Recognition: no theorem link
MolDeTox: Evaluating Language Model's Stepwise Fragment Editing for Molecular Detoxification
Pith reviewed 2026-05-13 05:19 UTC · model grok-4.3
The pith
Fragment-level molecule editing in language models raises structural validity and quality for detoxification tasks.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
MolDeTox evaluates general-purpose LLMs and VLMs on stepwise fragment editing for molecular detoxification. The central claim is that understanding and generating molecules at the fragment level improves structural validity and enhances the quality of generated molecules while supplying an interpretable benchmark for the detoxification process.
What carries the argument
The MolDeTox benchmark, built around stepwise fragment-editing tasks that isolate toxicity removal while preserving other molecular properties.
If this is right
- Fragment-level processing produces a higher fraction of chemically valid detoxified molecules than whole-structure approaches.
- Task-level analysis isolates which steps in detoxification models handle reliably and where they fail.
- The same general-purpose models show measurable gains when the input format shifts to fragments.
- The benchmark enables direct comparison of LLMs and VLMs under consistent toxicity-aware conditions.
Where Pith is reading between the lines
- The fragment approach could transfer to other molecular optimization goals such as potency or solubility changes.
- Wider use of stepwise fragment benchmarks might lower dependence on full experimental validation during early screening.
- Future versions would benefit from pairing the proxy toxicity scores with at least a subset of real assay results to test alignment.
Load-bearing premise
Proxy models for toxicity assessment are accurate enough and the benchmark's added data diversity and validity metrics fully overcome the shortcomings of earlier toxicity repair tests.
What would settle it
Running the same models on MolDeTox tasks with whole-molecule editing instead of fragment steps and finding no gain in structural validity or quality scores.
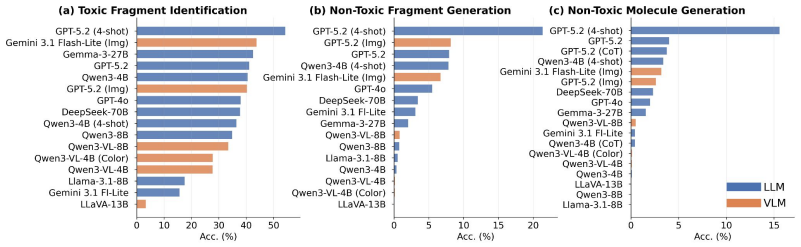
Figures













read the original abstract
Large Language Models (LLMs) and Vision Language Models (VLMs) have recently shown promising capabilities in various scientific domain. In particular, these advances have opened new opportunities in drug discovery, where the ability to understand and modify molecular structures is critical for optimizing drug properties such as efficacy and toxicity. However, existing models and benchmarks often overlook toxicity-related challenges, focusing primarily on general property optimization without adequately addressing safety concerns. In addition, even existing toxicity repair benchmarks suffer from limited data diversity, low structural validity of generated molecules, and heavy reliance on proxy models for toxicity assessment. To address these limitations, we propose MolDeTox, a novel benchmark for molecular detoxification, designed to enable fine-grained and reliable evaluation of toxicity-aware molecular optimization across stepwise tasks. We evaluate a wide range of general-purpose LLMs and VLMs under diverse settings, and demonstrate that understanding and generating molecules at the fragment-level improves structural validity and enhances the quality of generated molecules. Moreover, through detailed task-level performance analysis, MolDeTox provides an interpretable benchmark that enables a deeper understanding of the detoxification process. Our dataset is available at : https://huggingface.co/datasets/MolDeTox/MolDeTox
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proposes MolDeTox, a new benchmark for evaluating LLMs and VLMs on stepwise fragment-level editing tasks for molecular detoxification. It critiques prior toxicity repair benchmarks for limited data diversity, low structural validity, and proxy-model reliance, then claims that fragment-level molecular understanding and generation improves structural validity and overall quality of detoxified molecules while enabling interpretable task-level analysis. A range of general-purpose models is evaluated and the dataset is released.
