylogenetic study some of Crataegus L
Sharfni F, Seyedipour N, Mehregan I and Salimpour F (2013). Phylogenetic study some of Crataegus L. (Rosaceae, Pyreae) Species in Iran. Journal of Biodiversity and Environmental Sciences. 3(11) 1-11. Sharifnia F (2011). Crataegus grossidentata sp. nov. (Rosaceae, Pyreae), a new Hawthorn from northern Iran. Nordic Journal of Botany 29 534 –
work page 2013
-
[38]
Sultana, B., Anwar, F., and Ashraf, M. (2009). Effect of extraction solvent/technique on the antioxidant activity of selected medicinal plant extracts. Molecules, 14(6), 2167-2180. Sun, B., Huo, H. Z., Cai, A. H., Xie, Y. C., Li, H. Y., & Li, D. P. (2021). Determination of contents of eight phenolic acids in Malus doumeri fruit by HPLC. 07:1135–1144 Sun, ...
work page 2009
-
[39]
U., Khadivi, A., & Goodarzi, S
Tunç, Y., Yaman, M., Yılmaz, K. U., Khadivi, A., & Goodarzi, S. (2025). Elucidation of genetic diversity in Hawthorn (Crataegus azarolus L.) accessions naturally growing in the Eastern Mediterranean region of Türkiye using morphological, pomological, and inter simple sequence repeat markers. BMC Plant Biology, 25(1),
work page 2025
-
[40]
Uzun, A., Pinar, H., Yaman, M., Çakiroğlu, Y., Karakaya, A., Uysal, M., ... & Akol, S. (2025). Assessment of genetic diversity by multivariate analysis in Turkish Hawthorn (Crataegus azarolus) genetic resources. Genetic Resources and Crop Evolution, 1-13. Venskutonis, P. R. (2018). Phytochemical composition and bioactivities of Hawthorn (Crataegus spp.): ...
work page 2025
-
[41]
Venskutonis, P. R. (2018). Phytochemical composition and bioactivities of Hawthorn (Crataegus spp.): Review of recent research advances. Journal of food bioactives., 4, 69-87. Wang, B., Wu, X., Luo, D., Chen, J., Zhang, Y., Li, G., & Zhang, T. (2024). Genome-wide survey of Crataegus scabrifolia provides new insights into its genetic evolution and adaptati...
work page 2018
-
[42]
Wu, J., Peng, W., Qin, R., & Zhou, H
Available online: http://www.who.int/traditional- complementaryintegrative-medicine/publications/trm_strategy14_23/en/ (accessed on 5 May 2020). Wu, J., Peng, W., Qin, R., & Zhou, H. (2014). Crataegus pinnatifida: chemical constituents, pharmacology, and potential applications. Molecules, 19(2), 1685-1712. Wu, L., Cui, Y., Wang, Q., Xu, Z., Wang, Y., Lin,...
work page 2020
-
[43]
Zhang, J., Deng, Y., Tong, X., Liang, Y., Li, Y., & Deng, C. (2021). Nutrition and Health Function of Crataegus pinnatifida Bunge and Its Application Progres. Anhui Agric. Sci. Bull, 27, 116-118. Zhang, T., Qiao, Q., Du, X., Zhang, X., Hou, Y., Wei, X., ... & Dong, W. (2022). Cultivated Hawthorn (Crataegus pinnatifida var. major) genome sheds light on the...
work page 2021
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.