Recognition: no theorem link
Vine Copulas for Analyzing Multivariate Conditional Dependencies in Electronic Health Records Data
Pith reviewed 2026-05-10 18:20 UTC · model grok-4.3
The pith
Vine copulas decompose mixed electronic health records into tree structures that rank variables by conditional dependence and identify central ones for outcome prediction.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
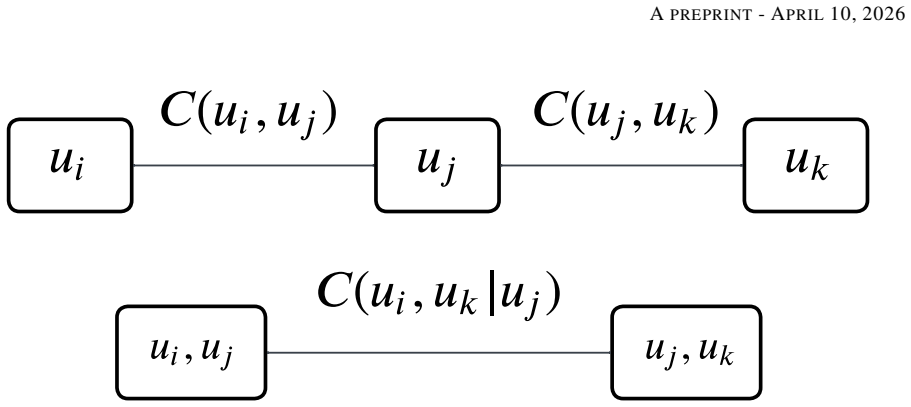
Vine copulas synthesize a multivariate distribution from many bivariate copulas organized into tree structures that represent conditional dependencies at successive hierarchical levels. The trees are then used to rank variables by conditional dependence and to extract subsets of central variables that exhibit local dependence. When applied to electronic health records, this decomposition identifies conditional links between co-morbid conditions and characterizes distinct patient cohorts, supplying data-driven explanations, visualizations, and a reduced variable set for prognostication.
What carries the argument
Vine copula tree structures that decompose the joint distribution into a sequence of bivariate conditional copulas arranged hierarchically.
If this is right
- Variables can be ordered by the strength of their conditional dependence on a chosen health outcome.
- Small sets of central variables with local dependence can be extracted to reduce the dimensionality of the analysis.
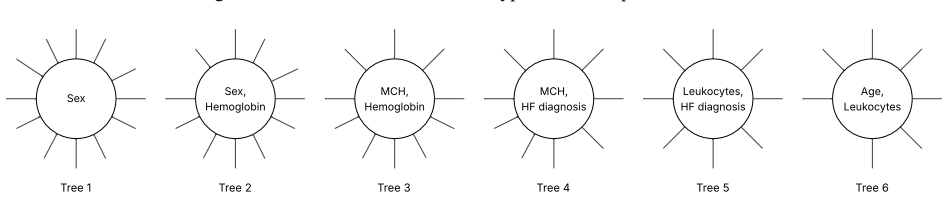
- Conditional dependence between co-morbid conditions becomes directly visible in the tree layout.
- The same trees can be applied to separate patient cohorts and tested for consistency across different record collections.
- The approach supplies both visual summaries and a principled basis for selecting variables before prognostic modeling.
Where Pith is reading between the lines
- The tree-based ranking could be inserted as a preprocessing step inside existing machine-learning pipelines that currently rely on correlation or mutual-information filters.
- Because the construction works with mixed data types, the same workflow might transfer to other domains that combine continuous sensor readings with ordinal or categorical labels.
- Longitudinal EHR streams could be re-estimated periodically to check whether the dependence trees remain stable or shift with changes in clinical practice.
Load-bearing premise
That vine copula trees can faithfully recover the conditional dependencies present in mixed-type electronic health records data.
What would settle it
On a public EHR dataset with documented strong dependencies, such as between diabetes diagnosis and repeated blood-glucose measurements, the vine copula trees either miss those links or yield no improvement in variable selection or cohort separation compared with simpler Gaussian-based methods.