Significance. If the fragment-level mechanism demonstrably improves detoxification outcomes, the benchmark could provide a useful, more interpretable resource for AI-assisted drug design. The focus on addressing data diversity and validity gaps in existing toxicity benchmarks is constructive, and releasing the dataset supports reproducibility.
major comments (2)
- [§4 and §5] §4 (Evaluation Methodology) and §5 (Results): The central claim that fragment-level editing enhances detoxification quality depends on the reliability of the proxy toxicity models used to label success. No correlation studies, calibration against experimental toxicity data, or wet-lab confirmation are reported, despite the abstract explicitly critiquing prior benchmarks for the same proxy reliance. If proxies misclassify toxicity (e.g., due to scaffold biases), gains in structural validity and proxy scores do not establish actual detoxification improvement.
- [§3] §3 (Benchmark Construction): Dataset statistics, diversity metrics, and validity rates for the MolDeTox split are not compared quantitatively to prior toxicity repair benchmarks, leaving the claim that the new benchmark overcomes their limitations unsupported by direct evidence.
minor comments (2)
- [Abstract] Abstract: Performance improvements are asserted without any numerical results, error bars, or dataset size summaries; a single sentence summarizing key metrics would improve clarity.
- [Figures/Tables] Figure and table captions: Ensure all figures reporting model comparisons include the exact proxy model names and thresholds used for toxicity labeling.
Simulated Author's Rebuttal
We thank the referee for the thoughtful and constructive comments on our manuscript. We address each major point below with honest responses and indicate where revisions will be made to strengthen the work.
read point-by-point responses
-
Referee: [§4 and §5] §4 (Evaluation Methodology) and §5 (Results): The central claim that fragment-level editing enhances detoxification quality depends on the reliability of the proxy toxicity models used to label success. No correlation studies, calibration against experimental toxicity data, or wet-lab confirmation are reported, despite the abstract explicitly critiquing prior benchmarks for the same proxy reliance. If proxies misclassify toxicity (e.g., due to scaffold biases), gains in structural validity and proxy scores do not establish actual detoxification improvement.
Authors: We agree that proxy-based toxicity assessment is a limitation of the current work, as it is for most computational molecular benchmarks. Our primary contributions focus on demonstrating improvements in structural validity (measured via independent cheminformatics checks such as RDKit sanitization) and on providing interpretable fragment-level task analysis, which do not rely on the toxicity proxies. The stepwise editing mechanism itself is shown to increase the fraction of valid outputs compared to direct generation baselines. We acknowledge the inconsistency with our critique of prior work and will revise §4 and §5 to include an expanded limitations subsection that explicitly discusses proxy reliability, cites known calibration studies from the literature where available, and clarifies that our validity gains are proxy-independent. We cannot add new wet-lab experiments, as this is a benchmark and evaluation paper rather than an experimental study. revision: partial
-
Referee: [§3] §3 (Benchmark Construction): Dataset statistics, diversity metrics, and validity rates for the MolDeTox split are not compared quantitatively to prior toxicity repair benchmarks, leaving the claim that the new benchmark overcomes their limitations unsupported by direct evidence.
Authors: We thank the referee for highlighting this gap. In the revised manuscript we will add a dedicated comparison subsection (or table) in §3 that quantitatively reports dataset statistics including number of molecules, average Tanimoto diversity, scaffold diversity, and pre- and post-filtering validity rates, directly juxtaposed against the prior toxicity repair benchmarks referenced in the introduction. This will provide the direct evidence requested and better substantiate our claims regarding improved diversity and validity. revision: yes
- Experimental wet-lab confirmation or new correlation studies against measured toxicity data, which lie outside the scope of this computational benchmark paper.
