Recognition: no theorem link
Pretraining Strategies and Scaling for ECG Foundation Models: A Systematic Study
Pith reviewed 2026-05-13 03:52 UTC · model grok-4.3
The pith
Structured state space models outperform transformers and CNNs for ECG foundation models because of their inductive biases rather than pretraining scale alone.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The authors establish that structured state space models deliver superior transferable representations for ECG signals compared with transformers and CNNs when pretrained with the same contrastive or non-contrastive objectives, and that this advantage persists and even strengthens as the pretraining corpus grows to 11 million samples. They conclude that the strong inductive biases of structured state space models, rather than pretraining scale or objective alone, are the primary driver of effective representation learning in this domain.
What carries the argument
Structured state space models, which embed explicit assumptions about the structure of sequential physiological signals to enable efficient modeling of long-range dependencies.
If this is right
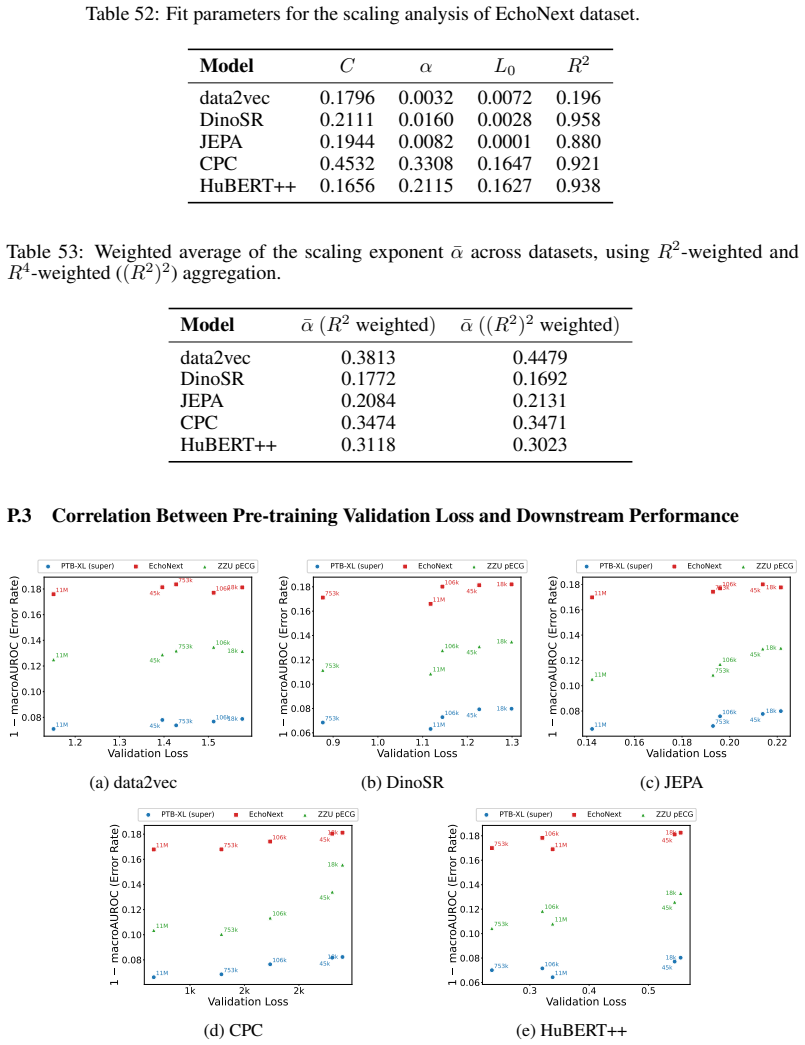
- Contrastive predictive coding produces the most transferable representations across the tested clinical tasks.
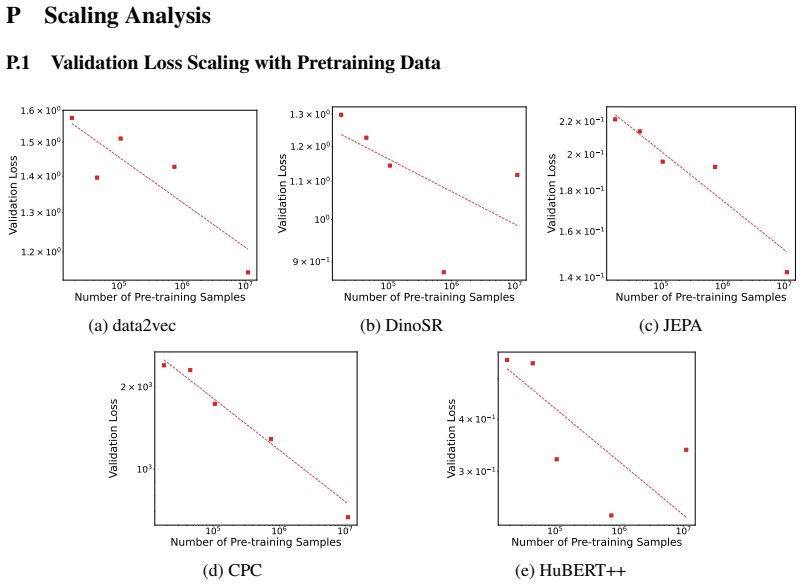
- Scaling the pretraining dataset to 11 million samples yields continued gains for most pretraining objectives.
