Recognition: unknown
ORBIT: Learning Gene Program Co-Activation Structure for Cell-Type-Stratified Pathway Rewiring Analysis in Single-Cell Transcriptomics
Pith reviewed 2026-05-08 01:38 UTC · model grok-4.3
The pith
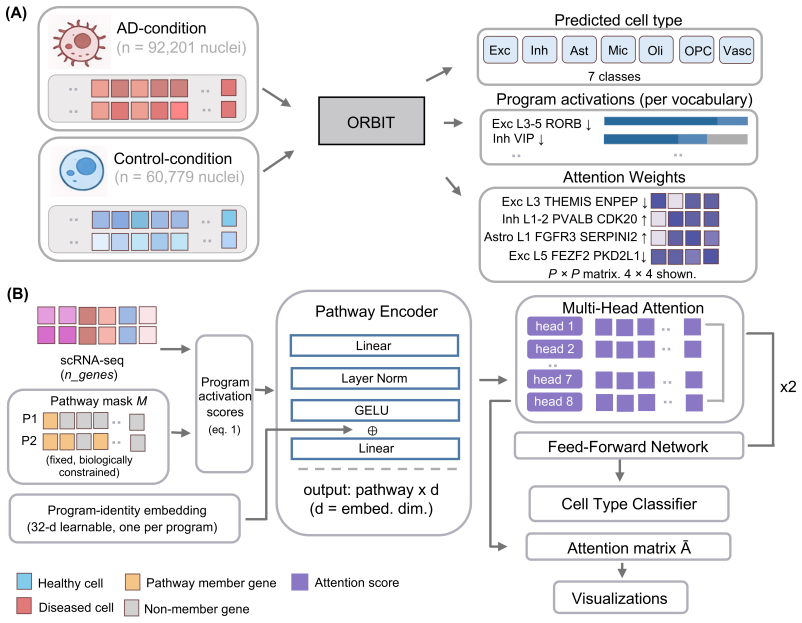
ORBIT learns asymmetric directional influences among gene programs from observational single-cell RNA-seq data alone by training a transformer to predict how other programs respond to simulated program removal.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
ORBIT recovers co-activation structure consistent with established Alzheimer's disease vulnerability signatures, identifies cell-type-specific rewiring invisible to differential expression, and achieves 0.984 macro F1 on cell-type classification from 220 pathway scores, which is within 0.3 points of a state-of-the-art classifier using all 22,088 genes.
What carries the argument
intervention-consistent training objective that trains the transformer to predict the change in every other gene program when one program is removed, so that the learned attention weights encode asymmetric directional influence rather than symmetric co-occurrence
If this is right
- Cell-type-stratified pathway rewiring becomes detectable in observational data without needing perturbation experiments for each cell type.
- Cell-type classification can be performed at high accuracy using only a few hundred pathway scores rather than the full transcriptome.
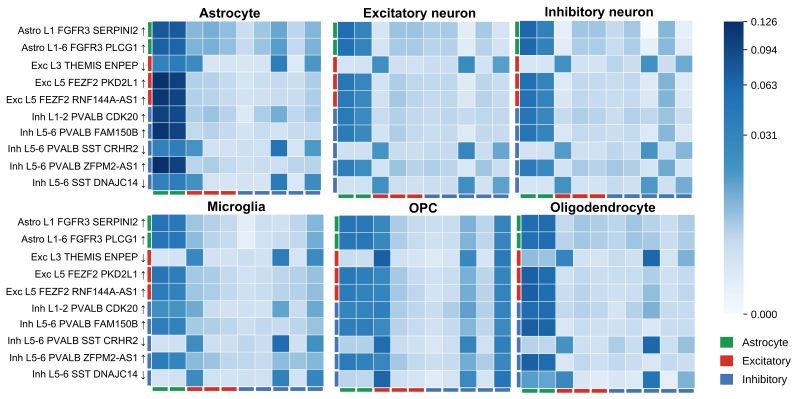
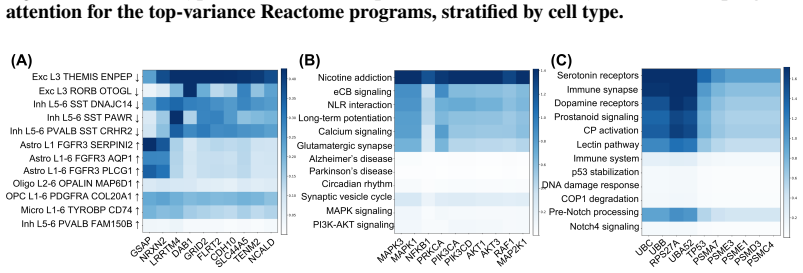
- Co-activation patterns recovered in Alzheimer's prefrontal cortex align with known vulnerability signatures and reveal rewiring not seen by standard differential expression.
- The same learned influence structure can be stratified by cell type to compare how program dependencies change across healthy and diseased states.