Circularity Check
No significant circularity in empirical benchmark evaluation
full rationale
The paper proposes the MolDeTox benchmark for stepwise fragment-level molecular detoxification using LLMs and VLMs, then reports empirical results showing improved structural validity and quality. No equations, derivations, fitted parameters, or predictions appear that reduce to inputs by construction. Claims rest on direct evaluation against the new dataset and proxy metrics rather than any self-referential logic or self-citation chain. Proxy toxicity model reliance is a methodological assumption about data quality, not a circular step. The work is self-contained as an empirical study with no load-bearing self-citations or ansatzes.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption LLMs and VLMs can meaningfully process and edit molecular structures represented as text (e.g., SMILES) or images for property optimization tasks
Reference graph
Works this paper leans on
-
[1]
Training a scientific reasoning model for chemistry.arXiv preprint arXiv:2506.17238, 2025
Siddharth M Narayanan, James D Braza, Ryan-Rhys Griffiths, Albert Bou, Geemi Wellawatte, Mayk Caldas Ramos, Ludovico Mitchener, Samuel G Rodriques, and Andrew D White. Training a scientific reasoning model for chemistry.arXiv preprint arXiv:2506.17238, 2025
-
[2]
ChemDFM-R: A Chemical Reasoning LLM Enhanced with Atomized Chemical Knowledge
Zihan Zhao, Bo Chen, Ziping Wan, Lu Chen, Xuanze Lin, Shiyang Yu, Situo Zhang, Da Ma, Zichen Zhu, Danyang Zhang, et al. Chemdfm-r: A chemical reasoning llm enhanced with atomized chemical knowledge.arXiv preprint arXiv:2507.21990, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[3]
Adibvafa Fallahpour, Andrew Magnuson, Purav Gupta, Shihao Ma, Jack Naimer, Arnav Shah, Haonan Duan, Omar Ibrahim, Hani Goodarzi, Chris J Maddison, et al. Bioreason: Incentivizing multimodal biological reasoning within a dna-llm model.arXiv preprint arXiv:2505.23579, 2025
-
[4]
Adibvafa Fallahpour, Arman Seyed-Ahmadi, Parsa Idehpour, Omar Ibrahim, Purav Gupta, Jack Naimer, Kevin Zhu, Arnav Shah, Shihao Ma, Abhinav Adduri, et al. Bioreason-pro: Advancing protein function prediction with multimodal biological reasoning.bioRxiv, pages 2026–03, 2026
work page 2026
-
[5]
Ana-Maria Istrate, Fausto Milletari, Fabrizio Castrotorres, Jakub M Tomczak, Michaela Torkar, Donghui Li, and Theofanis Karaletsos. rbio1-training scientific reasoning llms with biological world models as soft verifiers.bioRxiv, pages 2025–08, 2025
work page 2025
-
[6]
Chemvlm: Exploring the power of multimodal large language models in chemistry area
Junxian Li, Di Zhang, Xunzhi Wang, Zeying Hao, Jingdi Lei, Qian Tan, Cai Zhou, Wei Liu, Yaotian Yang, Xinrui Xiong, et al. Chemvlm: Exploring the power of multimodal large language models in chemistry area. InProceedings of the AAAI Conference on Artificial Intelligence, volume 39, pages 415–423, 2025
work page 2025
-
[7]
Deepan Adak, Yogesh Singh Rawat, and Shruti Vyas. Molvision: Molecular property prediction with vision language models.arXiv preprint arXiv:2507.03283, 2025
-
[8]
Jiajun Yu, Yizhen Zheng, Huan Yee Koh, Shirui Pan, Tianyue Wang, and Haishuai Wang. Collaborative expert llms guided multi-objective molecular optimization.arXiv preprint arXiv:2503.03503, 2025
-
[9]
Tung Nguyen and Aditya Grover. Lico: Large language models for in-context molecular optimization.arXiv preprint arXiv:2406.18851, 2024
-
[10]
Geyan Ye, Xibao Cai, Houtim Lai, Xing Wang, Junhong Huang, Longyue Wang, Wei Liu, and Xiangxiang Zeng. Drugassist: A large language model for molecule optimization.Briefings in Bioinformatics, 26(1):bbae693, 2025
work page 2025
-
[11]
Feiyang Cai, Jiahui Bai, Tao Tang, Guijuan He, Joshua Luo, Tianyu Zhu, Srikanth Pilla, Gang Li, Ling Liu, and Feng Luo. Mollangbench: A comprehensive benchmark for language-prompted molecular structure recognition, editing, and generation.arXiv preprint arXiv:2505.15054, 2025