- Architecture choice exerts a larger and more consistent effect on downstream performance than the choice of pretraining objective.
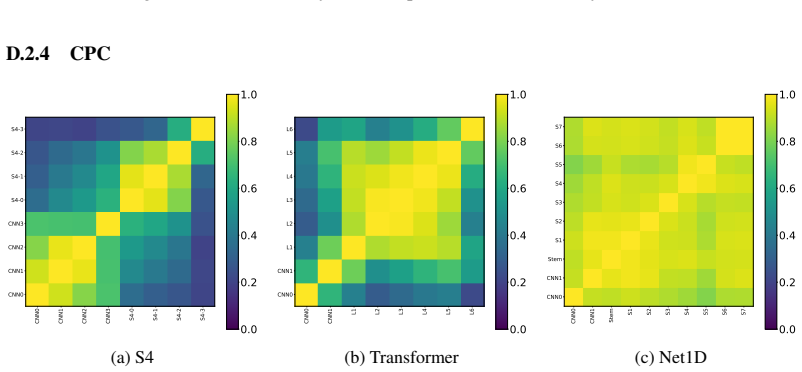
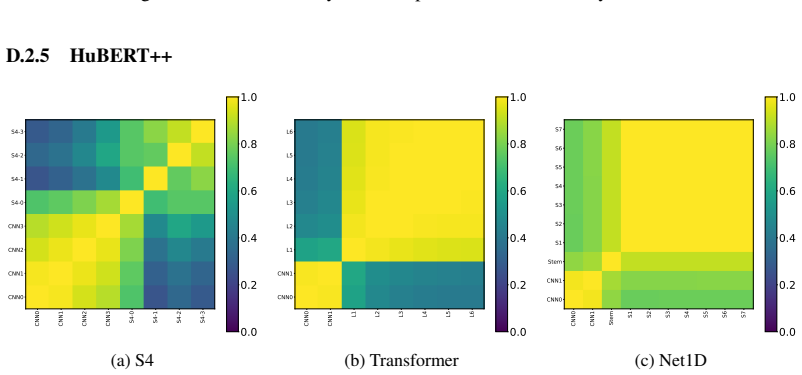
- The superiority of structured state space models holds across all five pretraining strategies examined.
Where Pith is reading between the lines
- The same inductive-bias advantage may favor structured state space models for other physiological time series such as EEG or PPG.
- Future foundation-model work in this area could test whether even larger state space models continue to improve without requiring proportionally larger datasets.
- Resource-constrained settings may achieve strong results by prioritizing state space architectures over simply collecting more unlabeled ECG data.
Load-bearing premise
That performance differences between architectures and pretraining methods are caused primarily by inductive biases rather than unmeasured differences in hyperparameters, preprocessing pipelines, or dataset-specific artifacts.
What would settle it
A controlled re-run in which transformers and CNNs are trained with identical hyperparameters, identical preprocessing, and the same random seeds as the state space models and still underperform on the same downstream tasks.
