Recognition: unknown
PanGuide3D: Cohort-Robust Pancreas Tumor Segmentation via Probabilistic Pancreas Conditioning and a Transformer Bottleneck
Pith reviewed 2026-05-09 22:09 UTC · model grok-4.3
The pith
PanGuide3D improves cross-cohort pancreas tumor segmentation by conditioning the tumor decoder on probabilistic pancreas maps and adding a transformer bottleneck.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
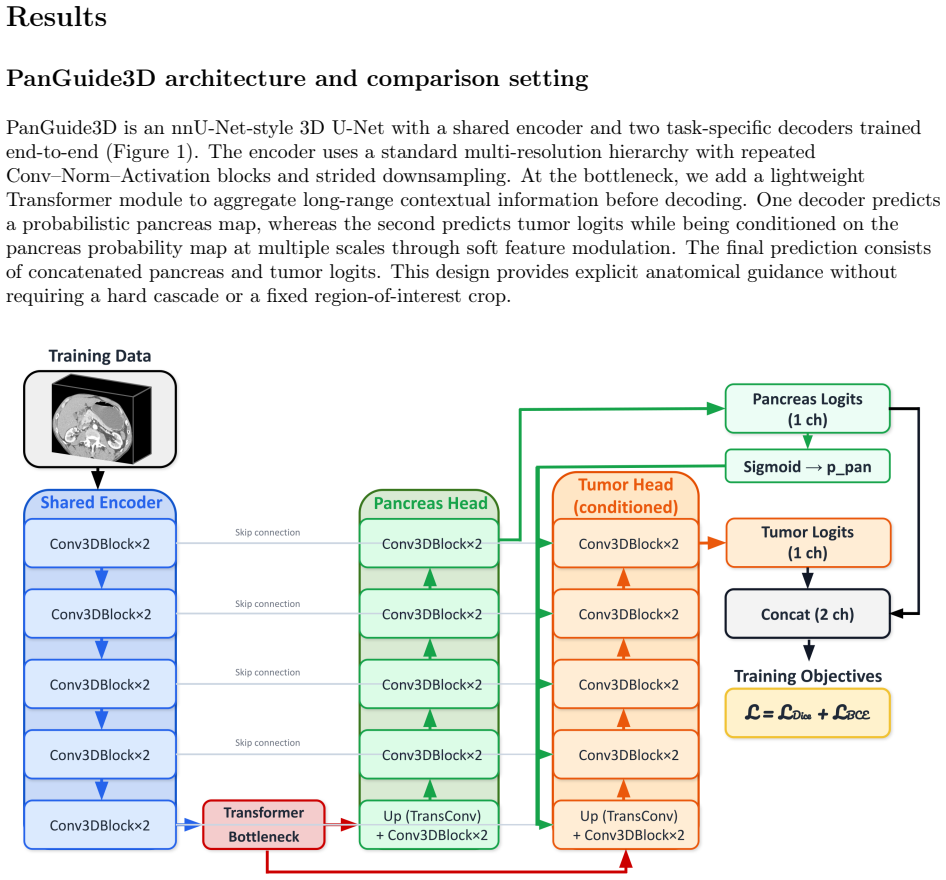
PanGuide3D consists of a shared 3D encoder, a pancreas decoder that outputs a probabilistic pancreas map, and a tumor decoder whose features are gated by this map at multiple scales through differentiable soft gating; a lightweight transformer is placed in the U-Net bottleneck to capture long-range context. Trained on the PanTS cohort and tested both in-cohort and on the out-of-cohort MSD Task07 pancreas data under matched preprocessing and training settings, the model records the highest tumor segmentation scores and the clearest gains in generalization, especially for small tumors and tumors in difficult anatomical positions, while producing fewer anatomically implausible false positives.
What carries the argument
Probabilistic pancreas map produced by a dedicated decoder and used to condition the tumor decoder via multi-scale soft gating, together with a transformer bottleneck
If this is right
- Higher tumor segmentation accuracy on small lesions and tumors in challenging anatomical locations.
- Fewer anatomically implausible false positives in the output masks.
- Better preservation of performance when the model is applied to a new cohort after training on PanTS.
- Overall highest tumor metrics among the models compared under the same evaluation protocol.
Where Pith is reading between the lines
- The same soft-gating idea could be used to condition segmentation of other abdominal structures on their own probabilistic priors.