-
[12]
Hao Li, He Cao, Bin Feng, Yanjun Shao, Xiangru Tang, Zhiyuan Yan, Li Yuan, Yonghong Tian, and Yu Li. Beyond chemical qa: Evaluating llm’s chemical reasoning with modular chemical operations.arXiv preprint arXiv:2505.21318, 2025. 10
-
[13]
Fei Lin, Ziyang Gong, Cong Wang, Tengchao Zhang, Yonglin Tian, Yining Jiang, Ji Dai, Chao Guo, Xiaotong Yu, Xue Yang, et al. Breaking bad molecules: are mllms ready for structure-level molecular detoxification?arXiv preprint arXiv:2506.10912, 2025
-
[14]
Gotta be safe: a new framework for molecular design.Digital Discovery, 3(4):796–804, 2024
Emmanuel Noutahi, Cristian Gabellini, Michael Craig, Jonathan SC Lim, and Prudencio Tossou. Gotta be safe: a new framework for molecular design.Digital Discovery, 3(4):796–804, 2024
work page 2024
-
[15]
Amit S Kalgutkar. Designing around structural alerts in drug discovery.Journal of medicinal chemistry, 63(12):6276–6302, 2019
work page 2019
-
[16]
Hyomin Kim, Yunhui Jang, and Sungsoo Ahn. Mt-mol: Multi agent system with tool-based reasoning for molecular optimization.Artificial Intelligence Repository, 2025
work page 2025
-
[17]
Hengzheng Yang, Jian Xiu, Weiqi Yan, Kaifeng Liu, Huizi Cui, Zhibang Wang, Qizheng He, Yilin Gao, and Weiwei Han. Large language models as tools for molecular toxicity prediction: Ai insights into cardiotoxicity.Journal of Chemical Information and Modeling, 65(5):2268–2282, 2025
work page 2025
-
[18]
Yi-Qi Chen, Tao Yu, Zheng-Qi Song, Chen-Yu Wang, Jiang-Tao Luo, Yong Xiao, Heng Qiu, Qing-Qing Wang, and Hai-Ming Jin. Application of large language models in drug-induced osteotoxicity prediction.Journal of Chemical Information and Modeling, 65(7):3370–3379, 2025
work page 2025
-
[19]
Cotox: Chain-of-thought-based molecular toxicity reasoning and prediction
Jueon Park, Yein Park, Minju Song, Soyon Park, Donghyeon Lee, Seungheun Baek, and Jaewoo Kang. Cotox: Chain-of-thought-based molecular toxicity reasoning and prediction. In2025 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), pages 4002–4007. IEEE, 2025
work page 2025
-
[20]
ToxReason: A Benchmark for Mechanistic Chemical Toxicity Reasoning via Adverse Outcome Pathway
Jueon Park, Wonjune Jang, Chanhwi Kim, Yein Park, and Jaewoo Kang. Toxreason: A benchmark for mechanistic chemical toxicity reasoning via adverse outcome pathway.arXiv preprint arXiv:2604.06264, 2026
work page internal anchor Pith review Pith/arXiv arXiv 2026
-
[21]
Quantifying the chemical beauty of drugs.Nature chemistry, 4(2):90–98, 2012
G Richard Bickerton, Gaia V Paolini, Jérémy Besnard, Sorel Muresan, and Andrew L Hopkins. Quantifying the chemical beauty of drugs.Nature chemistry, 4(2):90–98, 2012
work page 2012
-
[22]
Txgemma: Efficient and agentic llms for therapeutics
Eric Wang, Samuel Schmidgall, Paul F Jaeger, Fan Zhang, Rory Pilgrim, Yossi Matias, Joelle Barral, David Fleet, and Shekoofeh Azizi. Txgemma: Efficient and agentic llms for therapeutics. arXiv preprint arXiv:2504.06196, 2025
-
[23]
Shraddha Thakkar, Ting Li, Zhichao Liu, Leihong Wu, Ruth Roberts, and Weida Tong. Drug- induced liver injury severity and toxicity (dilist): binary classification of 1279 drugs by human hepatotoxicity.Drug discovery today, 25(1):201–208, 2020
work page 2020
-
[24]
Yanyan Qu, Ting Li, Zhichao Liu, Dongying Li, and Weida Tong. Dictrank: The largest reference list of 1318 human drugs ranked by risk of drug-induced cardiotoxicity using fda labeling.Drug Discovery Today, 28(11):103770, 2023
work page 2023
-
[25]
Skylar Connor, Ting Li, Yanyan Qu, Ruth A Roberts, and Weida Tong. Generation of a drug-induced renal injury list to facilitate the development of new approach methodologies for nephrotoxicity.Drug discovery today, 29(4):103938, 2024