Where Pith is reading between the lines
- The approach could be extended to other diseases where only observational single-cell data are available, allowing systematic mapping of program rewiring without new wet-lab perturbations.
- If the directional influences prove stable across datasets, the model could serve as a prior for designing targeted perturbation experiments that focus on the strongest predicted links.
- Because the output is a directed graph over programs rather than a list of differentially expressed genes, it may help prioritize upstream regulators in cell-type-specific disease mechanisms.
Load-bearing premise
The intervention-consistent training objective, which predicts changes after simulated program removal, accurately reflects true directional biological influences among gene programs even though no actual experimental perturbations were performed.
What would settle it
Apply ORBIT to a single-cell dataset that also contains matched perturbation experiments for the same gene programs and check whether the model's predicted directional influences match the observed post-perturbation expression changes.
Figures




read the original abstract
Gene programs co-activate within cells, but existing single-cell methods either treat programs independently or require experimental perturbation data to model their interactions. We introduce ORBIT, a self-supervised transformer that learns asymmetric dependencies among gene programs from observational single-cell RNA-sequencing data alone, quantifying how strongly each program influences every other program. The key mechanism is an intervention-consistent training objective: the model learns each program's directional influence on every other program by predicting how the others change when that program is removed, yielding attention weights that reflect asymmetric influence rather than symmetric co-occurrence. Applied to 191,890 prefrontal cortex nuclei across three pathway vocabularies, ORBIT recovers co-activation structure consistent with established Alzheimer's disease vulnerability signatures, identifies cell-type-specific rewiring invisible to differential expression, and achieves 0.984 macro F1 on cell-type classification from 220 pathway scores, which is within 0.3 points of a state-of-the-art classifier using all 22,088 genes.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces ORBIT, a self-supervised transformer that learns asymmetric dependencies among gene programs from observational scRNA-seq data alone. Its core mechanism is an intervention-consistent training objective that predicts downstream program changes after simulated removal of a program, with the resulting attention weights interpreted as directional influences. Applied to 191,890 prefrontal cortex nuclei and three pathway vocabularies, the work claims to recover co-activation structure consistent with established Alzheimer's disease vulnerability signatures, detect cell-type-specific rewiring invisible to differential expression, and achieve 0.984 macro F1 on cell-type classification from 220 pathway scores (within 0.3 points of a full-gene SOTA classifier).
Significance. If the directional-influence interpretation is substantiated, ORBIT would provide a practical route to infer cell-type-stratified pathway rewiring from observational data without perturbation experiments, addressing a clear gap in single-cell methods. The reported classification performance and post-hoc consistency with known AD signatures demonstrate immediate utility for dimensionality reduction and signature recovery; these strengths would be strengthened by explicit ablation and baseline comparisons.
major comments (2)
- [Methods (intervention-consistent training objective)] The intervention-consistent objective defines directional influence directly by the model's ability to predict removal effects (see abstract and methods description of the training loss). This creates a circularity risk: any predictive model of co-expression asymmetries can satisfy the objective without capturing true biological directionality, and no perturbation experiments are performed to test the learned directions against ground-truth interventions.
- [Results (Alzheimer's disease analysis and cell-type classification)] The claim that ORBIT identifies 'true directional biological influences' and recovers AD vulnerability signatures rests on post-hoc consistency checks rather than independent validation. The skeptic note correctly identifies that statistical asymmetries (cell-type confounders, technical artifacts, or symmetric correlations) can produce the same attention patterns; an ablation removing the intervention term or comparing against symmetric baselines would be required to isolate the contribution of the directional mechanism.
minor comments (2)
- [Abstract] The abstract states the classification result is 'within 0.3 points' of SOTA but does not report the exact baseline F1 value or the precise number of genes used in the comparator; adding this number would improve clarity.
- [Methods] The manuscript mentions 'three pathway vocabularies' but does not tabulate their sizes or overlap; a supplementary table listing the vocabularies and the 220 selected pathways would aid reproducibility.
Simulated Author's Rebuttal
We thank the referee for the constructive comments on the intervention-consistent objective and the need for stronger validation of directional influences. We address each point below, agree that ablations will strengthen the manuscript, and will incorporate them in revision.
read point-by-point responses
-
Referee: [Methods (intervention-consistent training objective)] The intervention-consistent objective defines directional influence directly by the model's ability to predict removal effects (see abstract and methods description of the training loss). This creates a circularity risk: any predictive model of co-expression asymmetries can satisfy the objective without capturing true biological directionality, and no perturbation experiments are performed to test the learned directions against ground-truth interventions.