Figures








read the original abstract
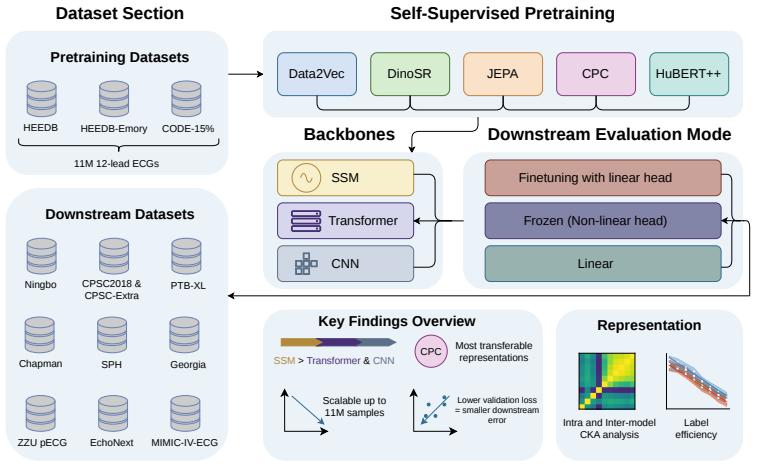
Specialized foundation models are beginning to emerge in various medical subdomains, but pretraining methodologies and parametric scaling with the size of the pretraining dataset are rarely assessed systematically and in a like-for-like manner. This work focuses on foundation models for electrocardiography (ECG) data, one of the most widely captured physiological time series world-wide. We present a comprehensive assessment of pretraining methodologies, covering five different contrastive and non-contrastive self-supervised learning objectives for ECG foundation models, and investigate their scaling behavior with pretraining dataset sizes up to 11M input samples, exclusively from publicly available sources. Pretraining strategy has a meaningful and consistent impact on downstream performance, with contrastive predictive coding (slightly ahead of JEPA) yielding the most transferable representations across diverse clinical tasks. Scaling pretraining data continues to yield meaningful improvements up to 11M samples for most objectives. We also compare model architectures across all pretraining methodologies and find evidence for a clear superiority of structured state space models compared to transformers and CNN models. We hypothesize that the strong inductive biases of structured state space models, rather than pretraining scale alone, are the primary driver of effective ECG representation learning, with important implications for future foundation model development in this and potentially other physiological signal domains.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript conducts a systematic empirical study of five self-supervised pretraining objectives (contrastive and non-contrastive) for ECG foundation models, using exclusively public data up to 11M samples. It reports scaling curves showing continued gains for most objectives, identifies contrastive predictive coding (slightly ahead of JEPA) as yielding the most transferable representations on downstream clinical tasks, and compares architectures (structured state space models vs. transformers vs. CNNs) across pretraining methods, finding clear SSM superiority. The central hypothesis is that SSM inductive biases, rather than pretraining scale alone, are the primary driver of effective ECG representation learning.
Significance. If the results hold under controlled conditions, the work supplies a valuable like-for-like benchmark for ECG pretraining strategies and scaling behavior that is currently rare in the domain. The architecture comparison, if shown to be scale-independent, would provide concrete evidence favoring strong inductive biases over pure scale in physiological time-series foundation models, with direct implications for model design choices in related medical signal domains.
major comments (2)
- [Architecture comparison results] The architecture comparison (reported as showing 'clear superiority' of SSMs across all pretraining methodologies) appears to have been performed at a single fixed scale rather than with per-architecture scaling curves at multiple data regimes. This design choice leaves the central hypothesis—that inductive biases rather than scale are primary—unsupported by direct evidence, as the performance gap could be an artifact of the particular (largest) scale chosen.
- [Downstream evaluation setup] The downstream clinical tasks used to evaluate transferability are not accompanied by an explicit justification or sensitivity analysis showing they are representative of real-world ECG use cases; without this, differences attributed to inductive biases could be confounded by unmeasured factors such as task-specific signal characteristics or preprocessing artifacts.
minor comments (2)
- [Abstract] The abstract states that five objectives were studied but does not name them; an explicit list would improve readability.
- [Figures] Figure captions and legends should consistently report the number of random seeds or runs used to compute means and error bars.
Simulated Author's Rebuttal
We thank the referee for the constructive feedback on our systematic study of ECG pretraining strategies and scaling. We address each major comment below with clarifications on our experimental choices and proposed revisions to strengthen the manuscript. We believe these changes will better support the central claims while acknowledging limitations in the current design.
read point-by-point responses
-
Referee: [Architecture comparison results] The architecture comparison (reported as showing 'clear superiority' of SSMs across all pretraining methodologies) appears to have been performed at a single fixed scale rather than with per-architecture scaling curves at multiple data regimes. This design choice leaves the central hypothesis—that inductive biases rather than scale are primary—unsupported by direct evidence, as the performance gap could be an artifact of the particular (largest) scale chosen.
Authors: We agree that performing architecture comparisons at multiple data regimes with dedicated scaling curves would provide stronger direct support for the hypothesis that SSM inductive biases are the primary driver independent of scale. Our current design compared architectures at the largest scale (11M samples) after observing continued gains from scaling the pretraining objectives, with the goal of evaluating representations under conditions where data scale has been maximized. The consistent SSM advantage across all five pretraining methods at this scale suggests the gap is not an artifact of a single objective. We will revise the manuscript to explicitly state the fixed scale used for architecture comparisons, add a dedicated limitations paragraph discussing the absence of per-architecture scaling curves, and include a forward-looking statement on the value of such experiments in future work. No new experiments are feasible within the revision timeline due to computational requirements. revision: partial
-
Referee: [Downstream evaluation setup] The downstream clinical tasks used to evaluate transferability are not accompanied by an explicit justification or sensitivity analysis showing they are representative of real-world ECG use cases; without this, differences attributed to inductive biases could be confounded by unmeasured factors such as task-specific signal characteristics or preprocessing artifacts.
