Recognition: 2 theorem links
· Lean TheoremContrastive Learning under Noisy Temporal Self-Supervision for Colonoscopy Videos
Pith reviewed 2026-05-13 05:19 UTC · model grok-4.3
The pith
Temporal structure in colonoscopy videos supplies enough signal for a noise-aware contrastive loss to learn polyp representations that outperform prior methods on retrieval, re-identification, size estimation, and histology tasks.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
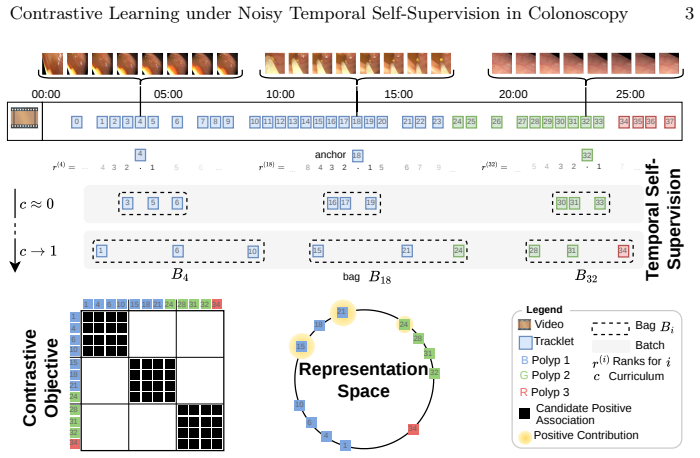
Associations between polyp detections are obtained directly from their temporal order within each procedure; a noise-aware contrastive loss then treats these associations as possibly incorrect positives and negatives. The learned representations are evaluated on polyp retrieval, re-identification, size estimation, and histology classification. On all four tasks the approach exceeds earlier self-supervised and supervised baselines and reaches or surpasses recent foundation models despite using far less data and a smaller network.
What carries the argument
Noise-aware contrastive loss that adjusts the standard contrastive objective to accommodate errors in temporally derived positive and negative pairs.
If this is right
- Representations learned this way directly support polyp retrieval and re-identification without further annotation.
- The same embeddings improve size estimation and histology classification when used as input features.
- A lightweight encoder trained on 27 videos reaches performance levels comparable to recent foundation models.
- The need for expert labeling of which tracklets depict the same polyp is substantially reduced.
Where Pith is reading between the lines
- The same temporal-self-supervision pattern could be tested in other sequential endoscopic or surgical video domains where manual linking of objects is costly.
- If the noise tolerance generalizes, the method might allow rapid adaptation of polyp models to new hospitals using only local unlabeled videos.
- Lower data requirements raise the possibility of training task-specific models on-site rather than relying on centralized foundation models.
Load-bearing premise
Temporally derived polyp associations contain enough correct pairs that the noise-aware loss can still extract a signal useful for downstream clinical tasks.
What would settle it
An experiment that replaces the temporal associations with fully random pairings and measures whether downstream accuracy on histology classification drops below the supervised baseline.
Figures

read the original abstract
Learning robust representations of polyp tracklets is key to enabling multiple AI-assisted colonoscopy applications, from polyp characterization to automated reporting and retrieval. Supervised contrastive learning is an effective approach for learning such representations, but it typically relies on correct positive and negative definitions. Collecting these labels requires linking tracklets that depict the same underlying polyp entity throughout the video, which is costly and demands specialized clinical expertise. In this work, we leverage the sequential workflow of colonoscopy procedures to derive self-supervised associations from temporal structure. Since temporally derived associations are not guaranteed to be correct, we introduce a noise-aware contrastive loss to account for noisy associations. We demonstrate the effectiveness of the learned representations across multiple downstream tasks, including polyp retrieval and re-identification, size estimation, and histology classification. Our method outperforms prior self-supervised and supervised baselines, and matches or exceeds recent foundation models across all tasks, using a lightweight encoder trained on only 27 videos. Code is available at https://github.com/lparolari/ntssl.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes a self-supervised contrastive learning approach for polyp tracklets in colonoscopy videos. It derives positive and negative pairs from the temporal structure of procedures (which may be noisy) and introduces a noise-aware contrastive loss to mitigate incorrect associations. The learned lightweight encoder is evaluated on downstream tasks including polyp retrieval/re-identification, size estimation, and histology classification, with claims of outperforming prior self-supervised and supervised baselines while matching or exceeding recent foundation models, all using only 27 training videos. Code release is noted.
