Recognition: unknown
Targeted Disruption of Hypernetworks via Spectral Partitioning
Pith reviewed 2026-05-08 02:35 UTC · model grok-4.3
The pith
Spectral partitioning of hyperedge overlaps reduces contagion more than random removal only in Erdős–Rényi hypergraphs.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The central discovery is that the effectiveness of using multiscale cut-persistence from spectral k-way clustering on s-line graphs to target hyperedges for removal is strongly dependent on the seed graph topology. In Erdős–Rényi derived hypergraphs, this targeting reduces the final infection size more than random hyperedge removal. However, in Watts–Strogatz and Barabási–Albert cases, random removal performs comparably or better, indicating that identifying structurally salient hyperedges via spectral overlap does not always lead to optimal contagion suppression.
What carries the argument
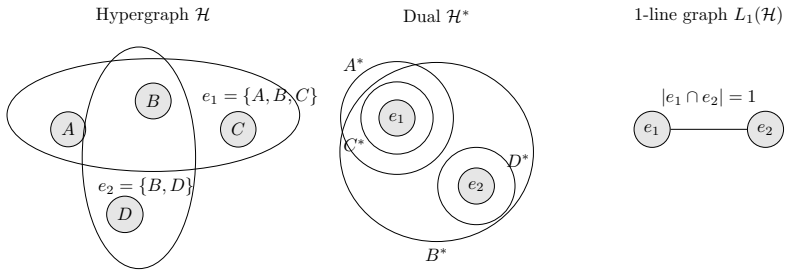
The multiscale cut-persistence score obtained from spectral k-way clustering of s-line graphs, which represent hyperedges as vertices connected if they overlap in at least s nodes, used to rank and remove hyperedges.
If this is right
- Targeted hyperedge removal based on spectral cuts can improve contagion control in certain random topologies but offers no advantage in small-world or scale-free ones.
- The structure of hyperedge overlaps can be analyzed at multiple scales using line graphs and clustering to identify important hyperedges.
- Random hyperedge removal remains a competitive baseline strategy across diverse hypergraph topologies.
- Further studies are needed to compare this approach against ensemble methods and more explicit higher-order contagion models.
Where Pith is reading between the lines
- Intervention designs for real hypernetworks should account for topology-specific responses rather than assuming uniform benefits from structural targeting.
- Testing the method on empirical hypergraphs from social or biological data could reveal whether the topology dependence holds outside synthetic cases.
- Integrating cut-persistence with other centrality measures might yield better performance across all topologies.
Load-bearing premise
The contagion dynamics observed on synthetic hypergraphs created by promoting cliques from standard graphs represent those of higher-order contagion processes on actual hypernetworks.
What would settle it
A simulation or observation in an Erdős–Rényi hypergraph where cut-persistence targeting fails to reduce final infection size below that achieved by random removal, or where random removal outperforms it consistently.
Figures






read the original abstract
We study hyperedge-removal strategies for suppressing contagion on synthetic hypergraphs. Hypergraphs are generated from Erd\H{o}s--R\'enyi, Barab\'asi--Albert, and Watts--Strogatz seed graphs by promoting maximal cliques to hyperedges. For each hypergraph, we construct \(s\)-line graphs whose vertices correspond to hyperedges and whose edges encode hyperedge overlap of size at least \(s\). Spectral \(k\)-way clustering of these \(s\)-line graphs yields a multiscale cut-persistence score used to rank hyperedges for removal. Simulations show that the effect of this intervention is strongly topology-dependent. In the reported Erd\H{o}s--R\'enyi case, cut-persistence targeting reduces final infection size more than random hyperedge removal. In the Watts--Strogatz and Barab\'asi--Albert cases, however, random removal is comparable to or better than cut-persistence targeting. These results suggest that spectral overlap structure can identify structurally salient hyperedges, but structural salience alone does not guarantee optimal contagion suppression. The study motivates further comparison with ensemble-level experiments and explicitly higher-order contagion models.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces a spectral method for ranking hyperedges for removal to suppress contagion on hypergraphs. Hypergraphs are generated synthetically by promoting maximal cliques from Erdős–Rényi, Barabási–Albert, and Watts–Strogatz seed graphs to hyperedges; s-line graphs are then built on overlaps of size at least s, and spectral k-way clustering produces a multiscale cut-persistence score for targeting. Simulations on these hypergraphs show topology-dependent outcomes: cut-persistence targeting reduces final infection size more than random removal in the ER case, while random removal is comparable or superior in the WS and BA cases. The authors conclude that structural salience identified by spectral overlap does not guarantee optimal dynamical suppression.
