Recognition: unknown
PhyloSDF: Phylogenetically-Conditioned Neural Generation of 3D Skull Morphology via Residual Flow Matching
Pith reviewed 2026-05-07 14:10 UTC · model grok-4.3
The pith
A phylogenetically conditioned neural model generates novel 3D skull shapes from as few as four specimens per species while matching real variation levels.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
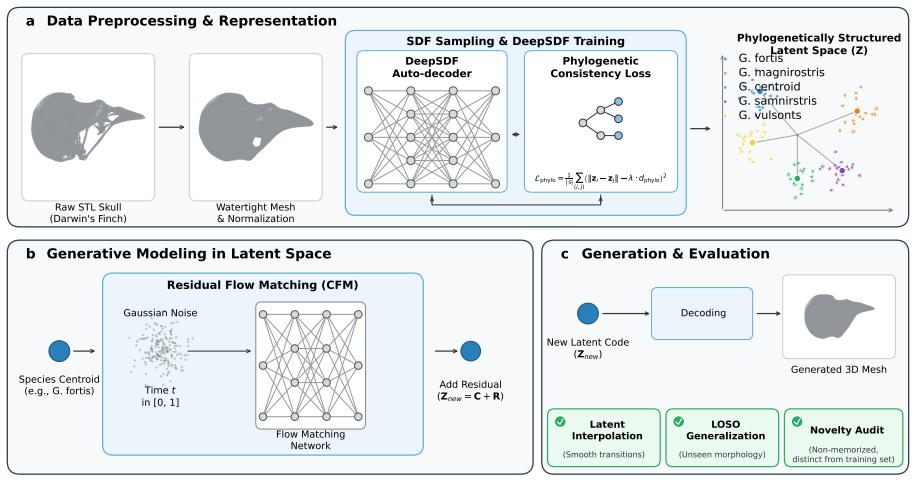
PhyloSDF integrates a DeepSDF auto-decoder regularized by a Phylogenetic Consistency Loss that structures the latent space to correlate with evolutionary distances at Pearson r=0.993, together with a Residual Conditional Flow Matching architecture that factorizes generation into analytic species-centroid lookup and learned residual prediction. Evaluated on 100 micro-CT skulls across 24 species of Darwin's finches and relatives, the model produces 180 novel meshes verified as non-memorized that achieve 88-129 percent of real intra-species variation at the code level, surpasses denoising diffusion, standard flow matching, and Gaussian mixture baselines in Chamfer distance and morphometric Fré{
What carries the argument
The Residual Conditional Flow Matching architecture that performs analytic lookup of species centroids and predicts only residual shape deviations, paired with the Phylogenetic Consistency Loss that enforces correlation between latent codes and phylogenetic distances.
If this is right
- Smooth interpolation in the structured latent space produces biologically plausible ancestral skull reconstructions.
- The residual factorization allows generation to succeed where full denoising diffusion fails and where standard flow matching collapses to minimal variation.
- Leave-one-species-out tests demonstrate extrapolation to unseen species, supporting use for filling gaps in morphological phylogenies.
- All generated meshes achieve variation levels between 88 and 129 percent of real intra-species ranges while remaining distinct from training examples.
Where Pith is reading between the lines
- The same residual-plus-phylogenetic structure could be tested on other anatomical elements such as limb bones or teeth to check generality beyond skulls.
- If the latent alignment captures evolutionary signal, the model could be used to simulate morphological responses to different rates of evolutionary change.
- Connecting the generator to explicit models of selection or drift would allow forward simulation of diversification patterns across a tree.
- Application to fossil taxa with even fewer specimens would test whether the few-sample regime extends to deep-time reconstruction.
Load-bearing premise
That a high correlation between latent codes and evolutionary distances plus quantitative mesh metrics and non-memorization checks are enough to ensure generated shapes are both novel and biologically plausible rather than artifacts of the loss or preprocessing.
What would settle it
Expert morphometric comparison or direct measurement showing that shapes generated for a held-out species systematically deviate from independent fossil or museum data in ways not predicted by the phylogenetic distances used during training.