- The transformer bottleneck may supply the global context that purely convolutional paths lose when patient demographics or scanner settings change.
- If the conditioning holds up, similar anatomical guidance layers could be added to existing 3D networks for other tumor types without redesigning the entire encoder.
Load-bearing premise
The observed gains in cross-cohort tumor performance are caused by the probabilistic pancreas conditioning and transformer bottleneck rather than by differences in optimization, data augmentation, or other unstated implementation choices.
What would settle it
An experiment that trains PanGuide3D and a plain 3D U-Net on identical PanTS data with the same optimizer, augmentation, and hyperparameters and then finds no improvement, or a drop, in tumor Dice score and patient-level detection rate for PanGuide3D when both are tested on the MSD Task07 set.
Figures








read the original abstract
Pancreatic tumor segmentation in contrast-enhanced computed tomography (CT) is clinically important yet technically challenging: lesions are often small, heterogeneous, and easily confused with surrounding soft tissue, and models that perform well on one cohort frequently degrade under cohort shift. Our goal is to improve cross-cohort generalization while keeping the model architecture simple, efficient, and practical for 3D CT segmentation. We introduce PanGuide3D, a cohort-robust architecture with a shared 3D encoder, a pancreas decoder that predicts a probabilistic pancreas map, and a tumor decoder that is explicitly conditioned on this pancreas probability at multiple scales via differentiable soft gating. To capture long-range context under distribution shift, we further add a lightweight Transformer bottleneck in the U-Net bottleneck representation. We evaluate cohort transfer by training on the PanTS (Pancreatic Tumor Segmentation) cohort and testing both in-cohort (PanTS) and out-of-cohort on MSD (Medical Segmentation Decathlon) Task07 Pancreas, using matched preprocessing and training protocols across strong baselines. We collect voxel-level segmentation metrics, patient-level tumor detection, subgroup analyses by tumor size and anatomical location, volume-conditioned performance analyses, and calibration measurements to assess reliability. Across the evaluated models, PanGuide3D achieves the best overall tumor performance and shows improved cross-cohort generalization, particularly for small tumors and challenging anatomical locations, while reducing anatomically implausible false positives. These findings support probabilistic anatomical conditioning as a practical strategy for improving cross-cohort robustness in an end-to-end model and suggest potential utility for contouring support, treatment planning, and multi-institutional studies.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces PanGuide3D, a 3D segmentation network with a shared encoder, a dedicated pancreas decoder that outputs a probabilistic pancreas map, and a tumor decoder conditioned on this map at multiple scales through differentiable soft gating. A lightweight transformer is inserted at the U-Net bottleneck to capture long-range context. The model is trained on the PanTS cohort and evaluated both in-cohort and on the held-out MSD Task07 Pancreas cohort using matched preprocessing and protocols. The central claim is that PanGuide3D achieves the highest tumor Dice, improved patient-level detection, better subgroup performance on small tumors and difficult anatomical sites, and fewer anatomically implausible false positives, with the gains attributed to the probabilistic conditioning and transformer bottleneck for enhanced cross-cohort robustness.
Significance. If the performance gains are causally linked to the proposed conditioning and bottleneck rather than implementation details, the work supplies a relatively simple, end-to-end architectural recipe for improving generalization under cohort shift in 3D medical segmentation. This addresses a practical barrier to deploying models across institutions and could support more reliable contouring tools, with the added benefit of explicit anatomical guidance that may reduce clinically unacceptable errors.
major comments (2)
- [Experimental Evaluation / Results] The evaluation section states that matched preprocessing and training protocols were used across baselines, yet no component-wise ablations (full PanGuide3D vs. baseline U-Net vs. conditioning-only vs. transformer-only) are reported under identical optimizer, augmentation, and random-seed conditions. Because the central claim attributes the cross-cohort tumor Dice, detection, and false-positive improvements on MSD Task07 specifically to the soft-gating pancreas conditioning and transformer bottleneck, the absence of these isolations leaves open the possibility that observed gains arise from unstated hyperparameter or data-handling differences (see skeptic note on unisolated factors).