Figures





read the original abstract
Electronic health records (EHR) store hundreds of demographic and laboratory variables from large patient populations. Traditional statistical methods have limited capacity in processing mixed-type data (continuous, ordinal) and capturing non-linear relationships in large multivariate data when oversimplified assumptions are made about the distribution (e.g., Gaussian) of disparate variables in EHR data. This paper addresses the limitations mentioned above by repurposing the vine copula method, which is primarily used to synthesize a multivariate distribution from many bivariate cumulative distribution functions (copulas). Vine copulas produce tree structures that represent bivariate conditional dependencies at varying hierarchical levels, decomposing a multivariate distribution. The tree structure is used to rank variables by conditional dependence and to identify a subset of central variables with local dependence, thus simplifying probabilistic mining of high-dimensional EHR data. The proposed application of vine copulas is used to identify conditional dependence between co-morbid conditions and is validated for characterizing different cohorts of EHR patients. The contribution of this paper is a novel approach to probabilistic mining and exploration of healthcare data that provides data-driven explanations, visualization, and variable selection to prognosticate a healthcare outcome. The source code is shared publicly.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes repurposing vine copula methods to model multivariate conditional dependencies in high-dimensional electronic health records (EHR) data containing mixed continuous and ordinal variables. It claims that the resulting vine tree structures can rank variables by their conditional dependence, identify central variables with local dependence, and thereby simplify probabilistic mining, visualization, and variable selection for characterizing patient cohorts and prognosticating healthcare outcomes. The approach is positioned as overcoming limitations of Gaussian-assuming methods, with public source code provided for reproducibility.
Significance. If the empirical validation holds, this work could offer a valuable tool for exploratory analysis of EHR data by providing interpretable dependency structures without strong distributional assumptions. The public release of source code is a strength that supports reproducibility and further application in healthcare data mining.
major comments (3)
- The manuscript claims that the proposed application 'is validated for characterizing different cohorts of EHR patients' (abstract) but supplies no quantitative results, performance metrics, error analysis, cohort definitions, dataset details, or comparisons to baselines, leaving the central claims of data-driven explanations and prognostication without demonstrated empirical support.
- In high-dimensional mixed EHR data, vine tree structure estimation proceeds via sequential pair-copula selection (typically maximum spanning tree on empirical Kendall's tau); the paper does not report bootstrap stability or reproducibility of the selected central nodes and hierarchy, which is load-bearing for the variable ranking and subset identification claims.
- The method applies standard vine copula decomposition to mixed continuous/ordinal variables but does not detail handling of mixed-pair copulas (e.g., via latent Gaussian transformations or adjusted empirical CDFs for ordinal margins), which directly affects the accuracy of conditional dependence capture asserted in the abstract.
minor comments (1)
- The abstract's contribution paragraph largely restates earlier sentences; consider tightening to highlight the specific novelty in the EHR application.
Simulated Author's Rebuttal
We thank the referee for the thoughtful and constructive report. The comments highlight important aspects of empirical validation, reproducibility, and technical detail that we address point by point below. We have prepared a revised manuscript that incorporates additional material to strengthen these areas.
read point-by-point responses
-
Referee: The manuscript claims that the proposed application 'is validated for characterizing different cohorts of EHR patients' (abstract) but supplies no quantitative results, performance metrics, error analysis, cohort definitions, dataset details, or comparisons to baselines, leaving the central claims of data-driven explanations and prognostication without demonstrated empirical support.
Authors: We agree that the current manuscript relies primarily on illustrative case studies rather than formal quantitative validation. In the revision we will expand the empirical section to include explicit cohort definitions, dataset characteristics (size, variable types, missingness), quantitative metrics (e.g., out-of-sample log-likelihood, variable-selection stability, and prognostic accuracy), and direct comparisons against Gaussian copula and mutual-information baselines. Error analysis will be added via cross-validation. revision: yes
-
Referee: In high-dimensional mixed EHR data, vine tree structure estimation proceeds via sequential pair-copula selection (typically maximum spanning tree on empirical Kendall's tau); the paper does not report bootstrap stability or reproducibility of the selected central nodes and hierarchy, which is load-bearing for the variable ranking and subset identification claims.
Authors: The referee correctly identifies a missing robustness check. We will add a bootstrap analysis (B = 500 resamples) that reports the selection frequency of the top-ranked central nodes and the stability of the first two tree levels. These results will be summarized in a new table and discussed in the context of variable-ranking reliability. revision: yes
-
Referee: The method applies standard vine copula decomposition to mixed continuous/ordinal variables but does not detail handling of mixed-pair copulas (e.g., via latent Gaussian transformations or adjusted empirical CDFs for ordinal margins), which directly affects the accuracy of conditional dependence capture asserted in the abstract.