work page 2024
-
[26]
Admet eval- uation in drug discovery
Shuangquan Wang, Huiyong Sun, Hui Liu, Dan Li, Youyong Li, and Tingjun Hou. Admet eval- uation in drug discovery. 16. predicting herg blockers by combining multiple pharmacophores and machine learning approaches.Molecular pharmaceutics, 13(8):2855–2866, 2016
work page 2016
-
[27]
Fang Du, Haibo Yu, Beiyan Zou, Joseph Babcock, Shunyou Long, and Min Li. hergcentral: a large database to store, retrieve, and analyze compound-human ether-a-go-go related gene channel interactions to facilitate cardiotoxicity assessment in drug development.Assay and drug development technologies, 9(6):580–588, 2011
work page 2011
-
[28]
Abdul Karim, Matthew Lee, Thomas Balle, and Abdul Sattar. Cardiotox net: a robust pre- dictor for herg channel blockade based on deep learning meta-feature ensembles.Journal of cheminformatics, 13(1):60, 2021. 11
work page 2021
-
[29]
Congying Xu, Feixiong Cheng, Lei Chen, Zheng Du, Weihua Li, Guixia Liu, Philip W Lee, and Yun Tang. In silico prediction of chemical ames mutagenicity.Journal of chemical information and modeling, 52(11):2840–2847, 2012
work page 2012
-
[30]
Predicting chemically-induced skin reactions
Vinicius M Alves, Eugene Muratov, Denis Fourches, Judy Strickland, Nicole Kleinstreuer, Carolina H Andrade, and Alexander Tropsha. Predicting chemically-induced skin reactions. part i: Qsar models of skin sensitization and their application to identify potentially hazardous compounds.Toxicology and applied pharmacology, 284(2):262–272, 2015
work page 2015
-
[31]
Ruili Huang, Menghang Xia, Dac-Trung Nguyen, Tongan Zhao, Srilatha Sakamuru, Jinghua Zhao, Sampada A Shahane, Anna Rossoshek, and Anton Simeonov. Tox21challenge to build predictive models of nuclear receptor and stress response pathways as mediated by exposure to environmental chemicals and drugs.Frontiers in Environmental Science, 3:85, 2016
work page 2016
-
[32]
Kaitlyn M Gayvert, Neel S Madhukar, and Olivier Elemento. A data-driven approach to predicting successes and failures of clinical trials.Cell chemical biology, 23(10):1294–1301, 2016
work page 2016
-
[33]
Henrike Veith, Noel Southall, Ruili Huang, Tim James, Darren Fayne, Natalia Artemenko, Min Shen, James Inglese, Christopher P Austin, David G Lloyd, et al. Comprehensive characteriza- tion of cytochrome p450 isozyme selectivity across chemical libraries.Nature biotechnology, 27(11):1050–1055, 2009
work page 2009
-
[34]
Martin Kondža, Josipa Buki´c, Ivan ´Cavar, and Biljana Tubi´c. Targeted but troubling: Cyp450 inhibition by kinase and parp inhibitors and its clinical implications.Drugs and drug candidates, 4(2):24, 2025
work page 2025
-
[35]
The sider database of drugs and side effects.Nucleic acids research, 44(D1):D1075–D1079, 2016
Michael Kuhn, Ivica Letunic, Lars Juhl Jensen, and Peer Bork. The sider database of drugs and side effects.Nucleic acids research, 44(D1):D1075–D1079, 2016
work page 2016
-
[36]
Derek Van Tilborg, Alisa Alenicheva, and Francesca Grisoni. Exposing the limitations of molecular machine learning with activity cliffs.Journal of chemical information and modeling, 62(23):5938–5951, 2022
work page 2022
-
[37]
Hajung Kim, Jueon Park, Junseok Choe, Sheunheun Baek, Hyeon Hwang, and Jaewoo Kang. Graphcliff: Short-long range gating for subtle differences but critical changes.arXiv preprint arXiv:2511.03170, 2025
-
[38]
Ziyi Yang, Shaohua Shi, Li Fu, Aiping Lu, Tingjun Hou, and Dongsheng Cao. Matched molecular pair analysis in drug discovery: methods and recent applications.Journal of Medicinal Chemistry, 66(7):4361–4377, 2023
work page 2023
-
[39]
Ch Sanjeev Kumar Dash, Ajit Kumar Behera, Satchidananda Dehuri, and Ashish Ghosh. An outliers detection and elimination framework in classification task of data mining.Decision Analytics Journal, 6:100164, 2023
