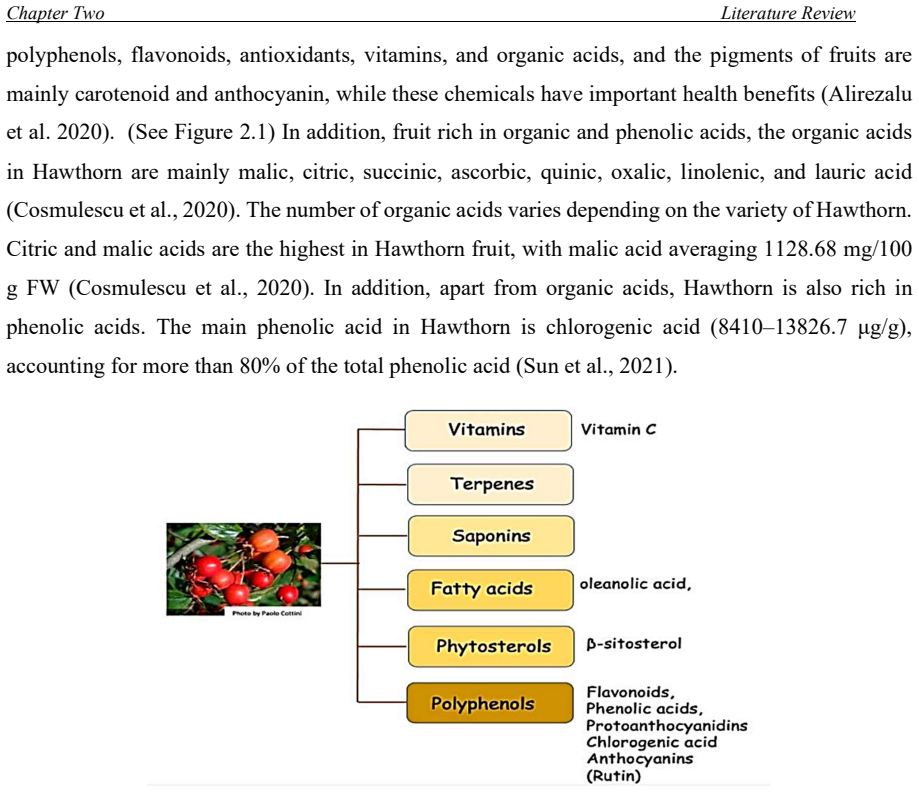
Kurdistan study finds seven hawthorn taxa with major fruit variation
Sixty-one accessions vary significantly in weight, size, seeds, pH and moisture, explained by eleven traits.
 full image
full image
Other Quantitative Biology
Work in quantitative biology that does not fit into the other q-bio classifications
Sixty-one accessions vary significantly in weight, size, seeds, pH and moisture, explained by eleven traits.
 full image
full image
Statin Recommendations among US Adults with the 2026 Dyslipidemia Guidelines
The outcome hinges on whether the new 30-year risk pathway is applied, expanding access most for adults aged 40-59.
 full image
full image
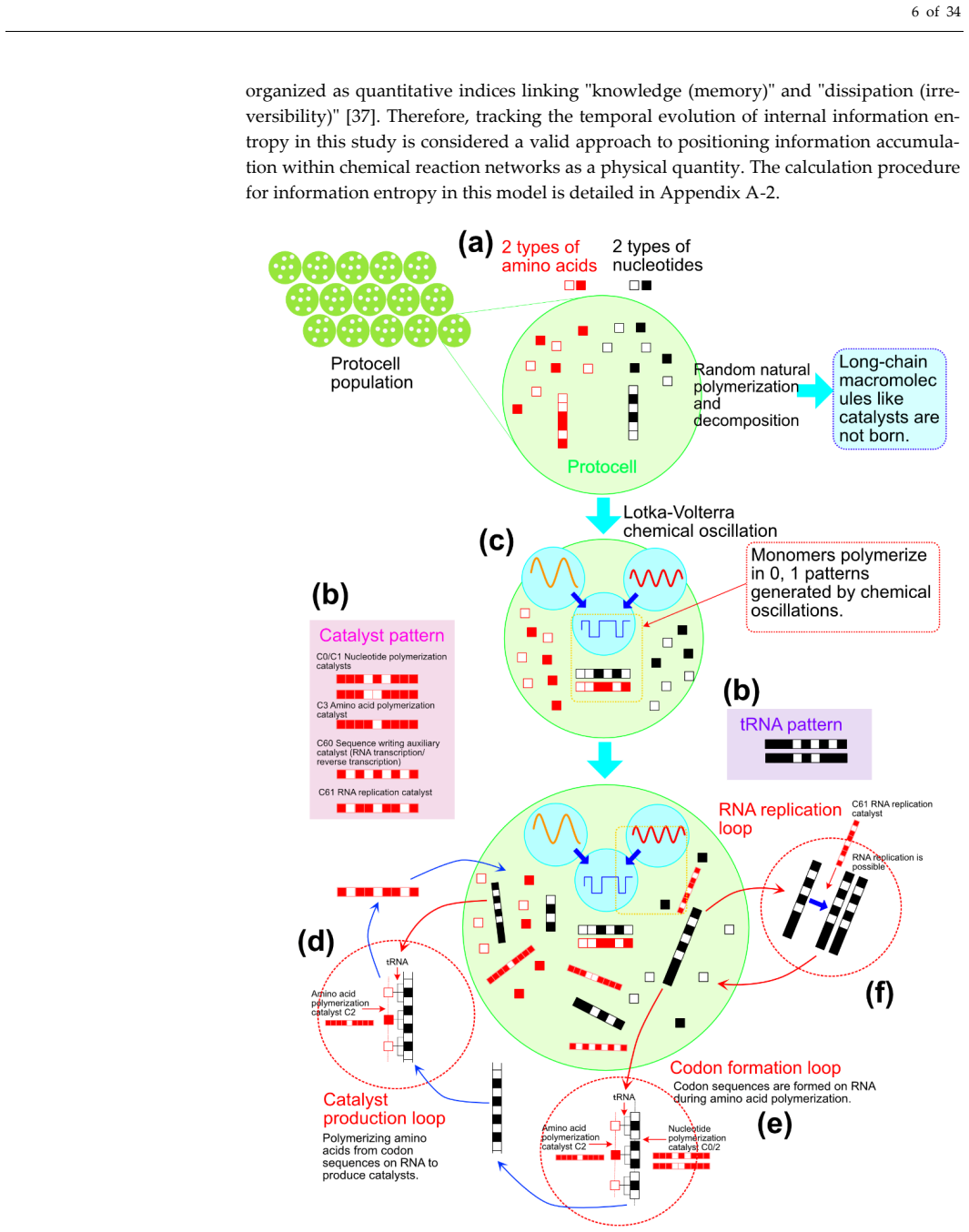
Simulations link superposed chemical oscillators to higher rates of functional polymer emergence compared with random selection.