Authors: We acknowledge the need for explicit technical description. The revised manuscript will contain a new subsection under Methods that specifies the marginal transformation procedure: continuous variables use the empirical CDF, ordinal variables use the adjusted empirical CDF with continuity correction, and mixed pairs are modeled with appropriate copula families (e.g., Gaussian or Frank) after these transformations. Relevant references on mixed copula estimation will be cited. revision: yes
Circularity Check
No circularity: standard vine copula decomposition applied to external EHR data
full rationale
The paper repurposes the established vine copula framework (tree-structured decomposition of multivariate distributions into bivariate copulas) on external EHR datasets for dependency ranking and variable selection. No derivation step reduces by construction to fitted parameters renamed as predictions, self-definitional loops, or load-bearing self-citations. The central claims rest on applying known vine properties to new data without the result being equivalent to the inputs by definition. This is the common non-circular case of methodological application.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Vine copulas can effectively model mixed continuous and ordinal data in EHR while capturing non-linear conditional dependencies without Gaussian distributional assumptions.
Reference graph
Works this paper leans on
-
[1]
T. Bouezmarni, M. Doukali, A. Taamouti, Copula-based estimation of health inequality measures, Journal of the Royal Statistical Society Series A: Statistics in Society (2025) qnaf039doi:10.1093/jrsssa/qnaf039. URLhttps://doi.org/10.1093/jrsssa/qnaf039
-
[2]
Frank, On the simultaneous associativity of f (x, y) and x+ y-f (x, y)., Aequationes Math
M. Frank, On the simultaneous associativity of f (x, y) and x+ y-f (x, y)., Aequationes Math. 19 (1) (1979) 141–160
1979
-
[3]
J. E. Black, Prognostic predictive model to estimate the risk of multiple chronic diseases: Constructing copulas using electronic medical record data, Master’s thesis, The University of Western Ontario (2018)
2018
-
[4]
Czado, T
C. Czado, T. Nagler, Vine copula based modeling, Annual Review of Statistics and Its Application 9 (1) (2022) 453–477
2022
-
[5]
N. Z. Petrakos, E. E. Moodie, N. Savy, A framework for generating realistic synthetic tabular data in a randomized controlled trial setting, Statistics in Medicine 44 (18-19) (2025) e70227
2025
-
[6]
L. Xu, M. Skoularidou, A. Cuesta-Infante, K. Veeramachaneni, Modeling tabular data using conditional gan, Advances in neural information processing systems 32 (2019)
2019
-
[7]
D. S. Watson, K. Blesch, J. Kapar, M. N. Wright, Adversarial random forests for density estimation and genera- tive modeling, in: Int. Conf. on Artificial Intel. and Statistics, PMLR, 2023, pp. 5357–5375
2023
-
[8]
A. M. Chu, C. Y . Ip, B. S. Lam, M. K. So, Vine copula statistical disclosure control for mixed-type data, Com- putational Statistics & Data Analysis 176 (2022) 107561
2022
-
[9]
¨O. S ¸ahin, Probabilistic patient risk profiling with pair-copula constructions, arXiv preprint arXiv:2506.13731 (2025). 9 APREPRINT- APRIL10, 2026
-
[10]
Dissmann, E
J. Dissmann, E. C. Brechmann, C. Czado, D. Kurowicka, Selecting and estimating regular vine copulae and application to financial returns, Computational Statistics & Data Analysis 59 (2013) 52–69
2013
-
[11]
of Us Research Program Investigators, J
A. of Us Research Program Investigators, J. C. Denny, J. L. Rutter, D. B. Goldstein, A. Philippakis, J. W. Smoller, G. Jenkins, E. Dishman, The ”all of us” research program, New England Journal of Medicine 381 (2019) 668–
2019
-
[12]
doi:10.1056/NEJMsr1809937
-
[13]