work page 2023
-
[40]
Christopher A Lipinski, Franco Lombardo, Beryl W Dominy, and Paul J Feeney. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings.Advanced drug delivery reviews, 23(1-3):3–25, 1997
work page 1997
-
[41]
Molecular properties that influence the oral bioavailability of drug candidates
Daniel F Veber, Stephen R Johnson, Hung-Yuan Cheng, Brian R Smith, Keith W Ward, and Ken- neth D Kopple. Molecular properties that influence the oral bioavailability of drug candidates. Journal of medicinal chemistry, 45(12):2615–2623, 2002
work page 2002
-
[42]
Aaron Hurst, Adam Lerer, Adam P Goucher, Adam Perelman, Aditya Ramesh, Aidan Clark, AJ Ostrow, Akila Welihinda, Alan Hayes, Alec Radford, et al. Gpt-4o system card.arXiv preprint arXiv:2410.21276, 2024
work page internal anchor Pith review Pith/arXiv arXiv 2024
-
[43]
Update to gpt-5 system card: Gpt-5.2, December 2025
OpenAI. Update to gpt-5 system card: Gpt-5.2, December 2025. URL https://cdn.openai. com/pdf/3a4153c8-c748-4b71-8e31-aecbde944f8d/oai_5_2_system-card.pdf
work page 2025
-
[44]
Gemini 3.1 flash lite model card, 2025
Google DeepMind. Gemini 3.1 flash lite model card, 2025. URL https://storage.googleapis.com/deepmind-media/Model-Cards/ Gemini-3-1-Flash-Lite-Model-Card.pdf. 12
work page 2025
-
[45]
An Yang, Anfeng Li, Baosong Yang, Beichen Zhang, Binyuan Hui, Bo Zheng, Bowen Yu, Chang Gao, Chengen Huang, Chenxu Lv, et al. Qwen3 technical report.arXiv preprint arXiv:2505.09388, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[46]
Aaron Grattafiori, Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Alex Vaughan, et al. The llama 3 herd of models.arXiv preprint arXiv:2407.21783, 2024
work page internal anchor Pith review Pith/arXiv arXiv 2024
-
[47]
Gemma Team, Aishwarya Kamath, Johan Ferret, Shreya Pathak, Nino Vieillard, Ramona Merhej, Sarah Perrin, Tatiana Matejovicova, Alexandre Ramé, Morgane Rivière, et al. Gemma 3 technical report.arXiv preprint arXiv:2503.19786, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[48]
DeepSeek-R1: Incentivizing Reasoning Capability in LLMs via Reinforcement Learning
Daya Guo, Dejian Yang, Haowei Zhang, Junxiao Song, Peiyi Wang, Qihao Zhu, Runxin Xu, Ruoyu Zhang, Shirong Ma, Xiao Bi, et al. Deepseek-r1: Incentivizing reasoning capability in llms via reinforcement learning.arXiv preprint arXiv:2501.12948, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[49]
Shuai Bai, Yuxuan Cai, Ruizhe Chen, Keqin Chen, Xionghui Chen, Zesen Cheng, Lianghao Deng, Wei Ding, Chang Gao, Chunjiang Ge, et al. Qwen3-vl technical report.arXiv preprint arXiv:2511.21631, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[50]
Visual instruction tuning.Advances in neural information processing systems, 36:34892–34916, 2023
Haotian Liu, Chunyuan Li, Qingyang Wu, and Yong Jae Lee. Visual instruction tuning.Advances in neural information processing systems, 36:34892–34916, 2023
work page 2023
-
[51]
Molrag: unlocking the power of large language models for molecular property prediction
Ziting Xian, Jiawei Gu, Lingbo Li, and Shangsong Liang. Molrag: unlocking the power of large language models for molecular property prediction. InProceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pages 15513–15531, 2025
work page 2025
-
[52]
Zaifei Yang, Hong Chang, Ruibing Hou, Shiguang Shan, and Xilin Chen. Knowmol: Advanc- ing molecular large language models with multi-level chemical knowledge.arXiv preprint arXiv:2510.19484, 2025
-
[53]
Iurii Sushko, Elena Salmina, Vladimir A Potemkin, Gennadiy Poda, and Igor V Tetko. Toxalerts: a web server of structural alerts for toxic chemicals and compounds with potential adverse reactions, 2012. 13 AT oxicityCliffConstruction Details Table A: Statistics ofT oxicityCliff, including the number of endpoints, molecules, toxicity cliff pairs, and result...
