Recognition: unknown
Donor-Aware scRNA-seq Benchmarks for IBD Classification
Pith reviewed 2026-05-09 16:43 UTC · model grok-4.3
The pith
Compartment-stratified CLR composition and GatedStructuralCFN embeddings classify IBD donors at AUROC 0.95-0.98 under strict donor-aware validation
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
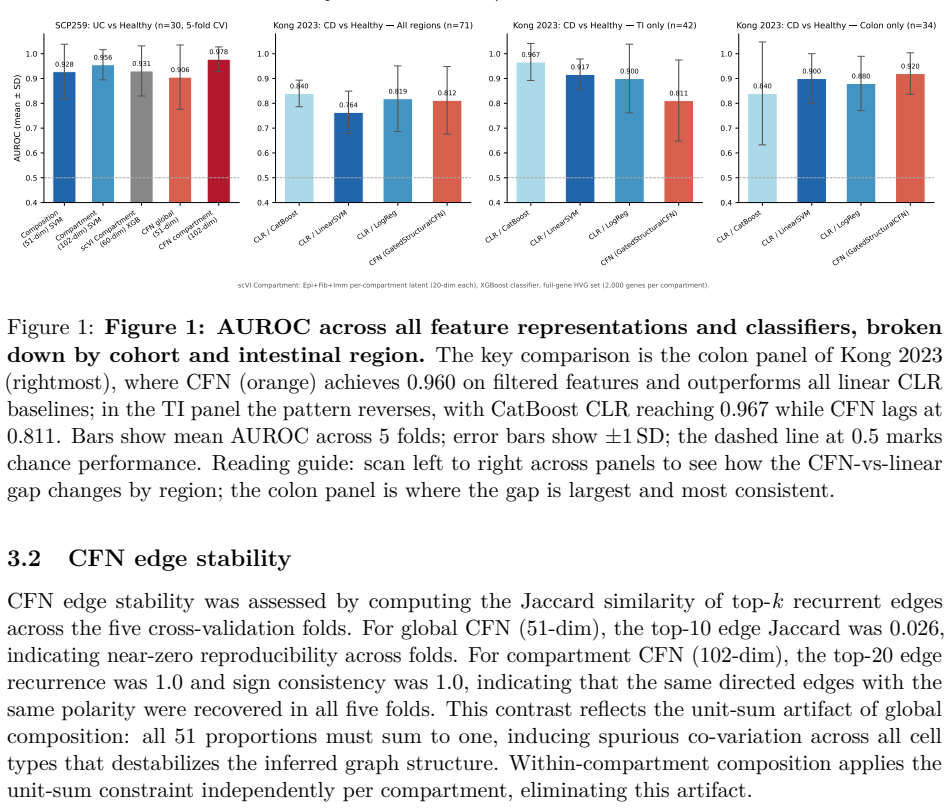
Compartment-stratified CLR transformed cell-type composition achieves AUROC 0.956 +/- 0.061 on SCP259 while GatedStructuralCFN on identical features reaches 0.978 +/- 0.050; in the Kong cohort CFN peaks at 0.960 +/- 0.055 in colon after filtering and exceeds linear CLR (0.900), with compartment-wise composition eliminating spurious unit-sum instability (Jaccard 0.026 versus top-20 recurrence of 1.0).
What carries the argument
Compartment-stratified CLR cell-type composition vectors fed into GatedStructuralCFN dependency embeddings, which extract stable inter-cell-type relations within each anatomical compartment while enforcing donor separation during training and testing.
If this is right
- Compartment stratification is required to remove spurious correlations induced by the unit-sum constraint in cell composition features.
- GatedStructuralCFN embeddings deliver a numerical edge over linear classifiers specifically in the colon region of Crohn's disease.
- Cross-dataset transfer between Crohn's and ulcerative colitis cohorts reaches only modest AUC (0.833) when limited to four shared cell types.