P. L. Sankar, L. S. Parker, The precision medicine initiative’s all of us research program: an agenda for research on its ethical, legal, and social issues, Genetics in Medicine 19 (7) (2017) 743–750
2017
-
[14]
Griffiths, A
C. Griffiths, A. Brock, C. Rooney, The impact of introducing icd-10 on trends in mortality from circulatory diseases in england and wales, Health Statistics Quarterly (22) (2004) 14–20
2004
-
[15]
C. Luo, Y . Zhu, Z. Zhu, R. Li, G. Chen, Z. Wang, A machine learning-based risk stratification tool for in-hospital mortality of intensive care unit patients with heart failure, Journal of Translational Medicine 20 (1) (2022) 136. doi:10.1186/s12967-022-03340-8
- [16]
-
[17]
J. Zhu, L. Hong, S. Yuan, X. Xu, J. Wei, H. Yin, Association between glucocorticoid use and all-cause mor- tality in critically ill patients with heart failure: A cohort study based on the mimic-iii database, Frontiers in Pharmacology 14 (2023) 1118551. doi:10.3389/fphar.2023.1118551
-
[18]
A. A. Huang, S. Y . Huang, Dendrogram of transparent feature importance machine learning statistics to classify associations for heart failure: A reanalysis of a retrospective cohort study of the medical information mart for intensive care iii (mimic-iii) database, PLOS ONE 18 (7) (2023) e0288819. doi:10.1371/journal.pone.0288819
-
[19]
Y . Hou, S. B. Rabbani, L. Hong, N. Diawara, M. D. Samad, Causal explainability of machine learning in heart failure prediction from electronic health records, in: 2025 IEEE Int Conf. on Information Reuse and Integration and Data Science (IRI), 2025, pp. 128–134. doi:10.1109/IRI66576.2025.00030
-
[20]
J. N. Claassen, E. E. Koks, M. C. de Ruiter, P. J. Ward, W. S. J¨ager, Vinecopulas: an open-source python package for vine copula modelling, J. of Open Source Software 9 (101) (2024) 6728
2024
-
[21]
Mining electronic health records to investigate effectiveness of ensemble deep clustering, in preparation (2026)
2026
-
[22]
S. W. Siddiqui, T. Ashok, N. Patni, M. Fatima, A. Lamis, K. K. Anne, Anemia and heart failure: a narrative review, Cureus 14 (7) (2022)
2022
-
[23]
Kleber, N
M. Kleber, N. Kozhuharov, Z. Sabti, B. Glatz, R. Isenreich, D. Wussler, A. Nowak, R. Twerenbold, P. Badertscher, C. Puelacher, et al., Relative hypochromia and mortality in acute heart failure, International journal of cardiology 286 (2019) 104–110
2019
-
[24]
Z. Zhu, S. Zhou, Leukocyte count and the risk of adverse outcomes in patients with hfpef, BMC Cardiovascular Disorders 21 (1) (2021) 333
2021
-
[25]
L. J. Drop, Ionized calcium, the heart, and hemodynamic function, Anesthesia & Analgesia 64 (4) (1985) 432– 451
1985
-
[26]
S. S. Franklin, D. Levy, Aging, blood pressure, and heart failure: what are the connections?, Hypertension 58 (5) (2011) 760–762
2011
-
[27]
Davies, L
M. Davies, L. Færch, O. K. Jeppesen, A. Pakseresht, S. D. Pedersen, L. Perreault, J. Rosenstock, I. Shimomura, A. Viljoen, T. A. Wadden, et al., Semaglutide 2· 4 mg once a week in adults with overweight or obesity, and type 2 diabetes (step 2): a randomised, double-blind, double-dummy, placebo-controlled, phase 3 trial, The Lancet 397 (10278) (2021) 971–984
2021
-
[28]
Figorilli, F
M. Figorilli, F. Velluzzi, S. Redolfi, Obesity and sleep disorders: A bidirectional relationship, Nutrition, Metabolism and Cardiovascular Diseases (2025) 104014
2025
-
[29]
E. Jemt, M. Ekstr ¨om, U. Ekelund, Outcomes in emergency department patients with dyspnea versus chest pain: a retrospective consecutive cohort study, Emergency Medicine International 2022 (1) (2022) 4031684
2022
-
[30]
E. M. Byrne, A. Timmerman, N. R. Wray, E. Agerbo, Sleep disorders and risk of incident depression: a popula- tion case–control study, Twin Research and Human Genetics 22 (3) (2019) 140–146. 10
2019
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.