Significance. If the empirical reversal holds under scrutiny, the work usefully illustrates the limits of purely structural targeting in higher-order contagion models and motivates ensemble comparisons. The explicit construction of s-line graphs and multiscale persistence scoring is a clear methodological contribution, and the use of three distinct seed topologies provides a controlled test of dependence on underlying structure.
major comments (2)
- [Hypergraph Generation] Hypergraph Generation section: the clique-promotion procedure forces every hyperedge to be a complete subgraph of the seed graph, which systematically correlates overlaps in a manner absent from many empirical hypernetworks (e.g., co-authorship or metabolic data). This construction artifact could produce the reported topology-dependent reversal as an artifact of the generator rather than a general property of hypergraph contagion; the central claim that “structural salience alone does not guarantee optimal contagion suppression” therefore requires validation on at least one alternative generator or real hypernetwork.
- [Simulation Results] Simulation Results section: the headline comparison (cut-persistence superior in ER; random comparable or better in WS/BA) rests on contagion simulations whose parameters (infection/recovery rates, initial seed size, number of runs, and error bars) are not reported with sufficient precision to reproduce the reversal. Without these details the topology dependence cannot be verified and remains load-bearing for the paper’s empirical conclusion.
minor comments (2)
- [Abstract] Abstract: the phrase “in the reported Erdős–Rényi case” is ambiguous without stating the specific ER parameters (p, n) used; adding these values would improve clarity.
- [Methods] Notation: the definition of the cut-persistence score is introduced without an explicit equation number; labeling it as Eq. (X) in the methods would aid cross-reference.
Simulated Author's Rebuttal
We thank the referee for the constructive comments, which highlight important aspects of generalizability and reproducibility. We respond to each major comment below and indicate the revisions we will make.
read point-by-point responses
-
Referee: Hypergraph Generation section: the clique-promotion procedure forces every hyperedge to be a complete subgraph of the seed graph, which systematically correlates overlaps in a manner absent from many empirical hypernetworks (e.g., co-authorship or metabolic data). This construction artifact could produce the reported topology-dependent reversal as an artifact of the generator rather than a general property of hypergraph contagion; the central claim that “structural salience alone does not guarantee optimal contagion suppression” therefore requires validation on at least one alternative generator or real hypernetwork.
Authors: We appreciate the referee's observation on the limitations of the clique-promotion generator. This method was selected specifically to produce hypergraphs whose higher-order structure directly inherits the topological properties of the seed graphs (ER, BA, WS), enabling a controlled test of how underlying connectivity affects spectral targeting efficacy. The resulting topology-dependent reversal demonstrates that structural salience identified via s-line graphs does not uniformly translate to dynamical advantage, which is consistent with the concern about generalizability. In the revised manuscript we will expand the Hypergraph Generation and Discussion sections with an explicit caveat on this construction choice, note that overlap correlations may differ in empirical data, and recommend validation against alternative generators (e.g., configuration-model hypergraphs) and real hypernetworks as a priority for follow-up work. We do not claim universality from the present synthetic results. revision: partial
-
Referee: Simulation Results section: the headline comparison (cut-persistence superior in ER; random comparable or better in WS/BA) rests on contagion simulations whose parameters (infection/recovery rates, initial seed size, number of runs, and error bars) are not reported with sufficient precision to reproduce the reversal. Without these details the topology dependence cannot be verified and remains load-bearing for the paper’s empirical conclusion.
Authors: We agree that the simulation parameters require fuller specification to support reproducibility. In the revised manuscript we will add a dedicated paragraph (and an appendix table) detailing the SIR contagion parameters: infection rate β, recovery rate γ, initial seed fraction, number of independent Monte Carlo realizations, and the precise definition of final infection size. All figures will be updated to display error bars (standard deviation across runs) and the text will state the exact numerical values used for each topology. revision: yes
Circularity Check
No circularity: empirical simulation comparison on generated hypergraphs
full rationale
The paper's workflow consists of (1) generating synthetic hypergraphs by promoting maximal cliques of ER/BA/WS seed graphs to hyperedges, (2) constructing s-line graphs on hyperedge overlaps, (3) performing spectral k-way clustering to compute a cut-persistence ranking, and (4) running contagion simulations to compare targeted removal against random removal. All reported outcomes (e.g., ER case shows targeting reduces infection size more than random; WS/BA cases show random comparable or superior) are direct numerical results of these simulations. No equation reduces a claimed prediction to a fitted input by construction, no uniqueness theorem is imported via self-citation, and no ansatz is smuggled. The derivation chain is therefore self-contained and externally falsifiable against the generated data and simulation runs.