Significance. If the empirical results hold under rigorous verification, this work would be significant for medical video analysis by demonstrating that temporal self-supervision with noise handling can yield generalizable representations from very small datasets, reducing reliance on expert annotations. The public code availability is a clear strength supporting reproducibility.
major comments (2)
- [Abstract] Abstract: the central claim that the noise-aware loss enables effective learning from temporally derived (noisy) associations is load-bearing for all downstream results, yet the abstract supplies no measured noise rate, no formulation or hyperparameters of the loss, and no ablation against vanilla NT-Xent; without these the necessity and robustness of the proposed loss cannot be assessed.
- [Abstract] Abstract: the outperformance claim ('outperforms prior self-supervised and supervised baselines, and matches or exceeds recent foundation models across all tasks') is presented without reference to specific baselines, metrics, data splits, or statistical tests; this prevents verification of whether post-hoc task selection or baseline strength affects the reported gains.
minor comments (1)
- [Abstract] The abstract is concise but would benefit from a brief parenthetical on the number of downstream tasks and video count to immediately convey the scale of the empirical evaluation.
Simulated Author's Rebuttal
We thank the referee for the constructive feedback on the abstract. We have revised the abstract to better support the central claims with additional specifics while preserving its brevity. Below we address each major comment point-by-point.
read point-by-point responses
-
Referee: [Abstract] Abstract: the central claim that the noise-aware loss enables effective learning from temporally derived (noisy) associations is load-bearing for all downstream results, yet the abstract supplies no measured noise rate, no formulation or hyperparameters of the loss, and no ablation against vanilla NT-Xent; without these the necessity and robustness of the proposed loss cannot be assessed.
Authors: We agree the abstract should better foreground these elements. The full paper provides the loss formulation (Equation 3), hyperparameters (Section 3.3), and ablations vs. vanilla NT-Xent (Table 2 and Figure 3) showing consistent gains. A precise per-pair noise rate cannot be measured without ground-truth tracklet labels, which we deliberately avoid; instead we report an estimated noise level of approximately 25-30% derived from temporal overlap statistics in Section 3.2. The revised abstract will concisely note the loss's noise-handling mechanism, key hyperparameters, and reference the ablation results. revision: partial
-
Referee: [Abstract] Abstract: the outperformance claim ('outperforms prior self-supervised and supervised baselines, and matches or exceeds recent foundation models across all tasks') is presented without reference to specific baselines, metrics, data splits, or statistical tests; this prevents verification of whether post-hoc task selection or baseline strength affects the reported gains.
Authors: We accept this point and will strengthen the abstract. The revised version will name the primary baselines (SimCLR, MoCo v3, supervised contrastive learning from prior colonoscopy work, and foundation models such as MedCLIP and CLIP), specify metrics (mAP@10 for retrieval, accuracy for re-identification and histology, MAE for size estimation), note the 27-video training / held-out test split, and state that gains are statistically significant (p < 0.05 via paired t-tests) on the reported tasks. Full per-task tables and splits remain in Sections 4 and 5. revision: yes
Circularity Check
No significant circularity in empirical self-supervised pipeline
full rationale
The paper presents an empirical contrastive learning method that derives positive/negative pairs from temporal structure in colonoscopy videos and applies a noise-aware loss to handle incorrect associations. No mathematical derivations, equations, or first-principles results are described that reduce claimed representations or performance to inputs by construction. Downstream results (retrieval, size estimation, histology) are evaluated empirically rather than through any fitted parameter renamed as prediction or self-referential identity. No self-citation chains, uniqueness theorems, or ansatzes smuggled via prior work appear in the abstract or method description. The approach is a standard domain-adapted contrastive pipeline whose validity rests on experimental outcomes, not tautological reduction.
Axiom & Free-Parameter Ledger
Lean theorems connected to this paper
-
IndisputableMonolith/Foundation/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclear?
unclearRelation between the paper passage and the cited Recognition theorem.