Authors: We acknowledge the concern about circularity. The objective requires not only predicting co-expression asymmetries but specifically simulating program removal and requiring accurate prediction of downstream effects; this interventional consistency distinguishes it from generic predictive models of symmetric correlations. We agree that direct perturbation validation would provide stronger evidence, but ORBIT is designed for observational scRNA-seq where such experiments are unavailable. We will add a limitations discussion and theoretical justification in the revised manuscript. revision: partial
-
Referee: [Results (Alzheimer's disease analysis and cell-type classification)] The claim that ORBIT identifies 'true directional biological influences' and recovers AD vulnerability signatures rests on post-hoc consistency checks rather than independent validation. The skeptic note correctly identifies that statistical asymmetries (cell-type confounders, technical artifacts, or symmetric correlations) can produce the same attention patterns; an ablation removing the intervention term or comparing against symmetric baselines would be required to isolate the contribution of the directional mechanism.
Authors: We agree that explicit ablations are needed to isolate the directional mechanism's contribution. In the revision we will add: (1) an ablation training without the intervention term using only standard self-supervision, and (2) comparisons to symmetric baselines including Pearson correlation networks and a standard transformer without intervention simulation. These will quantify performance and biological consistency gains from the directional objective. We will also revise language to describe 'inferred directional influences consistent with intervention semantics' rather than 'true directional biological influences'. revision: yes
- Direct experimental validation of learned directional influences against ground-truth perturbation data, as the method targets observational scRNA-seq and new wet-lab interventions are outside the current study's scope.
Circularity Check
Intervention-consistent objective defines directional influence by construction via removal prediction fit
specific steps
-
fitted input called prediction
[Abstract]
"the model learns each program's directional influence on every other program by predicting how the others change when that program is removed, yielding attention weights that reflect asymmetric influence rather than symmetric co-occurrence"
Directional influence is defined exactly as the model's ability to predict removal effects during self-supervised training on observational data; the attention weights are the fitted parameters that enable this prediction, so claiming they quantify true asymmetric program influences reduces to the training objective by construction without independent perturbation validation.
full rationale
The paper's core mechanism trains a transformer self-supervised on observational scRNA-seq counts to predict downstream program changes after simulated removal of a program; the resulting attention weights are then presented as asymmetric biological influences. This reduces the claimed 'directional influence' to the fitted parameters of the training objective itself (no external perturbations are performed to validate directionality). While downstream applications such as AD signature consistency and cell-type classification from pathway scores provide independent checks, the load-bearing claim that ORBIT recovers true cell-type-specific rewiring invisible to differential expression rests on the self-supervised fit. This matches the fitted-input-called-prediction pattern with partial circularity; the derivation is otherwise self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
free parameters (1)
- transformer architecture hyperparameters
axioms (1)
- domain assumption Observational single-cell RNA-seq data contains enough statistical structure for a removal-prediction objective to recover true directional biological influences.
invented entities (1)
-
intervention-consistent training objective
no independent evidence
Reference graph
Works this paper leans on
-
[1]
Melina A. Agosto and Theodore G. Wensel. Lrrtm4 is a member of the transsynaptic complex between rod photoreceptors and bipolar cells. Journal of Comparative Neurology, 529, 2021. doi:10.1002/cne.24944
-
[2]
SCENIC: single-cell regulatory network inference and clustering
Sara Aibar, Carmen Bravo Gonz\'alez-Blas, Thomas Moerman, V\^an Anh Huynh-Thu, Hana Imrichova, Gert Hulselmans, Florian Rambow, Jean-Christophe Marine, Pierre Geurts, Jan Aerts, Joost van den Oord, Zeynep Kalender Atak, Jasper Wouters, and Stein Aerts. SCENIC: single-cell regulatory network inference and clustering. Nature Methods, 14:1083--1086, 2017. do...
-
[3]
Glycosaminoglycans: Mechanisms and therapeutic potential in neurological diseases: A mini-review
Tahir Ali, Iram Murtaza, Hongling Guo, and Shupeng Li. Glycosaminoglycans: Mechanisms and therapeutic potential in neurological diseases: A mini-review. Biochemical and Biophysical Research Communications, 765, 2025. doi:10.1016/j.bbrc.2025.151861
-
[4]
Brandon J. Aubrey, Gemma L. Kelly, Ana Janic, Marco J. Herold, and Andreas Strasser. How does p53 induce apoptosis and how does this relate to p53-mediated tumour suppression? Cell Death and Differentiation, 25:104--113, 2018. doi:10.1038/cdd.2017.169
-
[5]
Asli Ayka and Ahmet \"O. S ehirli. The role of the SLC transporters protein in the neurodegenerative disorders. Clinical Psychopharmacology and Neuroscience, 18:174--187, 2020. doi:10.9758/cpn.2020.18.2.174
-
[6]
Alfonso E. Bencomo-Alvarez, Andres J. Rubio, Idaly M. Olivas, Mayra A. Gonzalez, Rebecca Ellwood, Carme Ripoll Fiol, Christopher A. Eide, Joshua J. Lara, Christian Barreto-Vargas, Luis F. Jave-Suarez, Georgios Nteliopoulos, Alistair G. Reid, Dragana Milojkovic, Brian J. Druker, Jane Apperley, Jamshid S. Khorashad, and Anna M. Eiring. Proteasome 26S subuni...