Authors: We appreciate this observation. The downstream tasks were chosen to span a range of clinically relevant ECG applications drawn from established benchmarks in the literature (e.g., arrhythmia classification, myocardial infarction detection, and rhythm analysis on datasets such as PTB-XL and others). We will add a new subsection in the experimental setup that provides explicit justification for each task, including references to prior ECG foundation model evaluations and clinical guidelines. We will also incorporate a sensitivity analysis (e.g., results under alternative preprocessing pipelines and task subsets) to demonstrate robustness. These additions will be included in the revised manuscript. revision: yes
Circularity Check
No circularity: purely empirical comparisons with no derivations or self-referential reductions
full rationale
This is an empirical study that evaluates five SSL objectives, scaling curves up to 11M samples, and architecture ablations (SSM vs transformer vs CNN) on held-out clinical downstream tasks using public ECG data. No equations, derivations, or 'predictions' are presented that could reduce to fitted inputs by construction. The hypothesis about SSM inductive biases is framed as an interpretation of experimental results rather than a mathematical claim. No self-citation chains are load-bearing for any central result, and the work does not rename known patterns or smuggle ansatzes. The design is self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Self-supervised learning objectives produce representations that transfer to downstream clinical tasks.
Reference graph
Works this paper leans on
-
[1]
On the Opportunities and Risks of Foundation Models
Rishi Bommasani, Drew A Hudson, Ehsan Adeli, Russ Altman, Simran Arora, Sydney von Arx, Michael S Bernstein, Jeannette Bohg, Antoine Bosselut, Emma Brunskill, et al. On the Opportunities and Risks of Foundation Models.arXiv preprint arXiv:2108.07258, 2021
work page internal anchor Pith review Pith/arXiv arXiv 2021
-
[2]
Richard J Chen, Tong Ding, Ming Y Lu, Drew FK Williamson, Guillaume Jaume, Andrew H Song, Bowen Chen, Andrew Zhang, Daniel Shao, Muhammad Shaban, et al. Towards a General-Purpose Foundation Model for Computational Pathology.Nature medicine, 30(3):850–862, 2024
work page 2024
-
[3]
Yukun Zhou, Mark A Chia, Siegfried K Wagner, Murat S Ayhan, Dominic J Williamson, Robbert R Struyven, Timing Liu, Moucheng Xu, Mateo G Lozano, Peter Woodward-Court, et al. A foundation model for generalizable disease detection from retinal images.Nature, 622(7981):156–163, 2023
work page 2023
-
[4]
Konstantinos C Siontis, Peter A Noseworthy, Zachi I Attia, and Paul A Friedman. Artificial intelligence- enhanced electrocardiography in cardiovascular disease management.Nature Reviews Cardiology, 18(7): 465–478, 2021
work page 2021
-
[5]
Nils Strodthoff, Patrick Wagner, Tobias Schaeffter, and Wojciech Samek. Deep learning for ECG analysis: Benchmarks and insights from PTB-XL.IEEE journal of biomedical and health informatics, 25(5): 1519–1528, 2020
work page 2020
-
[6]
R Sacha Bhatia and Paul Dorian. Screening for cardiovascular disease risk with electrocardiography.JAMA Internal Medicine, 178(9):1163–1164, 2018
work page 2018
-
[7]
Ivan C Rokos, William J French, Amal Mattu, Graham Nichol, Michael E Farkouh, James Reiffel, and Gregg W Stone. Appropriate cardiac cath lab activation: optimizing electrocardiogram interpretation and clinical decision-making for acute ST-elevation myocardial infarction.American heart journal, 160(6): 995–1003, 2010
work page 2010
-
[8]
Jun Li, Aaron D Aguirre, Valdery Moura Junior, Jiarui Jin, Che Liu, Lanhai Zhong, Chenxi Sun, Gari Clifford, M Brandon Westover, and Shenda Hong. An Electrocardiogram Foundation Model Built on over 10 Million Recordings.NEJM AI, 2(7):AIoa2401033, 2025