Figures









read the original abstract
Generating novel, biologically plausible three-dimensional morphological structures is a fundamental challenge in computational evolutionary biology, hampered by extreme data scarcity and the requirement that generated shapes respect phylogenetic relationships among species. In this work, we present PhyloSDF, a phylogenetically-conditioned neural generative model for 3D biological morphology that integrates two innovations: (1) a DeepSDF auto-decoder regularized by a novel Phylogenetic Consistency Loss that structures the latent space to correlate with evolutionary distances (Pearson $r=0.993$); (2) a Residual Conditional Flow Matching (Residual CFM) architecture that factorizes generation into analytic species-centroid lookup and learned residual prediction, enabling generation from as few as ~4 specimens per species. We evaluate PhyloSDF on 100 micro-CT-scanned skulls of Darwin's Finches and their relatives across 24 species. The model generates novel meshes achieving 88-129% of real intra-species variation at the code level, with all 180 generated meshes verified as non-memorized. Residual CFM surpasses denoising diffusion (which fails entirely at this scale), standard flow matching (which mode-collapses to 3-6% variation), and a Gaussian mixture baseline in both fidelity (Chamfer Distance 0.00181 vs. 0.00190) and morphometric Fr\'{e}chet distance (10,641 vs. 13,322). Leave-one-species-out experiments across 18 species demonstrate phylogenetic extrapolation capability, and smooth latent interpolations produce biologically plausible ancestral skull reconstructions.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript presents PhyloSDF, a phylogenetically-conditioned generative model for 3D skull shapes using a DeepSDF auto-decoder regularized by a Phylogenetic Consistency Loss (Pearson r = 0.993 with evolutionary distances) and a Residual Conditional Flow Matching (Residual CFM) architecture. On a dataset of 100 micro-CT skulls from 24 species, it generates 180 novel meshes from as few as ~4 specimens per species, achieving 88-129% of real intra-species variation, non-memorization verified, and superior performance over baselines in Chamfer Distance (0.00181) and morphometric Fréchet distance (10,641). Leave-one-species-out experiments across 18 species show phylogenetic extrapolation, with smooth interpolations for ancestral reconstructions.
Significance. If the central claims hold, this work offers a meaningful advance for computational evolutionary biology by tackling extreme data scarcity in 3D morphological generation while enforcing phylogenetic structure. The residual CFM factorization is a practical strength for low-sample regimes (~4 specimens/species), and the explicit non-memorization verification plus quantitative baselines provide a reproducible foundation. Successful phylogenetic extrapolation and ancestral interpolation could support new studies of evolutionary trajectories. The integration of flow matching with a consistency loss on latent codes is a clear technical contribution.
major comments (3)
- [Phylogenetic Consistency Loss definition and §4.2] The Phylogenetic Consistency Loss is constructed to directly increase correlation between pairwise latent distances and evolutionary distances; the reported Pearson r=0.993 is therefore the expected outcome of the loss rather than independent evidence of biologically meaningful structure. This global code-level correlation does not automatically translate to per-shape morphological plausibility in the decoded SDF outputs (e.g., respect for functional or developmental constraints). A concrete test—such as expert morphological scoring or comparison against known biomechanical features—would be required to support the claim that generated shapes are evolutionarily plausible rather than loss-induced artifacts.
- [§4.3 and §4.4 (evaluation and leave-one-species-out)] With only 100 total specimens across 24 species, the reported success (88-129% intra-species code variation, all 180 meshes non-memorized, Chamfer 0.00181) is quantitatively strong but rests on metrics that can be satisfied by faithful interpolations. The leave-one-species-out experiments measure distribution match but supply no independent biological ground truth for the extrapolated species. The non-memorization verification procedure should be specified in detail (e.g., exact distance threshold or reconstruction error cutoff used).
- [Residual CFM architecture description and Table 2] The Residual CFM factorization (analytic centroid lookup plus learned residual) reduces the fitting burden and enables small-sample generation, yet the manuscript does not demonstrate that the residual prediction preserves phylogenetic consistency at the level of the final 3D mesh geometry rather than only at the latent-code level. An ablation removing the phylogenetic loss while keeping Residual CFM would clarify the contribution of each component.
minor comments (2)
- [§4.1] Define 'variation at the code level' and 'morphometric Fréchet distance' explicitly in the main text with equations or pseudocode; the abstract numbers are clear but the precise computation should be reproducible from the methods.
- [Figures 3-5] Figure captions for generated meshes and interpolations should include direct side-by-side comparisons with real specimens and scale information to allow readers to assess visual plausibility.