-
[7]
Courtney Bull, Ryan M. Byrne, Natalie C. Fisher, Shania M. Corry, Raheleh Amirkhah, Jessica Edwards, Lily V. S. Hillson, Mark Lawler, Aideen E. Ryan, Felicity Lamrock, Philip D. Dunne, and Sudhir B. Malla. Dual gene set enrichment analysis (dualGSEA); an R function that enables more robust biological discovery and pre-clinical model alignment from transcr...
-
[8]
Feed-forward inhibition in the hippocampal formation
Gyorgy Buzsaki. Feed-forward inhibition in the hippocampal formation. Progress in Neurobiology, 22:131--153, 1984. doi:10.1016/0301-0082(84)90023-6
-
[9]
Connor, Jeffrey Petrozzino, Lucas D
John A. Connor, Jeffrey Petrozzino, Lucas D. Pozzo-Miller, and Satoru Otani. Calcium signals in long-term potentiation and long-term depression. Canadian Journal of Physiology and Pharmacology, 77(9):722--734, 1999. doi:10.1139/y99-065
-
[10]
scGPT: toward building a foundation model for single-cell multi-omics using generative AI
Haotian Cui, Chloe Wang, Hassaan Maan, Kuan Pang, Fengning Luo, Nan Duan, and Bo Wang. scGPT: toward building a foundation model for single-cell multi-omics using generative AI. Nature Methods, 21:1470--1480, 2024. doi:10.1038/s41592-024-02201-0
-
[11]
Colin J. Dinsmore and Philippe Soriano. MAPK and PI3K signaling: At the crossroads of neural crest development. Developmental Biology, 444, 2018. doi:10.1016/j.ydbio.2018.02.003
-
[12]
Cecilia Dom\'inguez Conde, Chuan Xu, Lorna B. Jarvis, Dan B. Rainbow, Steven B. Wells, Tom\'as Gomes, Sarah K. Howlett, Ondrej Suchanek, Krzysztof Polanski, Hamish W. King, Lira Mamanova, Ni Huang, Peter A. Szabo, Lucy Richardson, Liam Bolt, Eirini S. Fasouli, Krishnaa T. Mahbubani, Martin Prete, Liz Tuck, Nathan Richoz, et al. Cross-tissue immune cell an...
-
[13]
Transcriptome programs involved in the development and structure of the cerebellum
Donatella Farini, Daniela Marazziti, Maria Concetta Geloso, and Claudio Sette. Transcriptome programs involved in the development and structure of the cerebellum. Cellular and Molecular Life Sciences, 78, 2021. doi:10.1007/s00018-021-03911-w
-
[14]
Catherine Fleitas, Pau Marfull-Orom\'i, Disha Chauhan, Daniel del Toro, Blanca Peguera, Bahira Zammou, Daniel Rocandio, R\"udiger Klein, Carme Espinet, and Joaquim Egea. FLRT2 and FLRT3 cooperate in maintaining the tangential migratory streams of cortical interneurons during development. Journal of Neuroscience, 41, 2021. doi:10.1523/JNEUROSCI.0380-20.2021
-
[15]
Combinatorial prediction of therapeutic perturbations using causally inspired neural networks
Guadalupe Gonzalez, Xiang Lin, Isuru Herath, Kirill Veselkov, Michael Bronstein, and Marinka Zitnik. Combinatorial prediction of therapeutic perturbations using causally inspired neural networks. Nature Biomedical Engineering, 2025. doi:10.1038/s41551-025-01481-x
-
[16]
Marshall, Fei Chen, Feng Zhang, Tommy Kaplan, Aviv Regev, and Michal Schwartz
Naomi Habib, Cristin McCabe, Sedi Medina, Miriam Varshavsky, Daniel Kitsberg, Raz Dvir-Szternfeld, Gilad Green, Danielle Dionne, Lan Nguyen, Jamie L. Marshall, Fei Chen, Feng Zhang, Tommy Kaplan, Aviv Regev, and Michal Schwartz. Disease-associated astrocytes in Alzheimer's disease and aging. Nature Neuroscience, 23:701--706, 2020. doi:10.1038/s41593-020-0624-8
-
[18]
Chen Jin, Jiaoni Wang, Yumeng Wang, Bojun Jia, Xuefei Guo, Guanghui Yang, Peng Xu, Paul Greengard, Rui Zhou, and Yigong Shi. Modulation of amyloid precursor protein cleavage by -secretase activating protein through phase separation. Proceedings of the National Academy of Sciences, 119, 2022. doi:10.1073/pnas.2122292119
-
[19]
Gillespie, Imre Vastrik, Peter D'Eustachio, Eric W
Geeta Joshi-Tope, Matthew T. Gillespie, Imre Vastrik, Peter D'Eustachio, Eric W. Schmidt, Bernard de Bono, Bijay Jassal, Gopal R. Gopinath, Gao-Rong Wu, Lucy M. Matthews, Simon L. Lewis, Ewan Birney, and Lincoln Stein. Reactome: a knowledgebase of biological pathways. Nucleic Acids Research, 33, 2005. doi:10.1093/nar/gki072
-
[20]
KEGG: Kyoto Encyclopedia of Genes and Genomes