work page 2025
-
[9]
Sehun Kim. Learning general representation of 12-lead electrocardiogram with a joint-embedding predic- tive architecture.arXiv preprint arXiv:2410.08559, 2024
work page internal anchor Pith review Pith/arXiv arXiv 2024
-
[10]
Guiding Masked Representation Learning to Capture Spatio-Temporal Relationship of Electrocardiogram
Yeongyeon Na, Minje Park, Yunwon Tae, and Sunghoon Joo. Guiding Masked Representation Learning to Capture Spatio-Temporal Relationship of Electrocardiogram. InThe Twelfth International Conference on Learning Representations, 2024
work page 2024
-
[11]
Zero-shot ECG classification with multimodal learning and test-time clinical knowledge enhancement
Che Liu, Zhongwei Wan, Cheng Ouyang, Anand Shah, Wenjia Bai, and Rossella Arcucci. Zero-shot ECG classification with multimodal learning and test-time clinical knowledge enhancement. InProceedings of the 41st International Conference on Machine Learning, pages 31949–31963, 2024
work page 2024
-
[12]
Yuanyuan Tian, Zhiyuan Li, Yanrui Jin, Mengxiao Wang, Xiaoyang Wei, Liqun Zhao, Yunqing Liu, Jinlei Liu, and Chengliang Liu. Foundation model of ECG diagnosis: Diagnostics and explanations of any form and rhythm on ECG.Cell Reports Medicine, 5(12), 2024
work page 2024
-
[13]
Edoardo Coppola, Mattia Savardi, Mauro Massussi, Marianna Adamo, Marco Metra, and Alberto Signoroni. HuBERT-ECG as a self-supervised foundation model for broad and scalable cardiac applications.medRxiv, pages 2024–11, 2024
work page 2024
-
[14]
Ecg-fm: An open electrocardiogram foundation model.Jamia Open, 8(5):ooaf122, 2025
Kaden McKeen, Sameer Masood, Augustin Toma, Barry Rubin, and Bo Wang. Ecg-fm: An open electrocardiogram foundation model.Jamia Open, 8(5):ooaf122, 2025
work page 2025
-
[15]
Boosting Masked ECG-Text Auto-Encoders as Discrimi- native Learners
Manh Pham Hung, Aaqib Saeed, and Dong Ma. Boosting Masked ECG-Text Auto-Encoders as Discrimi- native Learners. InProceedings of the 42nd International Conference on Machine Learning, 2025
work page 2025
-
[16]
Riccardo Lunelli, Angus Nicolson, Samuel Martin Pröll, Sebastian Johannes Reinstadler, Axel Bauer, and Clemens Dlaska. BenchECG and xECG: a benchmark and baseline for ECG foundation models.arXiv preprint arXiv:2509.10151, 2025. 10
-
[17]
Alexis Nolin-Lapalme, Achille Sowa, Jacques Delfrate, Olivier Tastet, Denis Corbin, Merve Kulbay, Derman Ozdemir, Marie-Jeanne Noël, François-Christophe Marois-Blanchet, François Harvey, et al. Foundation models for electrocardiogram interpretation: clinical implications.European Heart Journal, page ehaf1119, 2026
work page 2026
-
[18]
Han Yu, Peikun Guo, and Akane Sano. ECG semantic integrator (ESI): A foundation ECG model pretrained with LLM-enhanced cardiological text.Transactions on Machine Learning Research, 2024. ISSN 2835-8856
work page 2024
-
[19]
PhysioWave: A Multi-Scale Wavelet-Transformer for Physiological Signal Representation
Yanlong Chen, Mattia Orlandi, Pierangelo Maria Rapa, Simone Benatti, Luca Benini, and Yawei Li. PhysioWave: A Multi-Scale Wavelet-Transformer for Physiological Signal Representation. InThe Thirty- ninth Annual Conference on Neural Information Processing Systems, 2025
work page 2025
-
[20]
Benchmarking ECG FMs: A Reality Check Across Clinical Tasks
M A Al-Masud, Juan Lopez Alcaraz, and Nils Strodthoff. Benchmarking ECG FMs: A Reality Check Across Clinical Tasks. InThe Fourteenth International Conference on Learning Representations, 2026
work page 2026
-
[21]
Zhijiang Wan, Qianhao Yu, Jia Mao, Wenfeng Duan, and Cheng Ding. OpenECG: Benchmarking ECG Foundation Models with Public 1.2 Million Records.arXiv preprint arXiv:2503.00711, 2025. URL https://arxiv.org/abs/2503.00711
-
[22]
Representation Learning with Contrastive Predictive Coding