Axiom & Free-Parameter Ledger
free parameters (2)
- overlap threshold s
- number of clusters k
axioms (2)
- domain assumption Spectral clustering on the s-line graph yields a meaningful multiscale cut-persistence score for hyperedge importance.
- domain assumption Clique promotion from ER/BA/WS graphs produces representative hypergraphs for contagion studies.
Reference graph
Works this paper leans on
-
[1]
kneed: Knee-point detection in python, 2020
Kevin Arvai. kneed: Knee-point detection in python, 2020. Python package implementing the Kneedle algorithm
2020
-
[2]
Cambridge university press, 2016
Albert-L´ aszl´ o Barab´ asi.Network science. Cambridge university press, 2016
2016
-
[3]
Emergence of scaling in random networks.Science, 286(5439):509–512, 1999
Albert-L´ aszl´ o Barab´ asi and R´ eka Albert. Emergence of scaling in random networks.Science, 286(5439):509–512, 1999
1999
-
[4]
Networks beyond pairwise interactions: structure and dynamics.Physics Reports, 874:1–92, 2020
Federico Battiston, Giulia Cencetti, Iacopo Iacopini, Vito Latora, Maxime Lucas, Alice Patania, Jean-Gabriel Young, and Giovanni Petri. Networks beyond pairwise interactions: structure and dynamics.Physics Reports, 874:1–92, 2020
2020
-
[5]
Benson, Rediet Abebe, Michael T
Austin R. Benson, Rediet Abebe, Michael T. Schaub, Ali Jadbabaie, and Jon Kleinberg. Simplicial closure and higher-order link prediction.Proceedings of the National Academy of Sciences, 115(48):E11221–E11230, 2018. 14
2018
-
[6]
Higher-order organization of complex networks.Science, 353(6295):163–166, 2016
Austin R Benson, David F Gleich, and Jure Leskovec. Higher-order organization of complex networks.Science, 353(6295):163–166, 2016
2016
-
[7]
Random walks on hypergraphs.Phys
Timoteo Carletti, Federico Battiston, Giulia Cencetti, and Duccio Fanelli. Random walks on hypergraphs.Phys. Rev. E, 101:022308, Feb 2020
2020
-
[8]
Hubert Chan, Anand Louis, Zhihao Gavin Tang, and Chenzi Zhang
T.-H. Hubert Chan, Anand Louis, Zhihao Gavin Tang, and Chenzi Zhang. Spectral properties of hypergraph laplacian and approximation algorithms.J. ACM, 65(3), March 2018
2018
-
[9]
Philip S. Chodrow. Configuration models of random hypergraphs.Journal of Complex Networks, 8(3):cnaa018, 2020
2020
-
[10]
On random graphs.Publicationes Mathematicae, 6:290–297, 1959
Paul Erd˝ os and Alfr´ ed R´ enyi. On random graphs.Publicationes Mathematicae, 6:290–297, 1959
1959
-
[11]
Rodr´ ıguez-Vel´ azquez
Ernesto Estrada and Juan A. Rodr´ ıguez-Vel´ azquez. Subgraph centrality in complex networks.Phys. Rev. E, 71:056103, May 2005
2005
-
[12]
Community detection in graphs.Physics Reports, 486(3–5):75–174, 2010
Santo Fortunato. Community detection in graphs.Physics Reports, 486(3–5):75–174, 2010
2010
-
[13]
Girvan and M
M. Girvan and M. E. J. Newman. Community structure in social and biological networks.Proceedings of the National Academy of Sciences, 99(12):7821–7826, 2002
2002
-
[14]
Hagberg, Daniel A
Aric A. Hagberg, Daniel A. Schult, and Pieter J. Swart. Exploring network structure, dynamics, and function using NetworkX. In Ga¨ el Varoquaux, Travis Vaught, and Jarrod Millman, editors,Proceedings of the 7th Python in Science Conference, pages 11–15, Pasadena, CA USA, 2008
2008
-
[15]
Hasenjager, Mark M
Matthew J. Hasenjager, Mark M. Bailey, and Nina H. Fefferman. Group composition influences diffusion dynamics via impacts on behavioural production.Animal Behaviour, 233:123482, 2026
2026
-
[16]
Hasenjager and Nina H