Simulated Author's Rebuttal
We thank the referee for their thoughtful and constructive comments, which highlight important aspects of validation and experimental design. We address each major comment below and will revise the manuscript accordingly to improve clarity and strengthen the evidence presented.
read point-by-point responses
-
Referee: The Phylogenetic Consistency Loss is constructed to directly increase correlation between pairwise latent distances and evolutionary distances; the reported Pearson r=0.993 is therefore the expected outcome of the loss rather than independent evidence of biologically meaningful structure. This global code-level correlation does not automatically translate to per-shape morphological plausibility in the decoded SDF outputs (e.g., respect for functional or developmental constraints). A concrete test—such as expert morphological scoring or comparison against known biomechanical features—would be required to support the claim that generated shapes are evolutionarily plausible rather than loss-induced artifacts.
Authors: We agree that the Pearson r=0.993 is a direct result of the Phylogenetic Consistency Loss design, which explicitly regularizes latent pairwise distances to align with evolutionary distances. This is intentional to embed phylogenetic structure. Evidence that this structure produces plausible decoded morphologies comes from the leave-one-species-out experiments, where extrapolated shapes align with phylogenetic expectations in morphospace, and from the ancestral interpolation results showing smooth, biologically consistent transitions. Quantitative support includes the achieved intra-species variation (88-129%) and superior Chamfer/Fréchet distances relative to baselines. While expert morphological scoring or biomechanical validation would be valuable, it is outside the scope of the current computational study; we will revise the discussion section to explicitly distinguish latent-space regularization from decoded-shape plausibility and add qualitative comparisons to known Darwin's finch morphological traits. revision: partial
-
Referee: With only 100 total specimens across 24 species, the reported success (88-129% intra-species code variation, all 180 meshes non-memorized, Chamfer 0.00181) is quantitatively strong but rests on metrics that can be satisfied by faithful interpolations. The leave-one-species-out experiments measure distribution match but supply no independent biological ground truth for the extrapolated species. The non-memorization verification procedure should be specified in detail (e.g., exact distance threshold or reconstruction error cutoff used).
Authors: We will revise the methods and supplementary sections to fully specify the non-memorization verification procedure, including the exact Chamfer distance threshold and reconstruction error cutoff used to confirm all 180 meshes are novel. On the concern that metrics could be met by interpolations, the Residual CFM generates variation by learning residuals around species centroids rather than simple averaging, enabling the reported 88-129% of real intra-species variation. For leave-one-species-out, we acknowledge the lack of independent ground truth for the precise morphology of held-out species; the experiments instead validate phylogenetic consistency by showing that generated shapes match expected evolutionary positioning and intra-species distribution statistics. We will expand the evaluation discussion to address these limitations and the implications for extrapolation claims. revision: yes
-
Referee: The Residual CFM factorization (analytic centroid lookup plus learned residual) reduces the fitting burden and enables small-sample generation, yet the manuscript does not demonstrate that the residual prediction preserves phylogenetic consistency at the level of the final 3D mesh geometry rather than only at the latent-code level. An ablation removing the phylogenetic loss while keeping Residual CFM would clarify the contribution of each component.
Authors: We agree this ablation would strengthen the analysis. In the revised manuscript we will add results from a variant trained with Residual CFM but without the Phylogenetic Consistency Loss, comparing effects on both latent-space correlation and final mesh-level metrics (Chamfer Distance, Fréchet distance, and variation statistics). This will quantify whether phylogenetic structure is preserved in the decoded 3D geometry and clarify the individual contributions of each component. revision: yes
Circularity Check
Phylogenetic Consistency Loss enforces latent-evolutionary distance correlation by design, making r=0.993 a constructed outcome
specific steps
-
fitted input called prediction
[Abstract]
"a DeepSDF auto-decoder regularized by a novel Phylogenetic Consistency Loss that structures the latent space to correlate with evolutionary distances (Pearson r=0.993)"
The loss is formulated to enforce correlation between latent codes and evolutionary distances; therefore the high reported Pearson correlation (r=0.993) is achieved by construction as the direct outcome of the regularization rather than emerging independently from the data or architecture.