Aaron van den Oord, Yazhe Li, and Oriol Vinyals. Representation learning with contrastive predictive coding.arXiv preprint arXiv:1807.03748, 2018
work page internal anchor Pith review Pith/arXiv arXiv 2018
-
[23]
Wei-Ning Hsu, Benjamin Bolte, Yao-Hung Hubert Tsai, Kushal Lakhotia, Ruslan Salakhutdinov, and Abdelrahman Mohamed. Hubert: Self-supervised speech representation learning by masked prediction of hidden units.IEEE/ACM transactions on audio, speech, and language processing, 29:3451–3460, 2021
work page 2021
-
[24]
Self-supervised learning from images with a joint-embedding predictive architecture
Mahmoud Assran, Quentin Duval, Ishan Misra, Piotr Bojanowski, Pascal Vincent, Michael Rabbat, Yann LeCun, and Nicolas Ballas. Self-supervised learning from images with a joint-embedding predictive architecture. InProceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pages 15619–15629, 2023
work page 2023
-
[25]
Attention is all you need.Advances in neural information processing systems, 30, 2017
Ashish Vaswani, Noam Shazeer, Niki Parmar, Jakob Uszkoreit, Llion Jones, Aidan N Gomez, Łukasz Kaiser, and Illia Polosukhin. Attention is all you need.Advances in neural information processing systems, 30, 2017
work page 2017
-
[26]
Efficiently Modeling Long Sequences with Structured State Spaces
Albert Gu, Karan Goel, and Christopher Re. Efficiently Modeling Long Sequences with Structured State Spaces. InInternational Conference on Learning Representations, 2022
work page 2022
-
[27]
Temesgen Mehari and Nils Strodthoff. Towards quantitative precision for ECG analysis: Leveraging state space models, self-supervision and patient metadata.IEEE journal of biomedical and health informatics, 27(11):5326–5334, 2023
work page 2023
-
[28]
Nils Strodthoff, Juan Miguel Lopez Alcaraz, and Wilhelm Haverkamp. Prospects for artificial intelligence- enhanced electrocardiogram as a unified screening tool for cardiac and non-cardiac conditions: an explo- rative study in emergency care.European Heart Journal - Digital Health, 5(4):454–460, 07 2024. ISSN 2634-3916. doi: 10.1093/ehjdh/ztae039. URLhttp...
-
[29]
Training compute-optimal large language models
Jordan Hoffmann, Sebastian Borgeaud, Arthur Mensch, Elena Buchatskaya, Trevor Cai, Eliza Rutherford, Diego de Las Casas, Lisa Anne Hendricks, Johannes Welbl, Aidan Clark, et al. Training compute-optimal large language models. InProceedings of the 36th International Conference on Neural Information Processing Systems, pages 30016–30030, 2022
work page 2022
-
[30]
Scaling Laws for Robust Comparison of Open Foundation Language-Vision Models and Datasets
Marianna Nezhurina, Tomer Porian, Giovanni Puccetti, Tommie Kerssies, Romain Beaumont, Mehdi Cherti, and Jenia Jitsev. Scaling Laws for Robust Comparison of Open Foundation Language-Vision Models and Datasets. InThe Thirty-ninth Annual Conference on Neural Information Processing Systems, 2025
work page 2025
-
[31]
Roformer: Enhanced transformer with rotary position embedding.Neurocomputing, 568:127063, 2024
Jianlin Su, Murtadha Ahmed, Yu Lu, Shengfeng Pan, Wen Bo, and Yunfeng Liu. Roformer: Enhanced transformer with rotary position embedding.Neurocomputing, 568:127063, 2024
work page 2024
-
[32]
Gaussian Error Linear Units (GELUs)
Dan Hendrycks and Kevin Gimpel. Gaussian error linear units (gelus).arXiv preprint arXiv:1606.08415, 2016
work page internal anchor Pith review Pith/arXiv arXiv 2016
-
[33]
Data2vec: A general framework for self-supervised learning in speech, vision and language
Alexei Baevski, Wei-Ning Hsu, Qiantong Xu, Arun Babu, Jiatao Gu, and Michael Auli. Data2vec: A general framework for self-supervised learning in speech, vision and language. InInternational conference on machine learning, pages 1298–1312. PMLR, 2022. 11
work page 2022
-
[34]