Minoru Kanehisa and Susumu Goto. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Research, 28:27--30, 2000. doi:10.1093/nar/28.1.27
-
[21]
Causal analysis approaches in ingenuity pathway analysis
Andreas Kr\"amer, Jeff Green, Jack Pollard, and Stuart Tugendreich. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics, 30:523--530, 2013. doi:10.1093/bioinformatics/btt703
-
[22]
Shun-Fat Lau, Han Cao, Amy K. Y. Fu, and Nancy Y. Ip. Single-nucleus transcriptome analysis reveals dysregulation of angiogenic endothelial cells and neuroprotective glia in Alzheimer's disease. Proceedings of the National Academy of Sciences, 117:25800--25809, 2020. doi:10.1073/pnas.2008762117
-
[23]
Zs\'ofia I. Laszl\'o and Zsolt Lele. Flying under the radar: Cdh2 (N-cadherin), an important hub molecule in neurodevelopmental and neurodegenerative diseases. Frontiers in Neuroscience, 16, 2022. doi:10.3389/fnins.2022.972059
-
[24]
Kun Leng, Emmy Li, Rana Eser, Antonia Piergies, Rene Sit, Michelle Tan, Norma Neff, Song Hua Li, Roberta Diehl Rodriguez, Claudia Kimie Suemoto, Renata Elaine Paraizo Leite, Alexander J. Ehrenberg, Carlos A. Pasqualucci, William W. Seeley, Salvatore Spina, Helmut Heinsen, Lea T. Grinberg, and Martin Kampmann. Molecular characterization of selectively vuln...
-
[25]
Pei-Yi Lin, Lulu Y. Chen, Man Jiang, Justin H. Trotter, Erica Seigneur, and Thomas C. S\"udhof. Neurexin-2: An inhibitory neurexin that restricts excitatory synapse formation in the hippocampus. Science Advances, 9, 2023. doi:10.1126/sciadv.add8856
-
[26]
Boix, Leyla Anne Akay, Ziting Xia, Jose Davila-Velderrain, Ayesha P
Hansruedi Mathys, Carles A. Boix, Leyla Anne Akay, Ziting Xia, Jose Davila-Velderrain, Ayesha P. Ng, Xueqiao Jiang, Ghada Abdelhady, Kyriaki Galani, Julio Mantero, Neil Band, Benjamin T. James, Sudhagar Babu, Fabiola Galiana-Melendez, Kate Louderback, Dmitry Prokopenko, Rudolph E. Tanzi, David A. Bennett, Li-Huei Tsai, and Manolis Kellis. Single-cell mult...
-
[27]
Bril, Ana Archvadze, Madalena B
Almudena Medina-Samam\'e, \"Eva Paller, Mateo R. Bril, Ana Archvadze, Madalena B. C. Sim\ oes-Abade, Patricia Esta\ nol-Cayuela, and Chloe LeMaoult. Role of neurexins in Alzheimer's disease. The Journal of Neuroscience, 43, 2023. doi:10.1523/jneurosci.0169-23.2023
-
[28]
Alzheimer's disease rewires gene coexpression networks coupling different brain regions
Sanga Mitra, Kailash BP, Srivatsan C R, Naga Venkata Saikumar, Philge Philip, and Manikandan Narayanan. Alzheimer's disease rewires gene coexpression networks coupling different brain regions. NPJ Systems Biology and Applications, 10(50), 2024. doi:10.1038/s41540-024-00376-y
-
[29]
Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer's disease
Samuel Morabito, Emily Miyoshi, Neethu Michael, Saba Shahin, Alessandra Cadete Martini, Elizabeth Head, Justine Silva, Kelsey Leavy, Mari Perez-Rosendahl, and Vivek Swarup. Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer's disease. Nature Genetics, 53:1143--1155, 2021. doi:10.1038/s41588-021-00894-z
-
[30]
Reproducibility of methods to detect differentially expressed genes from single-cell RNA sequencing
Tian Mou, Wenjiang Deng, Fengyun Gu, Yudi Pawitan, and Trung Nghia Vu. Reproducibility of methods to detect differentially expressed genes from single-cell RNA sequencing. Frontiers in Genetics, 10:1331, 2019. doi:10.3389/fgene.2019.01331
-
[31]
Elliott J. Mufson, Scott E. Counts, Sylvia E. Perez, and Stephen D. Ginsberg. Cholinergic system during the progression of Alzheimer's disease: therapeutic implications. Expert Review of Neurotherapeutics, 8, 2008. doi:10.1586/14737175.8.11.1703
-
[32]
Marcos Otero-Garcia, Sameehan U. Mahajani, Debia Wakhloo, Weijing Tang, Yue-Qiang Xue, Samuel Morabito, Jie Pan, Jane Oberhauser, Angela E. Madira, Tamara Shakouri, Yongning Deng, Thomas Allison, Zihuai He, William E. Lowry, Riki Kawaguchi, Vivek Swarup, and Inma Cobos. Molecular signatures underlying neurofibrillary tangle susceptibility in Alzheimer's d...
