Recognition: unknown
A Target-Free Harmonization Method for MRI
Pith reviewed 2026-05-10 14:49 UTC · model grok-4.3
The pith
TgtFreeHarmony harmonizes MRI images to an unseen target domain by searching a style manifold with Bayesian optimization driven by downstream task performance.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
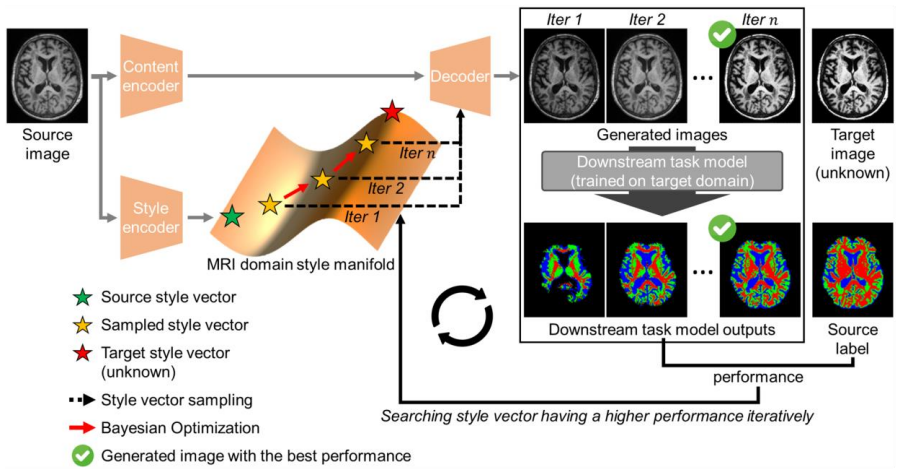
By training a disentanglement-based generator to separate content from style in MRI images and then using Bayesian optimization to select a point on the resulting style manifold according to how well a downstream model performs, source-domain images can be mapped to the appearance of an inaccessible target domain without any target data, data sharing, or direct style examples.
What carries the argument
A disentanglement-based generator that constructs a continuous manifold of MRI domain styles, searched by Bayesian optimization whose objective is the performance of a downstream task model trained exclusively on target-domain data.
If this is right
- Institutions can adjust their MRI data to match an external target style while keeping all scans inside their own secure environment.
- Models trained on one site's data become usable on images from other sites after local harmonization.
- Harmonization becomes practical for multi-center studies where patient data cannot leave the originating hospital.
- The same style-search process can be rerun if the downstream task or target requirements change.
Where Pith is reading between the lines
- This style-manifold search could be combined with federated learning so that even the optimization step avoids moving raw images.
- If the manifold is built from a larger set of scanners, the method might generalize to harmonizing toward styles never seen during manifold construction.
- The approach opens the possibility of on-device harmonization inside scanner consoles before images are stored.
Load-bearing premise
That maximizing downstream task performance on images transformed by a chosen style point will reliably recover the correct target appearance without altering biological content.
What would settle it
Downstream task accuracy on harmonized source images remains no better than on raw source images when both are evaluated against ground-truth labels from the target domain, or expert review detects loss of anatomical structures in the transformed images.
Figures




read the original abstract
In MRI, variations in scan parameters, sequence, or hardware can lead to discrepancies in image appearance, even for the same subject. These inconsistencies, known as domain shifts, can hinder image analysis and degrade the performance of deep learning models trained on data from specific target domains. MRI image harmonization aims to address these issues by aligning source domain images to the target domain images while preserving biological information such as anatomical structures. However, most existing harmonization approaches require access to both source and target domain data in training or test time. This dependence induces data sharing between institutions, raising concerns about patient privacy and substantially limiting the harmonization approaches that can be practically deployed in clinical settings. To overcome these limitations, we introduce TgtFreeHarmony, the harmonization framework tailored for target-free scenarios, eliminating the need for target domain data and any data sharing, enabling privacy-preserving harmonization directly within the source institution. Our approach estimates the target domain style by searching the manifold of MRI domain style constructed via a disentanglement-based generator using Bayesian optimization guided by the performance of a downstream task model, which is trained on target domain data. We evaluated our method on the brain tissue segmentation task across multiple institutes and demonstrated that it effectively harmonizes source images into target images, leading to improved downstream task performance. By enabling harmonization without any access to target-domain data, TgtFreeHarmony establishes a new direction of harmonization preserving data privacy that can be realistically deployed within clinical environments.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces TgtFreeHarmony, a target-free MRI harmonization framework. It builds a style manifold via a disentanglement-based generator and uses Bayesian optimization, guided solely by the performance of a downstream task model pre-trained on target-domain data, to select a style code for source images. The approach is evaluated on brain tissue segmentation across institutes and claims to achieve effective harmonization, improved task performance, and privacy preservation by eliminating any need for target data access or sharing.
Significance. If the empirical claims are substantiated, the work would be significant for enabling privacy-compliant harmonization in multi-institutional clinical settings where data sharing is restricted. It introduces a novel guidance mechanism (downstream-task performance on a style manifold) that could generalize to other imaging modalities and tasks, addressing a practical barrier in deploying domain-adapted models without compromising patient data confidentiality.
major comments (3)
- [Abstract] Abstract: The central claim of 'improved downstream task performance' and 'effective harmonization' is asserted without any quantitative metrics (e.g., Dice scores, Hausdorff distances), error bars, statistical significance tests, baseline comparisons, or ablation results. This absence leaves the magnitude and reliability of the improvement unverifiable and directly undermines assessment of whether the Bayesian optimization recovers a style inside the target distribution.