Alexander H Liu, Heng-Jui Chang, Michael Auli, Wei-Ning Hsu, and Jim Glass. Dinosr: Self-distillation and online clustering for self-supervised speech representation learning.Advances in Neural Information Processing Systems, 36:58346–58362, 2023
work page 2023
-
[35]
Marco Cuturi. Sinkhorn distances: Lightspeed computation of optimal transport.Advances in neural information processing systems, 26, 2013
work page 2013
-
[36]
Harvard- Emory ECG Database (version 5.0)
Zuzana Koscova, Valdery Moura Junior, Matthew Reyna, Shenda Hong, Aditya Gupta, Manohar Ghanta, Reza Sameni, Aaron Aguirre, Qiao Li, Sahar Zafar, Gari Clifford, and M Brandon Westover. Harvard- Emory ECG Database (version 5.0). Brain Data Science Platform, 2026
work page 2026
-
[37]
The harvard-emory ecg database
Zuzana Koscova, Qiao Li, Chad Robichaux, Valdery Moura Junior, Manohar Ghanta, Aditya Gupta, Jonathan Rosand, Aaron D Aguirre, Erik Reinertsen, Steven Song, et al. The harvard-emory ecg database. Scientific Data, 2026
work page 2026
-
[38]
Antônio H. Ribeiro, Gabriela M.M. Paixao, Emilly M. Lima, Manoel Horta Ribeiro, Marcelo M. Pinto Filho, Paulo R. Gomes, Derick M. Oliveira, Wagner Meira Jr, Thömas B Schon, and Antonio Luiz P. Ribeiro. CODE-15%: a large scale annotated dataset of 12-lead ECGs , June 2021
work page 2021
-
[39]
MIMIC-IV-ECG: Diagnostic Electrocardiogram Matched Subset.PhysioNet, September 2023
Brian Gow, Tom Pollard, Larry A Nathanson, Alistair Johnson, Benjamin Moody, Chrystinne Fernandes, Nathaniel Greenbaum, Jonathan W Waks, Parastou Eslami, Tanner Carbonati, Ashish Chaudhari, Elizabeth Herbst, Dana Moukheiber, Seth Berkowitz, Roger Mark, and Steven Horng. MIMIC-IV-ECG: Diagnostic Electrocardiogram Matched Subset.PhysioNet, September 2023. V...
work page 2023
-
[40]
V-JEPA: Latent Video Prediction for Visual Representation Learning
Antoine Bardes, Mehdi Mirza, Boris Oreshkin, Michael Auli, Ishan Misra, and Yann LeCun. V-JEPA: Latent Video Prediction for Visual Representation Learning. InInternational Conference on Learning Representations (ICLR), 2024
work page 2024
-
[41]
Matthew B McDermott, Haoran Zhang, Lasse H Hansen, Giovanni Angelotti, and Jack Gallifant. A closer look at AUROC and AUPRC under class imbalance.Advances in Neural Information Processing Systems, 37:44102–44163, 2024
work page 2024
-
[42]
Universal language model fine-tuning for text classification
Jeremy Howard and Sebastian Ruder. Universal language model fine-tuning for text classification. In Proceedings of the 56th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pages 328–339, 2018
work page 2018
-
[44]
URLhttps://arxiv.org/abs/2604.23385
work page internal anchor Pith review Pith/arXiv arXiv
-
[45]
Mathilde Caron, Hugo Touvron, Ishan Misra, Hervè Jègou, Julien Mairal, Piotr Bojanowski, and Armand Joulin. Emerging properties in self-supervised vision transformers.Proceedings of the IEEE/CVF international conference on computer vision, pages 9650–9660, 2021
work page 2021
-
[46]
Timothée Darcet, Federico Baldassarre, Maxime Oquab, Julien Mairal, and Piotr Bojanowski. Cluster and Predict Latents Patches for Improved Masked Image Modeling.Transactions on Machine Learning Research, 2025. ISSN 2835-8856. URLhttps://openreview.net/forum?id=Ycmz7qJxUQ
work page 2025
-
[47]
M. A. Reyna, N. Sadr, A. Gu, E. A. Perez Alday, C. Liu, S. Seyedi, A. Shah, and G. D. Clifford. Will Two Do? Varying Dimensions in Electrocardiography: The PhysioNet/Computing in Cardiology Challenge 2021 (version 1.0.3). https://doi.org/10.13026/34va-7q14, 2022. PhysioNet. RRID:SCR_007345
-
[48]
M. A. Reyna, N. Sadr, E. A. Perez Alday, A. Gu, A. J. Shah, C. Robichaux, A. B. Rad, A. Elola, S. Seyedi, S. Ansari, H. Ghanbari, Q. Li, A. Sharma, and G. D. Clifford. Will Two Do? Varying Dimensions in Electrocardiography: The PhysioNet/Computing in Cardiology Challenge 2021. In2021 Computing in Cardiology (CinC), pages 1–4, Brno, Czech Republic, 2021. d...