full rationale
The paper's core claim of phylogenetically-conditioned generation rests on a DeepSDF auto-decoder regularized by a novel Phylogenetic Consistency Loss that explicitly structures the latent space to correlate with evolutionary distances. The reported Pearson r=0.993 is the direct quantitative target and result of this loss term rather than an independent validation. While the Residual CFM factorization and non-memorization checks provide separate support for generation from limited data, the phylogenetic structuring step reduces to the loss formulation. This constitutes moderate circularity (fitted input called prediction) without invalidating the overall architecture. No self-citation chains, ansatz smuggling, or other load-bearing reductions appear in the abstract or described claims.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Evolutionary distances between species can be used as a supervisory signal to structure a latent shape space in a biologically meaningful way
Reference graph
Works this paper leans on
-
[1]
A virtual world of paleontology,
J. A. Cunningham, I. A. Rahman, S. Lautenschlager, E. J. Rayfield, and P. C. Donoghue, “A virtual world of paleontology,”Trends in Ecology & Evolution, vol. 29, no. 6, pp. 347–357, 2014
2014
-
[2]
Open data and digital morphology,
T. G. Davies, I. A. Rahman, S. Lautenschlager, J. A. Cunningham, R. J. Asher, P. M. Barrett, K. T. Bates, S. Bengtson, R. B. J. Benson, D. M. Boyer, J. Braga, J. A. Bright, L. P. A. M. Claessens, P. G. Cox, X.-P. Dong, A. R. Evans, P. L. Falkingham, M. Friedman, R. J. Garwood, A. Goswami, J. R. Hutchinson, N. S. Jeffery, Z. Johanson, R. Lebrun, C. Martíne...
2017
-
[3]
Evolution and development of shape: integrating quantitative approaches,
C. P. Klingenberg, “Evolution and development of shape: integrating quantitative approaches,”Nature Reviews Genetics, vol. 11, no. 9, pp. 623–635, 2010
2010
-
[4]
Micro-computed tomography for natural history specimens: a handbook of best practice protocols,
K. Keklikoglou, S. Faulwetter, E. Chatzinikolaou, P. Wils, J. Brecko, J. Kvaček, B. Metscher, and C. Arvanitidis, “Micro-computed tomography for natural history specimens: a handbook of best practice protocols,”European Journal of Taxonomy, no. 522, pp. 1–55, 2019. 20
2019
-
[5]
A practical guide to sliding and surface semilandmarks in morphometric analyses,
C. Bardua, R. N. Felice, A. Watanabe, A. C. Fabre, and A. Goswami, “A practical guide to sliding and surface semilandmarks in morphometric analyses,”Integrative Organismal Biology, vol. 1, no. 1, pp. 1–34, 2019
2019
-
[6]
How many specimens make a sufficient training set for automated three-dimensional feature extraction?,
J. M. Mulqueeney, A. Searle-Barnes, A. Brombacher, M. Sweeney, A. Goswami, and T. H. G. Ezard, “How many specimens make a sufficient training set for automated three-dimensional feature extraction?,” Royal Society Open Science, vol. 11, no. 6, p. rsos.240113, 2024
2024
-
[7]
Auto-Encoding Variational Bayes
D. P. Kingma and M. Welling, “Auto-encoding variational Bayes,”arXiv preprint arXiv:1312.6114, 2013
work page internal anchor Pith review arXiv 2013
-
[8]
Generative adversarial networks,
I. Goodfellow, J. Pouget-Abadie, M. Mirza, B. Xu, D. Warde-Farley, S. Ozair, A. Courville, and Y. Bengio, “Generative adversarial networks,”Communications of the ACM, vol. 63, no. 11, pp. 139–144, 2020
2020
-
[9]
Denoising diffusion probabilistic models,
J. Ho, A. Jain, and P. Abbeel, “Denoising diffusion probabilistic models,”Advances in Neural Information Processing Systems, vol. 33, pp. 6840–6851, 2020
2020
-
[10]
Flow matching for generative modeling,
Y. Lipman, R. T. Chen, H. Ben-Hamu, M. Nickel, and M. Le, “Flow matching for generative modeling,” in11th International Conference on Learning Representations (ICLR 2023), 2023
2023
-
[11]
On memorization in probabilistic deep generative models,