- [Results] Subgroup analyses by tumor size and anatomical location are described, but the manuscript does not provide the corresponding numerical tables or statistical tests (e.g., paired Wilcoxon or bootstrap confidence intervals) that would confirm the claimed superiority for small tumors. Without these, the assertion that gains are “particularly” pronounced in challenging subgroups remains difficult to verify quantitatively.
minor comments (2)
- [Methods] Clarify the exact formulation of the soft-gating operation (scale factors, temperature, and how gradients flow through the probabilistic map) in the methods section; the current description is high-level and could be made more reproducible.
- [Results] Add a short paragraph on calibration metrics (mentioned in the abstract) with explicit Brier or ECE values; this would strengthen the reliability claims.
Simulated Author's Rebuttal
We thank the referee for the constructive feedback on our experimental design and results presentation. The comments highlight important areas for strengthening the validation of our claims regarding the contributions of probabilistic pancreas conditioning and the transformer bottleneck to cross-cohort robustness. We address each major comment below and have revised the manuscript to incorporate the requested analyses and supporting evidence.
read point-by-point responses
-
Referee: [Experimental Evaluation / Results] The evaluation section states that matched preprocessing and training protocols were used across baselines, yet no component-wise ablations (full PanGuide3D vs. baseline U-Net vs. conditioning-only vs. transformer-only) are reported under identical optimizer, augmentation, and random-seed conditions. Because the central claim attributes the cross-cohort tumor Dice, detection, and false-positive improvements on MSD Task07 specifically to the soft-gating pancreas conditioning and transformer bottleneck, the absence of these isolations leaves open the possibility that observed gains arise from unstated hyperparameter or data-handling differences (see skeptic note on unisolated factors).
Authors: We agree that component-wise ablations under fully controlled conditions are required to isolate the effects of the proposed modules and rule out confounding factors. In the revised manuscript we have added these ablations, retraining the baseline U-Net, conditioning-only variant, transformer-only variant, and full PanGuide3D under identical optimizer, augmentation, and random-seed settings. The new results (added as Table 4 in the Experimental Evaluation section) show that each component contributes measurable gains in cross-cohort tumor Dice and detection on MSD Task07, with the full model achieving the largest improvement. This directly addresses the concern about unisolated factors. revision: yes
-
Referee: [Results] Subgroup analyses by tumor size and anatomical location are described, but the manuscript does not provide the corresponding numerical tables or statistical tests (e.g., paired Wilcoxon or bootstrap confidence intervals) that would confirm the claimed superiority for small tumors. Without these, the assertion that gains are “particularly” pronounced in challenging subgroups remains difficult to verify quantitatively.
Authors: We acknowledge that the original manuscript described subgroup trends without accompanying numerical tables or formal statistical tests. We have now added comprehensive tables reporting Dice scores, sensitivity, and false-positive rates stratified by tumor size (small < 2 cm, medium 2–4 cm, large > 4 cm) and by anatomical location. We further include paired Wilcoxon signed-rank tests and bootstrap confidence intervals (1,000 resamples) comparing PanGuide3D against the strongest baseline. These additions appear in the revised Results section and confirm statistically significant improvements (p < 0.05) that are indeed largest for small tumors and difficult anatomical sites. revision: yes
Circularity Check
No circularity: purely empirical evaluation on held-out cohorts
full rationale
The paper introduces an architecture (shared 3D encoder, probabilistic pancreas decoder with soft gating, tumor decoder conditioned on pancreas probability, plus transformer bottleneck) and reports direct empirical results: tumor Dice, detection, and false-positive metrics on MSD Task07 after training on PanTS, with matched preprocessing across baselines. No equations, derivations, or predictions are presented that reduce by construction to fitted parameters, self-citations, or ansatzes within the paper. All claims rest on observed performance differences under cohort shift, which are independent measurements rather than tautological re-statements of inputs. This matches the default expectation of a non-circular empirical study.
