Recognition: unknown
Bayesian Nonparametric Causal Inference for Quantile Residual Life: An Application to Alzheimer's Disease
Pith reviewed 2026-05-07 09:11 UTC · model grok-4.3
The pith
Elevated baseline amyloid shortens quantiles of remaining dementia-free time in Alzheimer's patients who stay event-free to landmark times.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Using Bayesian g-computation with an enriched Dirichlet process mixture model for the joint distribution of dementia onset times, amyloid exposure, and baseline covariates, the analysis shows that elevated baseline amyloid is associated with shorter quantiles of remaining dementia-free time than non-elevated amyloid among individuals dementia-free through landmark times, overall and within baseline diagnostic subgroups.
What carries the argument
Enriched Dirichlet process mixture model for the joint distribution of event times, exposure, and covariates, combined with Bayesian g-computation to obtain causal contrasts on quantile residual life.
If this is right
- Quantile residual life contrasts can be estimated at multiple landmark times without assuming a parametric survival form.
- The same framework yields subgroup-specific causal effects within baseline diagnostic categories.
- Data augmentation within the model permits valid inference when some baseline covariates are missing.
- Sensitivity parameters allow assessment of how strong residual confounding would need to be to overturn the observed contrasts.
Where Pith is reading between the lines
- The method could be applied to other censored time-to-event settings where interest lies in conditional remaining survival quantiles rather than hazard ratios.
- Landmark-specific results may help clinicians set time-bound prognostic thresholds for patients who have already survived initial follow-up.
- If the ignorability assumption holds only conditionally on richer covariate sets, future studies could incorporate additional biomarkers to tighten the causal contrast.
Load-bearing premise
Amyloid status is ignorable given the observed baseline covariates, with no substantial unmeasured confounding.
What would settle it
A dataset in which, after the same covariate adjustment and landmark conditioning, the estimated quantile residual life contrasts between elevated and non-elevated amyloid groups are statistically indistinguishable from zero.
Figures



read the original abstract
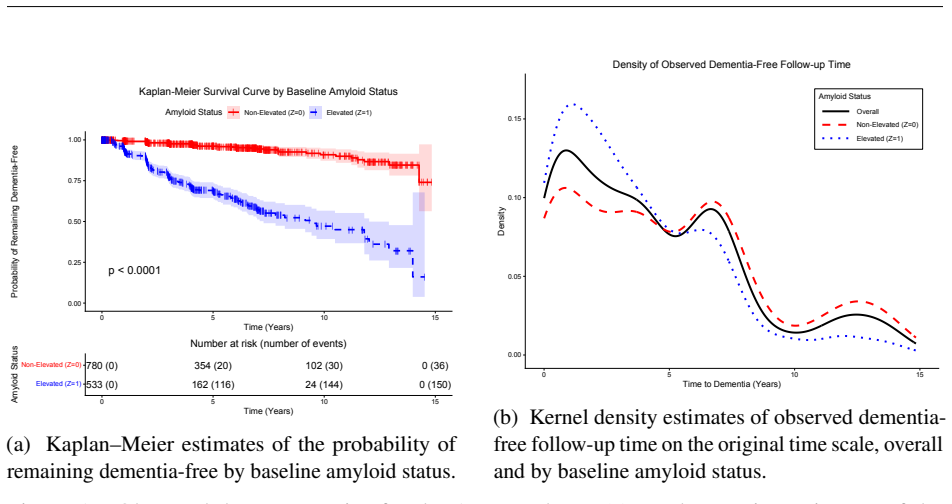
In Alzheimer's disease research, for individuals who remain dementia-free through a given follow-up time, an important clinical question is how much longer they are likely to remain dementia-free. Quantiles of this remaining time provide clinically interpretable prognostic milestones and can help characterize prognostic heterogeneity across baseline groups. We address this question in the Alzheimer's Disease Neuroimaging Initiative (ADNI), focusing on baseline amyloid status as the exposure. Estimation is challenging because amyloid status is observed rather than randomized, requiring adjustment for confounding, and because time to dementia onset is heterogeneous and heavily right-censored. We estimate causal contrasts in quantile residual life using a Bayesian nonparametric enriched Dirichlet process mixture model for the joint distribution of event times, exposure, and baseline covariates, with inference via Bayesian g-computation. The approach accommodates ignorable missing baseline covariates through data augmentation, supports inference across clinically relevant landmark times, and allows sensitivity analysis for residual unmeasured confounding. Simulation studies show good performance under complex heterogeneity and heavy censoring. In ADNI, elevated baseline amyloid was associated with shorter quantiles of remaining dementia-free time than non-elevated baseline amyloid among individuals who remained dementia-free through relevant landmark times, overall and within baseline diagnostic subgroups.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes a Bayesian nonparametric enriched Dirichlet process mixture model for the joint distribution of event times, exposure (baseline amyloid status), and baseline covariates. Causal contrasts in quantile residual life are estimated via Bayesian g-computation, with data augmentation for missing covariates. The method is evaluated in simulations under heterogeneity and heavy censoring, then applied to ADNI data to compare quantiles of remaining dementia-free time (overall and within diagnostic subgroups) for individuals remaining dementia-free at landmark times.