G. van den Burg and C. Williams, “On memorization in probabilistic deep generative models,”Advances in Neural Information Processing Systems, vol. 34, pp. 27916–27928, 2021
2021
-
[12]
Understanding deep learning (still) requires rethinking generalization,
C. Zhang, S. Bengio, M. Hardt, B. Recht, and O. Vinyals, “Understanding deep learning (still) requires rethinking generalization,”Communications of the ACM, vol. 64, no. 3, pp. 107–115, 2021
2021
-
[13]
Phylogenies and the comparative method,
J. Felsenstein, “Phylogenies and the comparative method,”The American Naturalist, vol. 125, no. 1, pp. 1–15, 1985
1985
-
[14]
Understanding posterior collapse in generative latent variable models,
J. Lucas, G. Tucker, R. Grosse, and M. Norouzi, “Understanding posterior collapse in generative latent variable models,” inICLR 2019 Workshop DeepGenStruct, 2019
2019
-
[15]
Effectively unbiased FID and inception score and where to find them,
M. J. Chong and D. Forsyth, “Effectively unbiased FID and inception score and where to find them,” in2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 6069–6078, 2020
2020
-
[16]
DeepSDF: Learning continuous signed distance functions for shape representation,
J. J. Park, P. Florence, J. Straub, R. Newcombe, and S. Lovegrove, “DeepSDF: Learning continuous signed distance functions for shape representation,” in2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 165–174, 2019
2019
-
[17]
Computing the Fréchet distance between two polygonal curves,
H. Alt and M. Godau, “Computing the Fréchet distance between two polygonal curves,”International Journal of Computational Geometry & Applications, vol. 5, no. 1–2, pp. 75–91, 1995
1995
-
[18]
Occupancy networks: Learning 3D reconstruction in function space,
L. Mescheder, M. Oechsle, M. Niemeyer, S. Nowozin, and A. Geiger, “Occupancy networks: Learning 3D reconstruction in function space,” in2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 4455–4465, 2019
2019
-
[19]
Learning implicit fields for generative shape modeling,
Z. Chen and H. Zhang, “Learning implicit fields for generative shape modeling,” in2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), (Los Alamitos, CA, USA), pp. 5932– 5941, IEEE Computer Society, June 2019
2019
-
[20]
Neural unsigned distance fields for implicit function learning,
J. Chibane, A. Mir, and G. Pons-Moll, “Neural unsigned distance fields for implicit function learning,” inProceedings of the 34th International Conference on Neural Information Processing Systems, (Red Hook, NY, USA), pp. 21638–21652, Curran Associates Inc., 2020
2020
-
[21]
Differentiable volumetric rendering: Learning implicit 3D representations without 3D supervision,
M. Niemeyer, L. Mescheder, M. Oechsle, and A. Geiger, “Differentiable volumetric rendering: Learning implicit 3D representations without 3D supervision,” in2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 3501–3512, 2020
2020
-
[22]
Learning representations and generative models for 3D point clouds,
P. Achlioptas, O. Diamanti, I. Mitliagkas, and L. Guibas, “Learning representations and generative models for 3D point clouds,” inInternational Conference on Machine Learning, pp. 40–49, PMLR, 2018
2018
-
[23]
PointFlow: 3D point cloud generation with continuous normalizing flows,
G. Yang, X. Huang, Z. Hao, M.-Y. Liu, S. Belongie, and B. Hariharan, “PointFlow: 3D point cloud generation with continuous normalizing flows,” in2019 IEEE/CVF International Conference on Computer Vision (ICCV), pp. 4540–4549, 2019
2019
-
[24]
VoxNet: A3Dconvolutionalneuralnetworkforreal-timeobjectrecognition,
D.MaturanaandS.Scherer, “VoxNet: A3Dconvolutionalneuralnetworkforreal-timeobjectrecognition,” in2015 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), pp. 922–928, 2015
2015
-
[25]
PolyGen: An autoregressive generative model of 3D meshes,