-
[33]
Kaminga, Shi Wu Wen, Xinyin Wu, Kwabena Acheampong, and Aizhong Liu
Xiongfeng Pan, Atipatsa C. Kaminga, Shi Wu Wen, Xinyin Wu, Kwabena Acheampong, and Aizhong Liu. Dopamine and dopamine receptors in Alzheimer's disease: A systematic review and network meta-analysis. Frontiers in Aging Neuroscience, 11, 2019. doi:10.3389/fnagi.2019.00175
-
[34]
Pandi-Perumal, Ilya Trakht, Srinivasan, D
Seithikurippu R. Pandi-Perumal, Ilya Trakht, Srinivasan, D. Warren Spence, Georges J. M. Maestroni, Nava Zisapel, and Daniel P. Cardinali. Physiological effects of melatonin: Role of melatonin receptors and signal transduction pathways. Progress in Neurobiology, 85:335--353, 2008. doi:10.1016/j.pneurobio.2008.04.001
-
[35]
Predicting transcriptional outcomes of novel multigene perturbations with GEARS
Yusuf Roohani, Kexin Huang, and Jure Leskovec. Predicting transcriptional outcomes of novel multigene perturbations with GEARS. Nature Biotechnology, 42:927--935, 2024. doi:10.1038/s41587-023-01905-6
-
[36]
Luisa Seguella and Brian D. Gulbransen. Enteric glial biology, intercellular signalling and roles in gastrointestinal disease. Nature Reviews Gastroenterology & Hepatology, 18:571--587, 2021. doi:10.1038/s41575-021-00423-7
-
[37]
Elaine H. Shen, Caroline C. Overly, and Allan R. Jones. The Allen Human Brain Atlas: comprehensive gene expression mapping of the human brain. Trends in Neurosciences, 35:711--714, 2012. doi:10.1016/j.tins.2012.09.005
-
[38]
Kai Silkwood, Emmanuel Dollinger, Joshua Gervin, Scott Atwood, Qing Nie, and Arthur D. Lander. Leveraging gene correlations in single cell transcriptomic data. BMC Bioinformatics, 25, 2024. doi:10.1186/s12859-024-05926-z
-
[39]
Mootha, Sayan Mukherjee, Benjamin L
Aravind Subramanian, Pablo Tamayo, Vamsi K. Mootha, Sayan Mukherjee, Benjamin L. Ebert, Michael A. Gillette, Amanda Paulovich, Scott L. Pomeroy, Todd R. Golub, Eric S. Lander, and Jill P. Mesirov. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences, 102(...
-
[40]
Priyanka Dey Talukdar and Urmi Chatterji. Transcriptional co-activators: emerging roles in signaling pathways and potential therapeutic targets for diseases. Signal Transduction and Targeted Therapy, 8, 2023. doi:10.1038/s41392-023-01651-w
-
[41]
Christina V. Theodoris, Ling Xiao, Anant Chopra, Mark D. Chaffin, Zeina R. Al Sayed, Matthew C. Hill, Helene Mantineo, Elizabeth M. Brydon, Zexian Zeng, X. Shirley Liu, and Patrick T. Ellinor. Transfer learning enables predictions in network biology. Nature, 618:616--624, 2023. doi:10.1038/s41586-023-06139-9
-
[42]
Michael Wainberg, Shea J. Andrews, and Shreejoy J. Tripathy. Shared genetic risk loci between Alzheimer's disease and related dementias, Parkinson's disease, and amyotrophic lateral sclerosis. Alzheimer's Research & Therapy, 15, 2023. doi:10.1186/s13195-023-01244-3
-
[44]
Martijn C. Wilde, Cassia R. Overk, John W. Sijben, and Eliezer Masliah. Meta-analysis of synaptic pathology in Alzheimer's disease reveals selective molecular vesicular machinery vulnerability. Alzheimer's & Dementia, 12:633--644, 2016. doi:10.1016/j.jalz.2015.12.005
-
[45]
Chenling Xu, Romain Lopez, Edouard Mehlman, Jeffrey Regier, Michael I. Jordan, and Nir Yosef. Probabilistic harmonization and annotation of single-cell transcriptomics data with deep generative models. Molecular Systems Biology, 17, 2021. doi:10.15252/msb.20209620
-
[46]
GABAergic inhibitory interneuron deficits in Alzheimer's disease: Implications for treatment
Yilan Xu, Manna Zhao, Yuying Han, and Heng Zhang. GABAergic inhibitory interneuron deficits in Alzheimer's disease: Implications for treatment. Frontiers in Neuroscience, 14, 2020. doi:10.3389/fnins.2020.00660
-
[47]
Yupu Xu, Yuzhou Wang, and Shisong Ma. SingleCellGGM enables gene expression program identification from single-cell transcriptomes and facilitates universal cell label transfer. Cell Reports Methods, 4, 2024. doi:10.1016/j.crmeth.2024.100813
-
[48]