Significance. If the results hold, the work offers a flexible BNP framework for causal quantile residual life analysis that directly handles heavy right-censoring, prognostic heterogeneity, and ignorable missing covariates while supporting g-computation. The simulation studies and the clinical relevance of the ADNI application to Alzheimer's prognosis are strengths.
major comments (2)
- [Abstract] Abstract: the manuscript states that the approach 'allows sensitivity analysis for residual unmeasured confounding,' yet provides no indication that this analysis was executed on the ADNI data or that the reported quantile differences remain stable under plausible residual confounding. In an observational cohort where amyloid status is not randomized, this omission is load-bearing for the causal interpretation of the g-computation results.
- [Results] ADNI application (results section): the central estimates arise from posterior inference on the joint model followed by g-computation, but the manuscript does not report detailed posterior predictive checks, model-fit diagnostics, or sensitivity to the enriched Dirichlet process hyperparameters for the real-data analysis. This limits verification that the reported associations are robust to the nonparametric modeling choices.
minor comments (1)
- [Methods] Notation for the quantile residual life functional and the g-computation step could be clarified with an explicit equation in the methods section to aid reproducibility.
Simulated Author's Rebuttal
We thank the referee for their constructive comments on our manuscript. We address each major comment below and describe the revisions we will make to strengthen the paper.
read point-by-point responses
-
Referee: [Abstract] Abstract: the manuscript states that the approach 'allows sensitivity analysis for residual unmeasured confounding,' yet provides no indication that this analysis was executed on the ADNI data or that the reported quantile differences remain stable under plausible residual confounding. In an observational cohort where amyloid status is not randomized, this omission is load-bearing for the causal interpretation of the g-computation results.
Authors: We agree that demonstrating the sensitivity analysis on the ADNI data is important for supporting the causal interpretation, given that amyloid status is observational. The abstract describes a capability of the Bayesian g-computation framework rather than a completed analysis. In the revised manuscript, we will conduct and report a sensitivity analysis for residual unmeasured confounding on the ADNI data, assessing stability of the quantile residual life contrasts under plausible levels of unmeasured confounding. revision: yes
-
Referee: [Results] ADNI application (results section): the central estimates arise from posterior inference on the joint model followed by g-computation, but the manuscript does not report detailed posterior predictive checks, model-fit diagnostics, or sensitivity to the enriched Dirichlet process hyperparameters for the real-data analysis. This limits verification that the reported associations are robust to the nonparametric modeling choices.