- [Method] Method (style manifold and optimization): The claim that Bayesian optimization on the disentanglement manifold will identify the target style rests on the untested assumption that the scalar downstream-task metric is maximized exclusively at the true target style point. No distribution-alignment loss, style-matching term, or post-hoc validation (e.g., FID to held-out target images or visual inspection) is described to rule out the possibility that the optimizer selects a non-target manifold point that boosts task accuracy while subtly altering anatomy. This is load-bearing for the target-free claim.
- [Experiments] Experiments: The evaluation across institutes is referenced but lacks details on dataset sizes, train/test splits, number of optimization iterations, hyperparameter sensitivity of the Bayesian search, or controls showing that biological content is preserved beyond the task metric. Without these, reproducibility and the risk of content leakage cannot be evaluated.
minor comments (2)
- [Abstract] The abstract contains several long, compound sentences that reduce readability; splitting the description of the optimization procedure would improve clarity.
- [Method] Notation for the style code and manifold parameters should be introduced consistently when first used in the method section to avoid ambiguity for readers unfamiliar with disentanglement literature.
Simulated Author's Rebuttal
We appreciate the referee's detailed feedback on our manuscript. We have carefully considered each comment and provide point-by-point responses below. Where appropriate, we will revise the manuscript to address the concerns raised.
read point-by-point responses
-
Referee: [Abstract] Abstract: The central claim of 'improved downstream task performance' and 'effective harmonization' is asserted without any quantitative metrics (e.g., Dice scores, Hausdorff distances), error bars, statistical significance tests, baseline comparisons, or ablation results. This absence leaves the magnitude and reliability of the improvement unverifiable and directly undermines assessment of whether the Bayesian optimization recovers a style inside the target distribution.
Authors: We agree that the abstract would benefit from including quantitative results to substantiate our claims. In the revised version of the manuscript, we will incorporate key performance metrics from our experiments, including Dice scores with error bars, comparisons to baseline methods, and mention of statistical tests. This will provide readers with a clearer understanding of the improvements achieved by TgtFreeHarmony. revision: yes
-
Referee: [Method] Method (style manifold and optimization): The claim that Bayesian optimization on the disentanglement manifold will identify the target style rests on the untested assumption that the scalar downstream-task metric is maximized exclusively at the true target style point. No distribution-alignment loss, style-matching term, or post-hoc validation (e.g., FID to held-out target images or visual inspection) is described to rule out the possibility that the optimizer selects a non-target manifold point that boosts task accuracy while subtly altering anatomy. This is load-bearing for the target-free claim.
Authors: The guidance mechanism relies on the pre-trained downstream task model, which has learned the target domain characteristics, to steer the optimization towards styles that enhance task performance. Given the disentangled nature of the generator, changes in style codes are intended to affect only the appearance (e.g., contrast, intensity) while preserving anatomical structures. We will add a more thorough discussion in the revised manuscript explaining this rationale and include additional ablation studies on the optimization process. However, direct post-hoc validation using metrics like FID on target images is not feasible without accessing target data, which would violate the target-free premise. Instead, we rely on the task performance as the indicator and will provide more indirect evidence of content preservation. revision: partial
-
Referee: [Experiments] Experiments: The evaluation across institutes is referenced but lacks details on dataset sizes, train/test splits, number of optimization iterations, hyperparameter sensitivity of the Bayesian search, or controls showing that biological content is preserved beyond the task metric. Without these, reproducibility and the risk of content leakage cannot be evaluated.
Authors: We will revise the experiments section to provide comprehensive details on the datasets used, including sizes and splits, the number of Bayesian optimization iterations, sensitivity analysis for hyperparameters, and additional controls or visualizations to demonstrate preservation of biological content. This will enhance the reproducibility of our results. revision: yes
- Direct validation against target domain distributions (such as FID scores) cannot be performed as it would require access to target data, contradicting the target-free approach.
Circularity Check
No significant circularity in the target-free harmonization derivation
full rationale
The paper's central method constructs a style manifold from a disentanglement-based generator and selects a style code via Bayesian optimization whose objective is the scalar performance of an independently trained downstream task model on target data. This chain does not reduce by construction to its inputs: the optimization is not a self-definition of the target style, the selected style is not a fitted parameter renamed as a prediction, and no self-citation or uniqueness theorem is invoked to force the result. The evaluation on brain tissue segmentation across institutes supplies an external benchmark, keeping the privacy-preserving claim independent of tautological equivalence.
Axiom & Free-Parameter Ledger
free parameters (2)
- Bayesian optimization hyperparameters
- Disentanglement generator parameters
axioms (2)
- domain assumption The disentanglement-based generator accurately separates domain-specific style from anatomical content in MRI images.
- domain assumption Performance of a downstream task model serves as a reliable indicator of harmonization quality.
invented entities (1)
-
MRI domain style manifold
no independent evidence
Reference graph
Works this paper leans on
-
[1]
Infogan: Interpretable representation learning by information maximizing generative adversarial nets
Chen, X., Duan, Y., Houthooft, R., Schulman, J., Sutskever, I., Abbeel, P., 2016. Infogan: Interpretable representation learning by information maximizing generative adversarial nets. Advances in neural information processing systems 29
2016
-
[2]
Ronneberger, O., Fischer, P., Brox, T., 2015. U -net: Convolutional networks for biomedical image segmentation, Medical Image Computing and Computer -Assisted Intervention – MICCAI 2015: 18th International Conference, Munich, Germany, October 5 -9, 2015,
2015
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.