Recognition: unknown
Robustness Evaluation of a Foundation Segmentation Model Under Simulated Domain Shifts in Abdominal CT: Implications for Health Digital Twin Deployment
Pith reviewed 2026-05-07 13:59 UTC · model grok-4.3
The pith
SAM exhibits stable spleen segmentation performance under simulated CT domain shifts
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
SAM (ViT-B) achieves a mean Dice score of 0.9145 on clean abdominal CT slices for spleen segmentation. When subjected to ten conditions of perturbations simulating inter-scanner variability, the absolute mean change in Dice remains below 0.01. Statistical analysis shows some small significant differences but McNemar tests indicate no significant rise in failure rates. These results support that SAM maintains stable segmentation behavior under moderate domain shifts.
What carries the argument
A slice-level robustness audit using ground-truth-derived bounding box prompts on 1,051 CT slices under five types of controlled perturbations to measure changes in Dice score.
If this is right
- SAM serves as a robust foundation baseline for medical image segmentation research.
- Health digital twins can use SAM for anatomical modeling with reduced concern for moderate imaging variability.
- Characterizing robustness under real-world variability is essential for trustworthy deployment of foundation models in medicine.
- Small-magnitude changes do not lead to increased segmentation failures in the tested conditions.
Where Pith is reading between the lines
- If the simulated perturbations capture key aspects of real scanner differences, SAM could reduce the need for per-center retraining in multi-site studies.
- Extending this audit to other organs and full 3D volumes could strengthen the case for broad deployment.
- Comparison with other foundation models might reveal whether this stability is unique to SAM or common across similar architectures.
Load-bearing premise
The five controlled perturbations sufficiently represent clinically realistic inter-scanner domain shifts in abdominal CT.
What would settle it
A study using actual multi-scanner abdominal CT datasets showing mean Dice changes exceeding 0.01 or increased failure rates would challenge the stability claim.
Figures
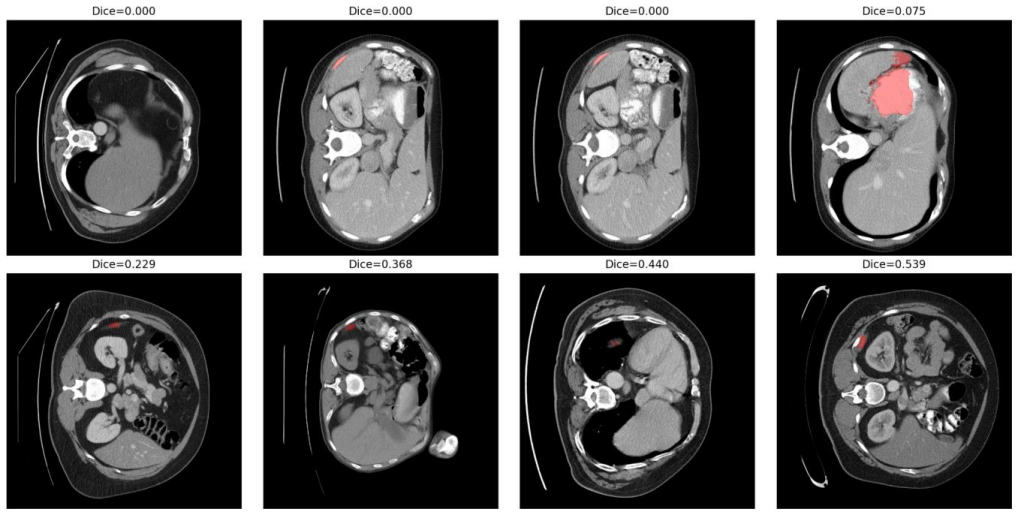

read the original abstract
Foundation segmentation models such as the Segment Anything Model (SAM) have demonstrated strong generalization across natural images; however, their robustness under clinically realistic medical imaging domain shifts remains insufficiently quantified. We present a systematic slice-level robustness audit of SAM (ViT-B) for spleen segmentation in abdominal CT using 1,051 nonempty slices from 41 volumes in the Medical Segmentation Decathlon. A standardized ground-truth-derived bounding-box protocol was used to isolate encoder robustness from prompt uncertainty. Controlled perturbations simulating inter-scanner variability, including Gaussian noise, blur, contrast scaling, gamma correction, and resolution mismatch, were applied across ten conditions. The clean baseline achieved a mean Dice score of 0.9145 (95% CI: [0.909, 0.919]) with a failure rate of 0.67%. Across all perturbations, the absolute mean {\Delta}Dice remained below 0.01. Paired Wilcoxon signed-rank tests with Benjamini-Hochberg false discovery rate correction identified statistically significant but small-magnitude changes under selected conditions, while McNemar analysis showed no significant increase in failure probability. These findings indicate that SAM exhibits stable segmentation behavior under moderate CT domain shifts, supporting its role as a robust foundation baseline for medical image segmentation research. As health digital twins increasingly incorporate foundation segmentation models for anatomical modeling and organ-level monitoring, formal characterization of robustness under real-world imaging variability is a necessary step toward trustworthy deployment.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper evaluates the robustness of the Segment Anything Model (SAM, ViT-B) for spleen segmentation in abdominal CT using 1,051 nonempty slices from 41 volumes in the Medical Segmentation Decathlon. A ground-truth-derived bounding-box prompt protocol isolates encoder performance. Five controlled perturbations (Gaussian noise, blur, contrast scaling, gamma correction, resolution mismatch) simulate inter-scanner domain shifts across ten conditions. Baseline mean Dice is 0.9145 (95% CI [0.909, 0.919]) with 0.67% failure rate. Absolute mean ΔDice remains below 0.01 under all perturbations. Paired Wilcoxon signed-rank tests with Benjamini-Hochberg FDR correction detect statistically significant but small-magnitude changes in selected conditions, while McNemar tests show no significant increase in failure probability. The authors conclude that SAM exhibits stable behavior under these moderate simulated shifts, supporting its role as a robust foundation baseline for medical segmentation and health digital twin applications.