-
[54]
Focus ontoxicity-associated fragment identification. - Identify the fragment(s) that are specific to the toxic molecule and are most likely responsible for the toxicity signal. - If multiple fragments are required, return all of them. - Preserve the original SAFE fragment format exactly
-
[55]
- If there are multiple fragments, concatenate them as a dot-separated SAFE string
Output format constraints: - Return the answer as thetoxic-only SAFE fragment string. - If there are multiple fragments, concatenate them as a dot-separated SAFE string. - Do not paraphrase fragment content or convert it into natural language
-
[56]
Response format: { "answer": "..." } HARD CONSTRAINTS: - Output ONLY the JSON object. - Do not include explanations, markdown, or extra text. - Do not add any extra keys. - The value of "answer" must be the toxic-only SAFE fragment string exactly. Table H:Task 1 system prompt for toxic fragment identification. 34 Task 2SYSTEM PROMPT You are a molecular to...
-
[57]
- Generate the fragment(s) that can replace the toxic fragment(s) while reducing toxicity
Focus onnon-toxic fragment generation. - Generate the fragment(s) that can replace the toxic fragment(s) while reducing toxicity. - If multiple fragments are required, return all of them. - Preserve the original SAFE fragment format exactly
-
[58]
- If there are multiple fragments, concatenate them as a dot-separated SAFE string
Output format constraints: - Return the answer as thenon-toxic-only SAFE fragment string. - If there are multiple fragments, concatenate them as a dot-separated SAFE string. - Do not paraphrase fragment content or convert it into natural language
-
[59]
Response format: { "answer": "..." } HARD CONSTRAINTS: - Output ONLY the JSON object. - Do not include explanations, markdown, or extra text. - Do not add any extra keys. - The value of "answer" must be the non-toxic-only SAFE fragment string exactly. Table I:Task 2 system prompt for non-toxic fragment generation. 35 Task 3 SMILES GenerationSYSTEM PROMPT ...
-
[60]
- Generate a chemically plausible non-toxic molecule
Focus onnon-toxic molecule generation. - Generate a chemically plausible non-toxic molecule. - Reduce toxicity while preserving the original molecular characteristics as much as possible. - Return the final molecule, not intermediate fragments
-
[61]
- Do not return SAFE fragments
Output format constraints: - Return the answer as asingle non-toxic molecule SMILES string. - Do not return SAFE fragments. - Do not return multiple candidates
-
[62]
Response format: { "answer": "..." } HARD CONSTRAINTS: - Output ONLY the JSON object. - Do not include explanations, markdown, or extra text. - Do not add any extra keys. - The value of "answer" must be the final non-toxic molecule SMILES string. Table J:Task 3 system prompt for direct non-toxic molecule generation. 36 Task 3 SAFE GenerationSYSTEM PROMPT ...
-
[63]
- Generate the full SAFE representation of the resulting non-toxic molecule
Focus onnon-toxic SAFE generation. - Generate the full SAFE representation of the resulting non-toxic molecule. - Reduce toxicity while preserving the original molecular characteristics as much as possible. - Return the complete molecule-level SAFE representation, not only edited fragments
-
[64]
- If multiple fragments are present, concatenate them as a dot-separated SAFE string
Output format constraints: - Return the answer as thefull non-toxic SAFE string. - If multiple fragments are present, concatenate them as a dot-separated SAFE string. - Do not paraphrase fragment content or convert it into natural language
-
[65]
Response format: { "answer": "..." } HARD CONSTRAINTS: - Output ONLY the JSON object. - Do not include explanations, markdown, or extra text. - Do not add any extra keys. - The value of "answer" must be the final full non-toxic SAFE string for the whole molecule. Table K:Task 3 system prompt for full non-toxic SAFE generation. 37 Task 3 Step-wise CoT SAFE...
-
[66]
- First identify the toxic fragment(s) most likely associated with toxicity
Usestep-wise reasoning. - First identify the toxic fragment(s) most likely associated with toxicity. - Then generate the corresponding non-toxic replacement fragment(s). - Finally generate the full non-toxic molecule as a SAFE string
-
[67]
- The JSON must include the final answer and all required intermediate fields
Output format constraints: - Return a single JSON object. - The JSON must include the final answer and all required intermediate fields. - Do not omit any required fields
-
[68]
Response format: { "answer": "...", "step1_only_toxic_safe_fragments": "...", "step1_reasoning": "...", "step2_only_nontoxic_safe_fragments": "...", "step2_reasoning": "...", "step3_reasoning": "..." } HARD CONSTRAINTS: - Output ONLY the JSON object. - Do not include markdown or extra text outside the JSON. - The value of "answer" must be the final full n...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.