C. Nash, Y. Ganin, S. M. A. Eslami, and P. Battaglia, “PolyGen: An autoregressive generative model of 3D meshes,” inProceedings of the 37th International Conference on Machine Learning(H. D. III and A. Singh, eds.), vol. 119 ofProceedings of Machine Learning Research, pp. 7220–7229, PMLR, 2020. 21
2020
-
[26]
DualSDF: Semantic shape manipulation using a two-level representation,
Z. Hao, H. Averbuch-Elor, N. Snavely, and S. Belongie, “DualSDF: Semantic shape manipulation using a two-level representation,” in2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), p. 7628–7638, IEEE, 2020
2020
-
[27]
LION: Latent point diffusion models for 3D shape generation,
A. Vahdat, F. Williams, Z. Gojcic, O. Litany, S. Fidler, K. Kreis,et al., “LION: Latent point diffusion models for 3D shape generation,”Advances in Neural Information Processing Systems, vol. 35, pp. 10021– 10039, 2022
2022
-
[28]
Phylogenetic principal components analysis and geometric morphometrics,
P. D. Polly, A. M. Lawing, A.-C. Fabre, and A. Goswami, “Phylogenetic principal components analysis and geometric morphometrics,”Hystrix, the Italian Journal of Mammalogy, vol. 24, pp. 33–41, Apr. 2013
2013
-
[29]
A registration and deep learning approach to automated landmark detection for geometric morphometrics,
J. Devine, J. D. Aponte, D. C. Katz, W. Liu, L. D. L. Vercio, N. D. Forkert, R. Marcucio, C. J. Percival, and B. Hallgrímsson, “A registration and deep learning approach to automated landmark detection for geometric morphometrics,”Evolutionary Biology, vol. 47, no. 3, pp. 246–259, 2020
2020
-
[30]
The phylogenetic effective sample size and jumps,
K. Bartoszek, “The phylogenetic effective sample size and jumps,”Mathematica Applicanda, vol. 46, July 2018
2018
-
[31]
Phylogenies and community ecology,
C. O. Webb, D. D. Ackerly, M. A. McPeek, and M. J. Donoghue, “Phylogenies and community ecology,” Annual Review of Ecology and Systematics, vol. 33, no. 1, pp. 475–505, 2002
2002
-
[32]
Inferring the historical patterns of biological evolution,
M. Pagel, “Inferring the historical patterns of biological evolution,”Nature, vol. 401, no. 6756, pp. 877–884, 1999
1999
-
[33]
Implicit geometric regularization for learning shapes,
A. Gropp, L. Yariv, N. Haim, M. Atzmon, and Y. Lipman, “Implicit geometric regularization for learning shapes,” inProceedings of the 37th International Conference on Machine Learning (ICML), pp. 3789–3799, 2020
2020
-
[34]
Level set methods and dynamic implicit surfaces,
S. Osher, R. Fedkiw, and K. Piechor, “Level set methods and dynamic implicit surfaces,”Applied Mechanics Reviews, vol. 57, no. 3, p. B15, 2004
2004
-
[35]
Marching cubes: A high resolution 3D surface construction algorithm,
W. E. Lorensen and H. E. Cline, “Marching cubes: A high resolution 3D surface construction algorithm,” inSeminal graphics: Pioneering efforts that shaped the field, pp. 347–353, 1998
1998
-
[36]
Phylogenetics and diversification of tanagers (Passeriformes: Thraupidae), the largest radiation of Neotropical songbirds,
K. J. Burns, A. J. Shultz, P. O. Title, N. A. Mason, F. K. Barker, J. Klicka, S. M. Lanyon, and I. J. Lovette, “Phylogenetics and diversification of tanagers (Passeriformes: Thraupidae), the largest radiation of Neotropical songbirds,”Molecular Phylogenetics and Evolution, vol. 75, pp. 41–77, 2014
2014
-
[37]
Phylogenetic relationships and morphological diversity in Darwin’s finches and their relatives,
K. J. Burns, S. J. Hackett, and N. K. Klein, “Phylogenetic relationships and morphological diversity in Darwin’s finches and their relatives,”Evolution, vol. 56, no. 6, pp. 1240–1252, 2002
2002
-
[38]
Building normalizing flows with stochastic interpolants,
M. S. Albergo and E. Vanden-Eijnden, “Building normalizing flows with stochastic interpolants,” in International Conference on Learning Representations (ICLR), 2023
2023
-
[39]
Improvingandgeneralizingflow-basedgenerativemodelswithminibatchoptimaltransport,