-
[49]
Jianwei Zheng, Jianming Zhang, Sidy Danioko, Hai Yao, Hangyuan Guo, and Cyril Rakovski. A 12-lead electrocardiogram database for arrhythmia research covering more than 10, 000 patients.Scientific Data, 7 (1), February 2020. ISSN 2052-4463. doi: 10.1038/s41597-020-0386-x. URL http://dx.doi.org/10. 1038/s41597-020-0386-x
-
[50]
A large-scale multi-label 12-lead electrocardiogram database with standardized diagnostic statements
Hui Liu, Dan Chen, Da Chen, Xiyu Zhang, Huijie Li, Lipan Bian, Minglei Shu, and Yinglong Wang. A large-scale multi-label 12-lead electrocardiogram database with standardized diagnostic statements. Scientific data, 9(1):272, 2022. doi: doi.org/10.1038/s41597-022-01403-5. URL https://doi.org/10. 1038/s41597-022-01403-5. 12
-
[51]
Hui Liu, Yinglong Wang, Da Chen, Xiyu Zhang, Huijie Li, Lipan Bian, Minglei Shu, and Dan Chen. A large-scale multi-label 12-lead electrocardiogram database with standardized diagnostic statements, 2022
work page 2022
-
[52]
PTB-XL, a large publicly available electrocardiography dataset (version 1.0.3)
Philipp Wagner, Nils Strodthoff, Ralf Bousseljot, Wojciech Samek, and Tobias Schaeffter. PTB-XL, a large publicly available electrocardiography dataset (version 1.0.3). https://doi.org/10.13026/ kfzx-aw45, 2022. PhysioNet. RRID:SCR_007345
work page 2022
-
[53]
PTB-XL, a large publicly available electrocardiography dataset.Scientific data, 7 (1):1–15, 2020
Patrick Wagner, Nils Strodthoff, Ralf-Dieter Bousseljot, Dieter Kreiseler, Fatima I Lunze, Wojciech Samek, and Tobias Schaeffter. PTB-XL, a large publicly available electrocardiography dataset.Scientific data, 7 (1):1–15, 2020. doi: 10.1038/s41597-020-0495-6
-
[54]
Jian Tan, Haoyi Fan, Jiawei Luo, Yanjie Zhou, Ning Wang, Xizheng Wang, Guizhi Liu, Chengyu Liu, and Zongmin Wang. A pediatric ECG database with disease diagnosis covering 11643 children.Scientific Data, 12(1):867, 2025. doi: 10.1038/s41597-025-05225-z
-
[55]
A pediatric ECG database with disease diagnosis covering 11643 children, 5 2025
Tan Jian, Haoyi Fan, Jiawei Luo, Yanjie Zhou, Ning Wang, Xizheng Wang, Guizhi Liu, Chengyu Liu, and Zongmin Wang. A pediatric ECG database with disease diagnosis covering 11643 children, 5 2025. URL https://doi.org/10.6084/m9.figshare.27078763.v1
-
[56]
EchoNext: A Dataset for Detecting Echocardiogram-Confirmed Structural Heart Disease from ECGs
Pierre Elias and Joshua Finer. EchoNext: A Dataset for Detecting Echocardiogram-Confirmed Structural Heart Disease from ECGs. PhysioNet, 2025. URLhttps://doi.org/10.13026/r9pp-3y42
-
[57]
John Weston Hughes, Linyuan Jing, Joshua Finer, Dustin Hartzel, Christopher Kelsey, Aaron Long, Daniel Rocha, Jeffrey Ruhl, Timothy Poterucha, and Pierre Elias. EchoNext-Mini: A Dataset and Baseline AI Model for Detecting Structural Heart Disease from Electrocardiograms.NEJM AI, 3(5), April 2026. ISSN 2836-9386. doi: 10.1056/aidbp2500516. URLhttp://dx.doi...
-
[58]
We use exponential moving averages of a student network as prediction target (CAPI Figure
-
[59]
like I-JEPA, HuBERT, CAPI
-
[60]
To avoiding backpropagation through the clustering, we therefore need another non-SGD cluster update
We use clustering as loss (CAPI Figure 4) like CAPI. To avoiding backpropagation through the clustering, we therefore need another non-SGD cluster update
-
[61]
We use Sinkhorn-Knopp like DINO and unlike DinoSR, which encourages equiparticipation and prevents collapse
-
[62]
We use granular, soft prediction targets (unlike DinoSR), which would also allow us to use different temperatures (as in DINO)
-
[63]
We use cluster assignmenta using Sinkhorn-Knopp optimal transport (unlike CAPI, which uses a quite ad-hoc procedure). This has the nice side effect that prediction target computa- tion and cluster updates happen consistently S.2 Pseudo-code We provide pseudo-code for the three most crucial components of the algorithm. 1@torch.no_grad() 2def get_ema_target...
work page 1987
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.