Matthew J. Hasenjager and Nina H. Fefferman. Social ageing and higher-order interactions: social selectiveness can enhance older individuals’ capacity to transmit knowledge.Philosophical Transactions of the Royal Society B: Biological Sciences, 379(1916):20220461, 2024
1916
-
[17]
Social science: Computational social science.Science, 323(5915):721–723, 2009
David Lazer, Alex Pentland, Lada Adamic, Sinan Aral, Albert-L´ aszl´ o Barab´ asi, Devon Brewer, Nicholas Christakis, Noshir Contractor, James Fowler, Myron Gutmann, Tony Jebara, Gary King, Michael Macy, Deb Roy, and Marshall Van Alstyne. Social science: Computational social science.Science, 323(5915):721–723, 2009
2009
-
[18]
Liu, Jesun Firoz, Sinan G
Xu T. Liu, Jesun Firoz, Sinan G. Aksoy, Ilya Amburg, Andrew Lumsdaine, Cliff Joslyn, Assefaw H. Gebremedhin, and Brenda Praggastis. High-order line graphs of non-uniform hypergraphs: Algorithms, applications, and experimental analysis. In2022 IEEE International Parallel and Distributed Processing Symposium (IPDPS), pages 852–862. IEEE, 2022
2022
-
[19]
Higher-order motif analysis in hypergraphs.Communications Physics, 5(1):79, 2022
Quintino Francesco Lotito, Federico Musciotto, Alberto Montresor, and Federico Battiston. Higher-order motif analysis in hypergraphs.Communications Physics, 5(1):79, 2022
2022
-
[20]
M. E. J. Newman. Modularity and community structure in networks.Proceedings of the National Academy of Sciences, 103(23):8577–8582, 2006
2006
-
[21]
On spectral clustering: Analysis and an algorithm
Andrew Y Ng, Michael I Jordan, and Yair Weiss. On spectral clustering: Analysis and an algorithm. InAdvances in neural information processing systems, pages 849–856, 2002
2002
-
[22]
Fast fragmentation of networks using module-based attacks.PLOS ONE, 10(11):e0142824, 2015
Bruno Requi˜ ao da Cunha, Juan Carlos Gonz´ alez-Avella, and Sebasti´ an Gon¸ calves. Fast fragmentation of networks using module-based attacks.PLOS ONE, 10(11):e0142824, 2015
2015
-
[23]
Shypar: A spectral coarsening approach to hypergraph partitioning, 2024
Hamed Sajadinia, Ali Aghdaei, and Zhuo Feng. Shypar: A spectral coarsening approach to hypergraph partitioning, 2024
2024
-
[24]
Finding a “kneedle” in a haystack: Detecting knee points in system behavior
Ville Satopaa, Jeannie Albrecht, David Irwin, and Barath Raghavan. Finding a “kneedle” in a haystack: Detecting knee points in system behavior. InProceedings of the 31st International Conference on Distributed Computing Systems Workshops, pages 166–171, 2011
2011
-
[25]
V. A. Traag, P. Van Dooren, and Y. Nesterov. Narrow scope for resolution-limit-free community detection.Phys. Rev. E, 84:016114, Jul 2011
2011
-
[26]
A tutorial on spectral clustering.Statistics and computing, 17(4):395–416, 2007
Ulrike Von Luxburg. A tutorial on spectral clustering.Statistics and computing, 17(4):395–416, 2007
2007
-
[27]
Small-World
Duncan J. Watts and Steven H. Strogatz. Collective dynamics of “Small-World” networks.Nature, 393(6684):440–442, 1998. 15
1998
-
[28]
Identifying critical edges in complex networks.Scientific Reports, 8(1):14469, 2018
En-Yu Yu, Duan-Bing Chen, and Jun-Yan Zhao. Identifying critical edges in complex networks.Scientific Reports, 8(1):14469, 2018
2018
-
[29]
Learning with hypergraphs: Clustering, classification, and embedding
Dengyong Zhou, Jiayuan Huang, and Bernhard Sch¨ olkopf. Learning with hypergraphs: Clustering, classification, and embedding. InAdvances in Neural Information Processing Systems 19, pages 1601–1608. MIT Press, 2006. 16
2006
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.