Significance. If the results hold, the work supplies a transparent, statistically controlled empirical baseline for SAM robustness in a medical imaging task on public data, with explicit perturbation definitions and a bounding-box protocol that cleanly separates encoder stability from prompt variability. The small observed effect sizes and lack of failure-rate increase provide a useful reference for foundation-model research. The study correctly emphasizes the need for formal robustness characterization prior to clinical deployment.
major comments (1)
- [Abstract] Abstract (perturbation definitions): The five perturbations are all global and stationary (uniform noise, global scaling, etc.). This choice is load-bearing for the implications claim because real inter-scanner CT variability typically includes non-stationary, spatially correlated effects such as vendor-specific reconstruction kernels, beam-hardening, and kVp/mAs-dependent artifacts. The observed |ΔDice| < 0.01 stability may therefore not generalize to actual multi-vendor clinical data, limiting support for health digital twin deployment.
minor comments (1)
- The manuscript would benefit from a short dedicated limitations paragraph explicitly discussing the scope of the chosen simulations relative to real scanner variability.
Simulated Author's Rebuttal
We thank the referee for the positive evaluation and the recommendation of minor revision. We address the single major comment point by point below.
read point-by-point responses
-
Referee: [Abstract] Abstract (perturbation definitions): The five perturbations are all global and stationary (uniform noise, global scaling, etc.). This choice is load-bearing for the implications claim because real inter-scanner CT variability typically includes non-stationary, spatially correlated effects such as vendor-specific reconstruction kernels, beam-hardening, and kVp/mAs-dependent artifacts. The observed |ΔDice| < 0.01 stability may therefore not generalize to actual multi-vendor clinical data, limiting support for health digital twin deployment.
Authors: We agree that the five perturbations are global and stationary, representing a controlled simplification that does not encompass the full spectrum of real inter-scanner effects such as spatially varying beam-hardening, vendor-specific kernels, or kVp/mAs artifacts. The experimental protocol was intentionally designed to isolate encoder behavior under reproducible, moderate intensity and resolution shifts using public data, yielding a transparent baseline rather than a comprehensive claim of robustness across all clinical domain shifts. The manuscript already qualifies the results as applying to 'moderate simulated shifts' and positions the work as supplying an empirical reference point. To directly address the concern, we will revise the abstract to more explicitly qualify the perturbation scope and add a limitations subsection in the discussion that acknowledges the gap to non-stationary multi-vendor data and calls for further validation on heterogeneous clinical cohorts. This revision clarifies the implications for health digital twin deployment without changing the reported quantitative findings. revision: yes
Circularity Check
No circularity: purely empirical robustness audit with no derivations or self-referential predictions
full rationale
The paper performs a direct empirical evaluation by applying five predefined perturbations to CT slices, computing Dice scores and failure rates against ground-truth bounding boxes, and reporting statistical tests on the observed differences. No equations, fitted parameters, predictions, or uniqueness claims are present that reduce to quantities defined by the authors' own choices. The central claim of stability (mean |ΔDice| < 0.01) follows immediately from the measured values under the stated conditions without any intermediate derivation or self-citation load-bearing step. The perturbations are explicitly described as controlled simulations rather than derived from a model, so the evaluation chain is self-contained and externally falsifiable via replication on the same dataset.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption The chosen perturbations adequately represent moderate inter-scanner variability in abdominal CT.