Authors: We acknowledge that explicit model diagnostics for the ADNI application would improve verification of robustness. The simulation studies validate performance under heterogeneity and heavy censoring, but we agree these do not substitute for real-data checks. In the revised manuscript, we will add posterior predictive checks, model-fit diagnostics, and sensitivity analyses to the enriched Dirichlet process hyperparameters for the ADNI data analysis. revision: yes
Circularity Check
No significant circularity in derivation chain
full rationale
The paper estimates causal contrasts in quantile residual life via posterior inference on a Bayesian nonparametric enriched Dirichlet process mixture model for the joint distribution of event times, exposure, and covariates, followed by Bayesian g-computation. This does not reduce any target quantity to a direct fit or self-definition by construction. The approach is validated through simulation studies under heterogeneity and censoring, and the central results follow from standard Bayesian updating and g-computation rather than tautological renaming or fitted-input-as-prediction. No load-bearing steps rely on self-citation chains that render the result equivalent to its inputs. The derivation remains self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Amyloid status is ignorable given observed baseline covariates
- domain assumption Censoring is non-informative conditional on observed data
Reference graph
Works this paper leans on
-
[1]
(a) If the current value of si is in one of these clusters, then draw values θ˚ and ω˚ from prior distributionsG θ 0 andG ω 0 fory-clusterstK ´i `1,
Sample cluster membership for each subjecti,s i “ psy i , sx i q. (a) If the current value of si is in one of these clusters, then draw values θ˚ and ω˚ from prior distributionsG θ 0 andG ω 0 fory-clusterstK ´i `1, . . . , K ´i `K newu. (b) If the current value of sy i is in an existing y-cluster (say, y-cluster k) but not an existing x-subcluster, then s...
-
[2]
, sy N u, updateθ ˚ k from P ` θ˚ k |y, z,x;s,θ ˚ ´k,ω ˚˘ 9G θ 0 pθ˚ kq ź i:sy i “k fpy|z,x;θ ˚ kq
Givens i and data, sampleθandωfrom their conditional distributions (a) For each uniquekins y “ ts y 1, . . . , sy N u, updateθ ˚ k from P ` θ˚ k |y, z,x;s,θ ˚ ´k,ω ˚˘ 9G θ 0 pθ˚ kq ź i:sy i “k fpy|z,x;θ ˚ kq . For the continuous outcome with θ˚ “ pβ ˚, σ2,˚q, the base measures for θ are Gθ 0 pβq “ Npaβ, σ2cβBβq, and Gθ 0 pσ2q “Scale-Inv-χ 2paσ, bσq. Set a...
-
[3]
Use aθ “b θ “a ω “ bω “1
Update hyperparameters αθ and αωpθq: We specify Gammapa θ, bθq. Use aθ “b θ “a ω “ bω “1. (a) First drawξ„Beta ` αθ `1, N ˘ and drawα θ from αθ „ϱGammapa θ `K, b θ ´logpξqq ` p1´ϱqGammapa θ `K´1, b θ ´logpξqq where ϱ“ ´ aθ`K´1 Npb θ´logpξqq ¯ M ´ 1` aθ`K´1 Npb θ´logpξqq ¯ “ aθ`K´1 Npb θ´logpξqq`paθ`K´1q (Escobar & West 1995). (b) To updateα ω, use Metropo...
1995
-
[4]
, K`1u with probabilities proportional to tn1,
Draw a cluster assignment sy pmq P t1, . . . , K`1u with probabilities proportional to tn1, . . . , nK, αθu, whereKis the current number of occupied main clusters
-
[5]
, Kk `1u with probabilities proportional to tn1|k,
Draw sx pmq P t1, . . . , Kk `1u with probabilities proportional to tn1|k, . . . , nKk|k, αωukďK (an existing main cluster) where k“s y pmq. If k“K`1 (a newly generated main cluster), set sx pmq “1
-
[6]
If a new subcluster was generated in the previous step, its parametersω ˚ sx pmq|sy pmq are first drawn from the prior distribution
Draw a synthetic covariate vectorxpmq from f ´ x;ω ˚ sx pmq|sy pmq ¯ . If a new subcluster was generated in the previous step, its parametersω ˚ sx pmq|sy pmq are first drawn from the prior distribution. Page 44 of 53 Using the generated set of M synthetic values ␣ xpmq,s pmq (M m“1 where spmq “ ´ sy pmq, sx pmq ¯ , the expected conditional survival funct...
2012
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.