Fan Yang, Wenchuan Wang, Fang Wang, Yuan Fang, Duyu Tang, Junzhou Huang, Hui Lu, and Jianhua Yao. scBERT as a large-scale pretrained deep language model for cell type annotation of single-cell RNA-seq data. Nature Machine Intelligence, 4:852--866, 2022. doi:10.1038/s42256-022-00534-z
-
[49]
Reproducibility of methods to detect differentially expressed genes from single-cell
Mou, Tian and Deng, Wenjiang and Gu, Fengyun and Pawitan, Yudi and Vu, Trung Nghia , journal =. Reproducibility of methods to detect differentially expressed genes from single-cell. 2019 , doi =
2019
-
[50]
Proceedings of the National Academy of Sciences , volume =
Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles , author =. Proceedings of the National Academy of Sciences , volume =. 2005 , doi =
2005
-
[51]
and Fisher, Natalie C
Bull, Courtney and Byrne, Ryan M. and Fisher, Natalie C. and Corry, Shania M. and Amirkhah, Raheleh and Edwards, Jessica and Hillson, Lily V. S. and Lawler, Mark and Ryan, Aideen E. and Lamrock, Felicity and Dunne, Philip D. and Malla, Sudhir B. , journal =. Dual gene set enrichment analysis (. 2024 , doi =
2024
-
[52]
2024 , doi =
Xu, Yupu and Wang, Yuzhou and Ma, Shisong , journal =. 2024 , doi =
2024
-
[53]
BMC Bioinformatics , volume =
Leveraging gene correlations in single cell transcriptomic data , author=. BMC Bioinformatics , volume =. 2024 , doi =
2024
-
[54]
and Pasqualucci, Carlos A
Leng, Kun and Li, Emmy and Eser, Rana and Piergies, Antonia and Sit, Rene and Tan, Michelle and Neff, Norma and Li, Song Hua and Rodriguez, Roberta Diehl and Suemoto, Claudia Kimie and Leite, Renata Elaine Paraizo and Ehrenberg, Alexander J. and Pasqualucci, Carlos A. and Seeley, William W. and Spina, Salvatore and Heinsen, Helmut and Grinberg, Lea T. and...
2021
-
[55]
Progress in Neurobiology , volume =
Buzsaki, Gyorgy , title =. Progress in Neurobiology , volume =. 1984 , doi =
1984
-
[56]
and Eysert, F
Mary, A. and Eysert, F. and Checler, F. and Chami, M. , title =. Molecular Psychiatry , volume =. 2022 , doi =
2022
-
[57]
Biochemical and Biophysical Research Communications , volume =
Ali, Tahir and Murtaza, Iram and Guo, Hongling and Li, Shupeng , title =. Biochemical and Biophysical Research Communications , volume =. 2025 , doi =
2025
-
[58]
and Mecca, C
Arcuri, C. and Mecca, C. and Bianchi, R. and Giambanco, I. and Donato, R. , title =. Frontiers in Molecular Neuroscience , volume =. 2017 , doi =
2017
-
[59]
Lau, Shun-Fat and Cao, Han and Fu, Amy K. Y. and Ip, Nancy Y. , title =. Proceedings of the National Academy of Sciences , volume =. 2020 , doi =
2020
-
[60]
and Wakhloo, Debia and Tang, Weijing and Xue, Yue-Qiang and Morabito, Samuel and Pan, Jie and Oberhauser, Jane and Madira, Angela E
Otero-Garcia, Marcos and Mahajani, Sameehan U. and Wakhloo, Debia and Tang, Weijing and Xue, Yue-Qiang and Morabito, Samuel and Pan, Jie and Oberhauser, Jane and Madira, Angela E. and Shakouri, Tamara and Deng, Yongning and Allison, Thomas and He, Zihuai and Lowry, William E. and Kawaguchi, Riki and Swarup, Vivek and Cobos, Inma , title =. Neuron , volume...
2022
-
[61]
NPJ Systems Biology and Applications , volume =
Mitra, Sanga and Kailash BP and R, Srivatsan C and Naga Venkata Saikumar and Philip, Philge and Narayanan, Manikandan , title =. NPJ Systems Biology and Applications , volume =. 2024 , doi =
2024
-
[62]
Signal Transduction and Targeted Therapy , volume =
Talukdar, Priyanka Dey and Chatterji, Urmi , title =. Signal Transduction and Targeted Therapy , volume =. 2023 , doi =
2023
-
[63]
, title =
Seguella, Luisa and Gulbransen, Brian D. , title =. Nature Reviews Gastroenterology & Hepatology , volume =. 2021 , doi =
2021
-
[64]
and Akay, Leyla Anne and Xia, Ziting and Davila-Velderrain, Jose and Ng, Ayesha P
Mathys, Hansruedi and Boix, Carles A. and Akay, Leyla Anne and Xia, Ziting and Davila-Velderrain, Jose and Ng, Ayesha P. and Jiang, Xueqiao and Abdelhady, Ghada and Galani, Kyriaki and Mantero, Julio and Band, Neil and James, Benjamin T. and Babu, Sudhagar and Galiana-Melendez, Fabiola and Louderback, Kate and Prokopenko, Dmitry and Tanzi, Rudolph E. and ...