Axiom & Free-Parameter Ledger
free parameters (2)
- Network weights
- Soft-gating scale factors
axioms (2)
- domain assumption A probabilistic pancreas map supplies useful spatial guidance for tumor segmentation under cohort shift
- domain assumption A lightweight transformer bottleneck captures long-range context that improves robustness to distribution shift
Reference graph
Works this paper leans on
-
[1]
State-of-the-art and challenges in pancreatic ct segmentation: A systematic review of u-net and its variants.IEEE Access, 12:78726–78742, 2024
Chaohui Zhang, Anusha Achuthan, and Galib Muhammad Shahriar Himel. State-of-the-art and challenges in pancreatic ct segmentation: A systematic review of u-net and its variants.IEEE Access, 12:78726–78742, 2024
2024
-
[2]
A systematic review on leveraging artificial intelligence for pancreatic cancer diagnosis.Science, 20:100268, 2026
Sonia Suneja, Rajneesh Talwar, and Manvinder Sharma. A systematic review on leveraging artificial intelligence for pancreatic cancer diagnosis.Science, 20:100268, 2026
2026
-
[3]
Hajra Arshad, Felipe Lopez-Ramirez, Florent Tixier, Philippe Soyer, Satomi Kawamoto, Elliot K Fishman, and Linda C Chu. Radiomics in early detection of pancreatic ductal adenocarcinoma: a close look at its current status and challenges to clinical implementation.Canadian Association of Radiologists Journal, 77(1):107–118, 2026
2026
-
[4]
Early detection of pancreatic cancer on computed tomography: advancements with deep learning.Radiology Advances, 2(5):umaf028, 2025
Felipe Lopez-Ramirez, Emir A Syailendra, Florent Tixier, Satomi Kawamoto, Elliot K Fishman, and Linda C Chu. Early detection of pancreatic cancer on computed tomography: advancements with deep learning.Radiology Advances, 2(5):umaf028, 2025
2025
-
[5]
Advances on pancreas segmentation: a review.Multimedia Tools and Applications, 79(9):6799–6821, 2020
Xu Yao, Yuqing Song, and Zhe Liu. Advances on pancreas segmentation: a review.Multimedia Tools and Applications, 79(9):6799–6821, 2020
2020
-
[6]
Automatic segmentation of pancreas and pancreatic tumor: a review of a decade of research.IEEE Access, 11:108727–108745, 2023
Harshal Ghorpade et al. Automatic segmentation of pancreas and pancreatic tumor: a review of a decade of research.IEEE Access, 11:108727–108745, 2023
2023
-
[7]
Qiu et al
D. Qiu et al. A deep learning-based cascade algorithm for pancreatic tumor segmentation.Frontiers in Oncology, 14:1328146, 2024
2024
-
[8]
Ghorpade, S
H. Ghorpade, S. Kolhar, J. Jagtap, and J. Chakraborty. An optimized two stage U-Net approach for segmentation of pancreas and pancreatic tumor.MethodsX, 13:102995, 2024
2024
-
[9]
Mahmoudi et al
T. Mahmoudi et al. Segmentation of pancreatic ductal adenocarcinoma (PDAC) and surrounding vessels in CT images using deep convolutional neural networks and texture descriptors.Scientific Reports, 12:3092, 2022
2022
-
[10]
Jaeger, Simon A
Fabian Isensee, Paul F. Jaeger, Simon A. A. Kohl, Jens Petersen, and Klaus H. Maier-Hein. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation.Nature Methods, 18:203–211, 2021. 17
2021
-
[11]
U-Net: Convolutional networks for biomedical image segmentation
Olaf Ronneberger, Philipp Fischer, and Thomas Brox. U-Net: Convolutional networks for biomedical image segmentation. InMedical Image Computing and Computer-Assisted Intervention (MICCAI), pages 234–241. Springer, 2015
2015
-
[12]
Attention is all you need
Ashish Vaswani et al. Attention is all you need. InAdvances in Neural Information Processing Systems, volume 30, 2017
2017
-
[13]
Li et al
W. Li et al. PanTS: the pancreatic tumor segmentation dataset. InThirty-ninth Annual Conference on Neural Information Processing Systems Datasets and Benchmarks Track, 2025
2025
-
[14]
UNETR: Transformers for 3d medical image segmentation
Ali Hatamizadeh et al. UNETR: Transformers for 3d medical image segmentation. InProceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, pages 574–584, 2022
2022
-
[15]
Wu et al
C. Wu et al. Towards generalist foundation model for radiology by leveraging web-scale 2d & 3d medical data.Nature Communications, 16:7866, 2025
2025
-
[16]
The medical segmentation decathlon.Nature Communications, 13:4128, 2022
Michela Antonelli et al. The medical segmentation decathlon.Nature Communications, 13:4128, 2022. 18
2022
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.