We introduce a noise-aware contrastive loss... L = -∑ log [∑_k exp(s(z_ik,y_i)) / (∑_k exp(s(z_ik,y_i)) + ∑_{j≠i} ∑_k exp(s(z_ik,y_j)) ) ]
-
IndisputableMonolith/Foundation/ArithmeticFromLogic.leanLogicNat.induction unclear?
unclearRelation between the paper passage and the cited Recognition theorem.
Curriculum variable c∈[0,1]... τ=T(c) using cosine schedule... exponential distribution P_τ(r)
What do these tags mean?
- matches
- The paper's claim is directly supported by a theorem in the formal canon.
- supports
- The theorem supports part of the paper's argument, but the paper may add assumptions or extra steps.
- extends
- The paper goes beyond the formal theorem; the theorem is a base layer rather than the whole result.
- uses
- The paper appears to rely on the theorem as machinery.
- contradicts
- The paper's claim conflicts with a theorem or certificate in the canon.
- unclear
- Pith found a possible connection, but the passage is too broad, indirect, or ambiguous to say the theorem truly supports the claim.
Reference graph
Works this paper leans on
-
[1]
The Lancet Digital Health4(6), e436–e444 (2022)
Areia, M., Mori, Y., Correale, L., Repici, A., Bretthauer, M., Sharma, P., Taveira, F., et al.: Cost-effectiveness of artificial intelligence for screening colonoscopy: A modelling study. The Lancet Digital Health4(6), e436–e444 (2022)
work page 2022
-
[2]
V-JEPA 2: Self-Supervised Video Models Enable Understanding, Prediction and Planning
Assran, M., Bardes, A., Fan, D., Garrido, Q., Howes, R., Komeili, M., Muckley, M.J., Rizvi, A., et al.: V-JEPA 2: Self-supervised video models enable understand- ing, prediction and planning. arXiv preprint arXiv:2506.09985 (2025)
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[3]
Barbano, C.A., Dufumier, B., Tartaglione, E., Grangetto, M., Gori, P.: Unbiased supervised contrastive learning. In: ICLR (2023)
work page 2023
-
[4]
Batic, D., Holm, F., Özsoy, E., Czempiel, T., Navab, N.: EndoViT: Pretraining visiontransformersonalargecollectionofendoscopicimages.InternationalJournal of Computer Assisted Radiology and Surgery19(6), 1085–1091 (2024)
work page 2024
-
[5]
In: In- ternational Conference on Machine Learning
Bengio, Y., Louradour, J., Collobert, R., Weston, J.: Curriculum learning. In: In- ternational Conference on Machine Learning. pp. 41–48 (2009)
work page 2009
-
[6]
Scientific Data11(1), 539 (2024)
Biffi, C., Antonelli, G., Bernhofer, S., Hassan, C., Hirata, D., Iwatate, M., Maieron, A., Salvagnini, P., Cherubini, A.: REAL-Colon: A dataset for developing real-world AI applications in colonoscopy. Scientific Data11(1), 539 (2024)
work page 2024
-
[7]
In: International Conference on Machine Learning
Chen, T., Kornblith, S., Norouzi, M., Hinton, G.: A simple framework for con- trastive learning of visual representations. In: International Conference on Machine Learning. pp. 1597–1607 (2020)
work page 2020
-
[8]
In: Association for Compu- tational Linguistics
Devlin, J., Chang, M.W., Lee, K., Toutanova, K.: BERT: Pre-training of deep bidirectional transformers for language understanding. In: Association for Compu- tational Linguistics. pp. 4171–4186 (2019)
work page 2019
-
[9]
In: International Conference on Learning Representations (2021)
Dosovitskiy, A., Beyer, L., Kolesnikov, A., Weissenborn, D., Zhai, X., Unterthiner, T., Dehghani, M., Minderer, M., Heigold, G., Gelly, S., Uszkoreit, J., Houlsby, N.: An image is worth 16x16 words: Transformers for image recognition at scale. In: International Conference on Learning Representations (2021)