2024
-
[65]
International Conference on Learning Representations (ICLR) Poster , year=
Towards Best Practices of Activation Patching in Language Models: Metrics and Methods , author =. International Conference on Learning Representations (ICLR) Poster , year=
-
[66]
Frontiers in Neuroscience , volume =
Xu, Yilan and Zhao, Manna and Han, Yuying and Zhang, Heng , title =. Frontiers in Neuroscience , volume =. 2020 , doi =
2020
-
[67]
A ttention is not E xplanation
Jain, Sarthak and Wallace, Byron C. A ttention is not E xplanation. Proceedings of the 2019 Conference of the North A merican Chapter of the Association for Computational Linguistics: Human Language Technologies, Volume 1 (Long and Short Papers). 2019. doi:10.18653/v1/N19-1357
-
[68]
Attention is not not explanation
Wiegreffe, Sarah and Pinter, Yuval. Attention is not not Explanation. Proceedings of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP). 2019. doi:10.18653/v1/D19-1002
-
[69]
Bioinformatics , volume =
Krämer, Andreas and Green, Jeff and Pollard, Jack and Tugendreich, Stuart , title =. Bioinformatics , volume =. 2013 , doi =
2013
-
[70]
Canadian Journal of Physiology and Pharmacology , volume =
Connor, John A and Petrozzino, Jeffrey and Pozzo-Miller, Lucas D and Otani, Satoru , title =. Canadian Journal of Physiology and Pharmacology , volume =. 1999 , doi =
1999
-
[71]
Predicting transcriptional outcomes of novel multigene perturbations with
Roohani, Yusuf and Huang, Kexin and Leskovec, Jure , journal =. Predicting transcriptional outcomes of novel multigene perturbations with. 2024 , doi =
2024
-
[72]
Nature Biomedical Engineering , year =
Combinatorial prediction of therapeutic perturbations using causally inspired neural networks , author=. Nature Biomedical Engineering , year =
-
[73]
2024 , doi =
Cui, Haotian and Wang, Chloe and Maan, Hassaan and Pang, Kuan and Luo, Fengning and Duan, Nan and Wang, Bo , journal =. 2024 , doi =
2024
-
[74]
Science , volume =
Cross-tissue immune cell analysis reveals tissue-specific features in humans , author=. Science , volume =. 2022 , doi =
2022
-
[75]
2022 , doi =
Yang, Fan and Wang, Wenchuan and Wang, Fang and Fang, Yuan and Tang, Duyu and Huang, Junzhou and Lu, Hui and Yao, Jianhua , journal =. 2022 , doi =
2022
-
[76]
Nature , volume =
Transfer learning enables predictions in network biology , author=. Nature , volume =. 2023 , doi =
2023
-
[77]
Molecular Systems Biology , volume =
Xu, Chenling and Lopez, Romain and Mehlman, Edouard and Regier, Jeffrey and Jordan, Michael I and Yosef, Nir , title =. Molecular Systems Biology , volume =. 2021 , doi =
2021
-
[78]
Single-nucleus chromatin accessibility and transcriptomic characterization of
Morabito, Samuel and Miyoshi, Emily and Michael, Neethu and Shahin, Saba and Alessandra Cadete Martini and Head, Elizabeth and Silva, Justine and Leavy, Kelsey and Perez-Rosendahl, Mari and Vivek Swarup , journal =. Single-nucleus chromatin accessibility and transcriptomic characterization of. 2021 , doi =
2021
-
[79]
Progress in Neurobiology , volume =
Physiological effects of melatonin: Role of melatonin receptors and signal transduction pathways , author =. Progress in Neurobiology , volume =. 2008 , doi =
2008
-
[80]
and Wen, Shi Wu and Wu, Xinyin and Acheampong, Kwabena and Liu, Aizhong , title =
Pan, Xiongfeng and Kaminga, Atipatsa C. and Wen, Shi Wu and Wu, Xinyin and Acheampong, Kwabena and Liu, Aizhong , title =. Frontiers in Aging Neuroscience , volume =. 2019 , doi =
2019
-
[81]
The role of the SLC transporters protein in the neurodegenerative disorders , journal =
Ayka, Asli and. The role of the SLC transporters protein in the neurodegenerative disorders , journal =. 2020 , doi =
2020
-
[82]
and Overk, Cassia R
Wilde, Martijn C. and Overk, Cassia R. and Sijben, John W. and Masliah, Eliezer , title =. Alzheimer's & Dementia , volume =. 2016 , doi =
2016
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.