A. Tong, K. Fatras, N. Malkin, G. Huguet, Y. Zhang, J. Rector-Brooks, G. Wolf, and Y. Bengio, “Improvingandgeneralizingflow-basedgenerativemodelswithminibatchoptimaltransport,”Transactions on Machine Learning Research, pp. 1–34, 2024
2024
-
[40]
The Fréchet distance between multivariate normal distributions,
D. C. Dowson and B. V. Landau, “The Fréchet distance between multivariate normal distributions,” Journal of Multivariate Analysis, vol. 12, no. 3, pp. 450–455, 1982
1982
-
[41]
Cranial shape evolution in adaptive radiations of birds: comparative morphometrics of Darwin’s finches and Hawaiian honeycreepers,
M. Tokita, W. Yano, H. F. James, and A. Abzhanov, “Cranial shape evolution in adaptive radiations of birds: comparative morphometrics of Darwin’s finches and Hawaiian honeycreepers,”Philosophical Transactions of the Royal Society B: Biological Sciences, vol. 372, no. 1713, p. 20150481, 2017
2017
-
[42]
Geometry and dynamics link form, function, and evolution of finch beaks,
S. Al-Mosleh, G. P. T. Choi, A. Abzhanov, and L. Mahadevan, “Geometry and dynamics link form, function, and evolution of finch beaks,”Proceedings of the National Academy of Sciences, vol. 118, no. 46, p. e2105957118, 2021
2021
-
[43]
Beak morphometry and morphogenesis across avian radiations,
S. Mosleh, G. P. T. Choi, G. M. Musser, H. F. James, A. Abzhanov, and L. Mahadevan, “Beak morphometry and morphogenesis across avian radiations,”Proceedings of the Royal Society B: Biological Sciences, vol. 290, no. 2007, p. 20230420, 2023
2007
-
[44]
Adam: A method for stochastic optimization,
D. P. Kingma and J. Ba, “Adam: A method for stochastic optimization,” inInternational Conference on Learning Representations (ICLR), 2015
2015
-
[45]
Towards understanding convergence and generalization of AdamW,
P. Zhou, X. Xie, Z. Lin, and S. Yan, “Towards understanding convergence and generalization of AdamW,” IEEE Transactions on Pattern Analysis and Machine Intelligence, vol. 46, no. 9, pp. 6486–6493, 2024
2024
-
[46]
Rethinking network design and local geometry in point cloud: A simple residual MLP framework,
X. Ma, C. Qin, H. You, H. Ran, and Y. Fu, “Rethinking network design and local geometry in point cloud: A simple residual MLP framework,” inInternational Conference on Learning Representations (ICLR), 2022. 22
2022
-
[47]
Gaussian Error Linear Units (GELUs)
D. Hendrycks and K. Gimpel, “Gaussian error linear units (GELUs),”arXiv preprint arXiv:1606.08415, 2016
work page internal anchor Pith review arXiv 2016
-
[48]
GANs trained by a two time-scale update rule converge to a local Nash equilibrium,
M. Heusel, H. Ramsauer, T. Unterthiner, B. Nessler, and S. Hochreiter, “GANs trained by a two time-scale update rule converge to a local Nash equilibrium,”Advances in Neural Information Processing Systems, vol. 30, 2017
2017
-
[49]
Efficient Score Pre-computation for Diffusion Models via Cross-Matrix Krylov Projection
K. Lau, A. S. Na, and J. W. L. Wan, “Efficient score pre-computation for diffusion models via cross-matrix Krylov projection,”arXiv preprint arXiv:2511.17634, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[50]
Denoising diffusion implicit models,
J. Song, C. Meng, and S. Ermon, “Denoising diffusion implicit models,” inInternational Conference on Learning Representations (ICLR), 2021
2021
-
[51]
Sexual dimorphism,
D. W. Frayer and M. H. Wolpoff, “Sexual dimorphism,”Annual Review of Anthropology, vol. 14, pp. 429–473, 1985
1985
-
[52]
Exogenous dna loading into extracellular vesicles via electroporation is size-dependent and enables limited gene delivery,
T. N. Lamichhane, R. S. Raiker, and S. M. Jay, “Exogenous dna loading into extracellular vesicles via electroporation is size-dependent and enables limited gene delivery,”Molecular Pharmaceutics, vol. 12, no. 10, p. 3650–3657, 2015. 23 Supplementary Information Layer Input dim Output dim Notes Linear 164 + 3 = 67512 Weight norm + ReLU Linear 2–4 512 512 W...
2015
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.