work page 2021
-
[10]
Dufumier, B., Barbano, C.A., Louiset, R., Duchesnay, E., Gori, P.: Integrating Prior Knowledge in Contrastive Learning with Kernel. In: ICML (2023)
work page 2023
-
[11]
Gastrointestinal Endoscopy93(1), 77–85 (2021) 10 L
Hassan, C., Spadaccini, M., Iannone, A., Maselli, R., Jovani, M., Chandrasekar, V.T., Antonelli, G., Yu, H., Areia, M., et al.: Performance of artificial intelligence in colonoscopy for adenoma and polyp detection: A systematic review and meta- analysis. Gastrointestinal Endoscopy93(1), 77–85 (2021) 10 L. Parolari et al
work page 2021
-
[12]
In: IEEE/CVF Conference on Computer Vision and Pattern Recognition
He, K., Fan, H., Wu, Y., Xie, S., Girshick, R.: Momentum contrast for unsupervised visual representation learning. In: IEEE/CVF Conference on Computer Vision and Pattern Recognition. pp. 9729–9738 (2020)
work page 2020
-
[13]
In: Conference on Computer Vision and Pattern Recognition
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: Conference on Computer Vision and Pattern Recognition. pp. 770–778 (2016)
work page 2016
-
[14]
Gaussian Error Linear Units (GELUs)
Hendrycks, D., Gimpel, K.: Gaussian error linear units. In: arXiv preprint arXiv:1606.08415 (2016)
work page internal anchor Pith review Pith/arXiv arXiv 2016
-
[15]
Intrator, Y., Aizenberg, N., Livne, A., Rivlin, E., Goldenberg, R.: Self-supervised polypre-identificationincolonoscopy.In:MedicalImageComputingandComputer Assisted Intervention. pp. 590–600 (2023)
work page 2023
-
[16]
Medical Image Analysis108, 103873 (2026)
Jaspers, T.J.M., de Jong, R.L.P.D., Li, Y., Kusters, C.H.J., Bakker, F.H.A., van Jaarsveld, R.C., Kuiper, G.M., van Hillegersberg, R., Ruurda, J.P., Brinkman, W.M., Pluim, J.P.W., et al.: Scaling up self-supervised learning for improved sur- gical foundation models. Medical Image Analysis108, 103873 (2026)
work page 2026
-
[17]
International Journal of Automa- tion and Computing19(6), 531–549 (2022)
Ji, G.P., Xiao, G., Chou, Y.C., Fan, D.P., Zhao, K., Chen, G., Gool, L.V.: Video polyp segmentation: A deep learning perspective. International Journal of Automa- tion and Computing19(6), 531–549 (2022)
work page 2022
-
[18]
In: Advances in Neural Information Processing Systems (2020)
Khosla, P., Teterwak, P., Wang, C., Sarna, A., Tian, Y., Isola, P., Maschinot, A., Liu, C., Krishnan, D.: Supervised contrastive learning. In: Advances in Neural Information Processing Systems (2020)
work page 2020
-
[19]
PLoS One16(8), e0255809 (2021)
Li, K., Fathan, M.I., Patel, K., Zhang, T., Zhong, C., Bansal, A., Rastogi, A., Wang, J.S., Wang, G.: Colonoscopy polyp detection and classification: Dataset creation and comparative evaluations. PLoS One16(8), e0255809 (2021)
work page 2021
-
[20]
In: International Conference on Learning Representations (2019)
Loshchilov, I., Hutter, F.: Decoupled weight decay regularization. In: International Conference on Learning Representations (2019)
work page 2019
-
[21]
In: Conference on Computer Vision and Pattern Recognition
Miech, A., Alayrac, J.B., Smaira, L., Laptev, I., Sivic, J., Zisserman, A.: End- to-end learning of visual representations from uncurated instructional videos. In: Conference on Computer Vision and Pattern Recognition. pp. 9876–9886 (2020)
work page 2020
-
[22]
Gastrointestinal Endoscopy93(4), 960–967 (2021)
Misawa, M., ei Kudo, S., Mori, Y., Hotta, K., Ohtsuka, K., Matsuda, T., Saito, S., Kudo, T., Baba, T., Ishida, F., et al.: Development of a computer-aided detection system for colonoscopy and a publicly accessible large colonoscopy video database. Gastrointestinal Endoscopy93(4), 960–967 (2021)