 full image
full image
Framework builds harmonized representations from genetic, environmental and brain data to support longitudinal and event-based prediction of
 full image
full image
BioVeil MATRIX: Uncovering and categorizing vulnerabilities of agentic biological AI scientists
Scaffolding on Biomni raises scores on WMDP biology proxies; new taxonomy with 10 categories maps the agentic risks.
 full image
full image
Tumor containment as an anti-percolation process
Tumor containment framed as anti-percolation, with simulations indicating partial independence of malignant area from connectivity metrics…
Personalizing Cancer Models under Data Scarcity via Parameter Decomposition
Decomposing parameters lets the population-level part serve as a prior so only the patient-specific part needs fitting to limited new data.
 full image
full image
Entropy-Dominated Temporal Vocal Dynamics as Digital Biomarkers for Depression Detection
Unpredictability in speech timing and pitch raises AUC from 0.593 to 0.646 on the DAIC-WOZ dataset under strict validation.
 full image
full image
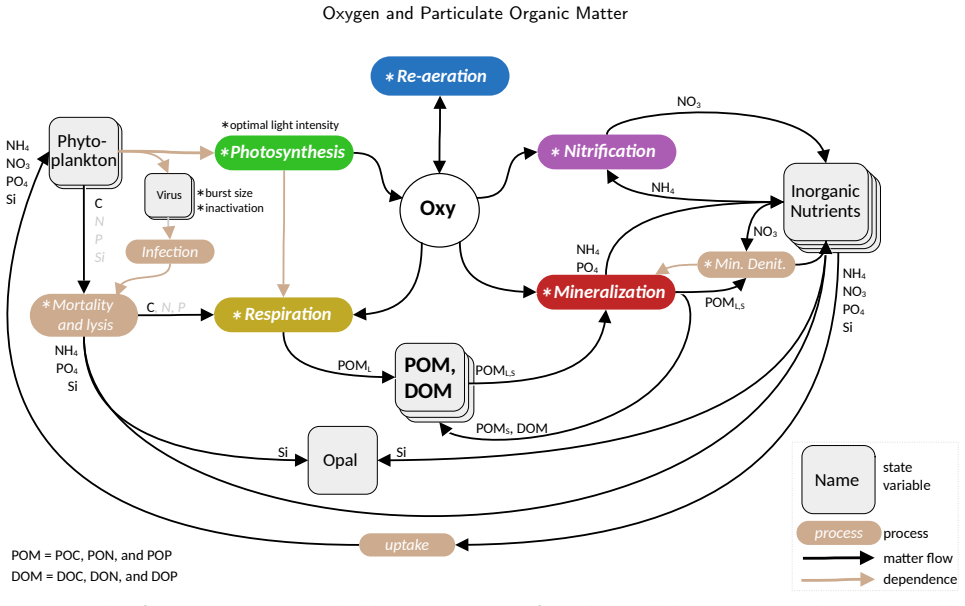
Uniform sensitivities in oxygen models underestimate particulate organic carbon and overestimate nutrients across seasons in low-oxygenwater
 full image
full image
Differential Analysis of Microbial Interaction Networks
Gene-family association analysis detects interaction shifts between men and women that abundance changes alone do not capture.
 full image
full image
Multi-stage model stabilizes features with network analysis, uses CNN on disease maps, and beats standard ML on classification.
 full image
full image
National survey analysis identifies a high-risk group where 77 percent of women with adverse reproductive burdens already meet multimorbidty
 full image
full image
StackFeat: a convergent algorithm for optimal predictor selection in genomic data
StackFeat combines signed coefficients and selection frequencies across repeated cross-validations to converge on reliable predictors that,
 full image
full image
Energy gradients as potential drivers of pre-cellular chemical organization
Simulations show strong coupled gradients in pH, redox and temperature overcome diffusion to create stable confined chemical states.
Mathematical modeling and intuition in microbiology: a perspective
They require consistent hypotheses, support testable forecasts, extract unmeasured parameters, and guide choices of model detail for any set
 full image
full image
Lipophilicity, molecular weight and polarity emerge as the key features driving accuracy in database-screened predictions
 full image
full image
Baseline glycemia exhibits non-random, history-dependent variation across repeated meals
Repeated identical meals produce non-random baseline changes scaled to the last post-meal rise, indicating memory in glucose control.