- domain assumption Ground-truth-derived bounding boxes isolate encoder robustness from prompt uncertainty.
Reference graph
Works this paper leans on
-
[1]
Segment Anything,
A. Kirillov et al., “Segment Anything,” in Proc. IEEE/CVF ICCV, 2023, pp. 4015-4026
2023
-
[2]
J. Ma and B. Wang, “Segment Anything in Medical Images,” Nature Commun., vol. 15, no. 1, Art. no. 654, Jan. 2024, doi: 10.1038/s41467-024-44824-z
-
[3]
Can You Trust Your Model’s Uncertainty? Evaluating Predictive Uncertainty Under Dataset Shift,
Y. Ovadia et al., “Can You Trust Your Model’s Uncertainty? Evaluating Predictive Uncertainty Under Dataset Shift,” in Proc. NeurIPS, 2019
2019
-
[4]
Adversarial Attacks on Medical Machine Learning,
S. G. Finlayson et al., “Adversarial Attacks on Medical Machine Learning,” Science, vol. 363, no. 6433, pp. 1287 -1289, Mar. 2019
2019
-
[5]
Machine Learning with Multi-Site Imaging Data: An Empirical Study on the Impact of Scanner Effects,
B. Glocker et al., “Machine Learning with Multi-Site Imaging Data: An Empirical Study on the Impact of Scanner Effects,” arXiv:1910.04597, 2019
-
[6]
A. L. Simpson et al., “A Large Annotated Medical Image Dataset for the Development and Evaluation of Segmentation Algorithms,” arXiv:1902.09063, 2019
-
[7]
Digital Twins for Health: A Scoping Review,
E. Bjornsson et al., “Digital Twins for Health: A Scoping Review,” npj Digital Medicine, vol. 6, no. 1, Art. no. 44, Mar. 2023, doi: 10.1038/s41746-023-00778-8
-
[8]
nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation,
F. Isensee, P. F. Jaeger, S. A. A. Kohl, J. Petersen, and K. H. Maier -Hein, “nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation,” Nature Methods, vol. 18, pp. 203 -211, 2021, doi: 10.1038/s41592-020- 01008-z
-
[9]
Segment anything model for medical image analysis: An experimental study,
M. A. Mazurowski, H. Dong, H. Gu, J. Yang, N. Konz, and Y. Zhang, “Segment anything model for medical image analysis: An experimental study,” Medical Image Analysis, vol. 89, p. 102918, 2023, doi: 10.1016/j.media.2023.102918
-
[10]
The multimodal brain tumor image segmentation benchmark (BRATS),
B. H. Menze et al., “The multimodal brain tumor image segmentation benchmark (BRATS),” IEEE Trans. Med. Imaging, vol. 34, no. 10, pp. 1993-2024, 2015, doi: 10.1109/TMI.2014.2377694
-
[11]
N. Heller et al., “The KiTS23 Challenge: 500 kidney tumor cases with accompanying task hierarchy,” arXiv:2307.01984, 2023
-
[12]
A multi -organ nucleus segmentation challenge,
N. Kumar et al., “A multi -organ nucleus segmentation challenge,” IEEE Trans. Med. Imaging, vol. 39, no. 5, pp. 1380 - 1391, 2019, doi: 10.1109/TMI.2019.2947628
-
[13]
Tizabi, Florian Buber, Eva Röth, Doro Ambros-Glasser, Björn Menze, and Annette Kopp-Schneider
L. Maier-Hein et al., “Metrics reloaded: Recommendations for image analysis validation,” Nature Methods, vol. 21, pp. 195-212, 2024, doi: 10.1038/s41592-023-02151-z
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.