work page 2021
-
[23]
Neuro- computing423, 721–734 (2021)
Nogueira-Rodríguez, A., Domínguez-Carbajales, R., López-Fernández, H., Águeda Iglesias, Cubiella, J., Fdez-Riverola, F., Reboiro-Jato, M., Glez-Peña, D.: Deep neural networks approaches for detecting and classifying colorectal polyps. Neuro- computing423, 721–734 (2021)
work page 2021
-
[24]
Transactions on Machine Learning Research2024(2024)
Oquab, M., Darcet, T., Moutakanni, T., Vo, H.V., Szafraniec, M., Khalidov, V., Fernandez, P., Haziza, D., Massa, F., El-Nouby, A., Assran, M., Ballas, N., Galuba, W., Howes, R., Huang, P.Y., et al.: DINOv2: Learning robust visual features with- out supervision. Transactions on Machine Learning Research2024(2024)
work page 2024
-
[25]
In: Medical Image Computing and Computer Assisted Intervention (2025)
Parolari, L., Cherubini, A., Ballan, L., Biffi, C.: Temporally-aware supervised con- trastive learning for polyp counting in colonoscopy. In: Medical Image Computing and Computer Assisted Intervention (2025)
work page 2025
-
[26]
In: IEEE International Symposium on Biomedical Imaging
Parolari, L., Cherubini, A., Ballan, L., Biffi, C.: Towards polyp counting in full- procedure colonoscopy videos. In: IEEE International Symposium on Biomedical Imaging. pp. 1–5 (2025)
work page 2025
-
[27]
Sekiguchi, M., Mizuguchi, Y., Kawagoe, R., Saito, Y.: Artificial intelligence and its impact on the quality of endoscopy reports. Digestive Endoscopy38(1), e70057 (2026) Contrastive Learning under Noisy Temporal Self-Supervision in Colonoscopy 11
work page 2026
-
[28]
Siméoni, O., Vo, H.V., Seitzer, M., Baldassarre, F., Oquab, M., Jose, C., Khalidov, V., Szafraniec, M., Yi, S.E., Ramamonjisoa, M., Massa, F., Haziza, D., Wehrstedt, L., Wang, J., et al.: Dinov3. arXiv preprint arXiv:2508.10104 (2025)
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[29]
Scientific Data12(1), 918 (2025)
Song, Y., Du, S., Wang, R., Liu, F., Lin, X., Chen, J., Li, Z., Li, Z., Yang, L., Zhang, Z., Yan, H., Zhang, Q., Qian, D., Li, X.: Polyp-Size: A precise endoscopic dataset for AI-driven polyp sizing. Scientific Data12(1), 918 (2025)
work page 2025
-
[30]
Advances in Neural Information Processing Systems30(2017)
Vaswani, A., Shazeer, N., Parmar, N., Uszkoreit, J., Jones, L., Gomez,A.N.,Łukasz Kaiser, Polosukhin, I.: Attention is all you need. Advances in Neural Information Processing Systems30(2017)
work page 2017
-
[31]
In: Medical Image Computing and Computer Assisted Intervention
Wang, Z., Liu, C., Zhang, S., Dou, Q.: Foundation model for endoscopy video analysis via large-scale self-supervised pre-train. In: Medical Image Computing and Computer Assisted Intervention. pp. 101–111 (2023)
work page 2023
-
[32]
IEEE Journal of Biomedical and Health Informatics29(5), 3526–3536 (2025)
Wang, Z., Liu, C., Zhu, L., Wang, T., Zhang, S., Dou, Q.: Improving foundation model for endoscopy video analysis via representation learning on long sequences. IEEE Journal of Biomedical and Health Informatics29(5), 3526–3536 (2025)
work page 2025
-
[33]
In: IEEE International Conference on Acoustics, Speech and Signal Processing
Xiang, S., Liu, C., Ruan, J., Cai, S., Du, S., Qian, D.: VT-ReID: Learning discrimi- native visual-text representation for polyp re-identification. In: IEEE International Conference on Acoustics, Speech and Signal Processing. pp. 3170–3174 (2024)
work page 2024
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.