- Edge stability analysis shows compartment-wise features produce fully recurrent top dependencies whereas global composition does not.
Where Pith is reading between the lines
- The regional performance gap between colon and ileum suggests that separate models per intestinal compartment may be needed for optimal clinical translation.
- Verification of cell-type and compartment labels with orthogonal assays such as spatial transcriptomics would strengthen the benchmark reliability.
- The same donor-aware, compartment-stratified workflow could be applied directly to other multi-region tissues or non-IBD inflammatory conditions.
Load-bearing premise
Cell-type annotations and compartment labels are accurate and consistent across the two cohorts, and the donor-aware cross-validation fully blocks any information leakage between training and test donors.
What would settle it
Re-run the full pipeline after deliberately swapping compartment labels on a subset of cells or after allowing donor overlap in the train-test splits and measure whether AUROC falls below 0.90.
Figures

read the original abstract
Donor-level disease classification from single-cell RNA sequencing (scRNA-seq) requires strict donor-aware cross-validation: naive pipelines that split cells randomly conflate training and test donors, inflating reported performance through pseudoreplication. We present a donor-aware benchmark evaluating three feature representations across two independent IBD cohorts: centered log-ratio (CLR) transformed cell-type composition, GatedStructuralCFN dependency embeddings, and scVI variational autoencoder latent embeddings. The cohorts are the SCP259 ulcerative colitis atlas (UC vs. Healthy, n=30 donors, 51 cell types) and the Kong 2023 Crohn's disease atlas (CD vs. Healthy, n=71 donors, 55-68 cell types across three intestinal regions). Compartment-stratified CLR composition achieves AUROC 0.956 +/- 0.061 on SCP259; GatedStructuralCFN on the same features achieves 0.978 +/- 0.050. In the Kong cohort, CFN achieves its best performance in the colon region (0.960 +/- 0.055 after feature filtering), exceeding linear CLR (0.900 +/- 0.100), while terminal ileum classification is dominated by linear models (CatBoost CLR 0.967 +/- 0.075 vs. CFN 0.811 +/- 0.164). Cross-dataset transfer (CD->UC, four shared cell types) achieves AUC 0.833 with XGBoost CLR; the reverse direction performs at chance. CFN edge stability analysis shows that compartment-wise composition eliminates spurious unit-sum-induced instability present in global composition (Jaccard 0.026 vs. top-20 recurrence 1.0). CFN shows a consistent numerical advantage over linear models in the colon region of CD (AUROC 0.960 vs. 0.900), though no inter-method comparison reached statistical significance at n<=34 donors per region. Compartment-aware feature construction is critical for both classification performance and structural interpretability. Code: https://github.com/Jonathan-321/sfn-scrna-study
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper benchmarks donor-aware classification of IBD (UC vs healthy in SCP259; CD vs healthy in Kong) from scRNA-seq using three feature sets: compartment-stratified CLR cell-type compositions, GatedStructuralCFN dependency embeddings, and scVI latents. It stresses strict donor-level cross-validation to avoid pseudoreplication, reports AUROCs (e.g., 0.956 for CLR and 0.978 for CFN on SCP259; 0.960 for CFN in Kong colon after filtering), notes non-significant differences at small donor counts (n≤34), and highlights compartment-aware construction for performance and edge stability.
Significance. If the donor-aware splits and label harmonization hold, the work supplies a useful empirical reference for scRNA-seq disease classification, demonstrating that compartment stratification mitigates composition-induced instability and that structural embeddings can numerically outperform linear baselines in specific regions, while underscoring the limits of statistical power with current cohort sizes.
major comments (3)
- [Methods] Methods (donor-aware CV implementation): the manuscript does not specify whether feature filtering, normalization, or any global preprocessing steps (explicitly mentioned for the Kong colon results) were performed inside or outside the donor-level folds; any global step would introduce leakage and undermine the central claim that the reported AUROCs (0.956–0.978) are unbiased.
- [Results] Results (Kong cohort, colon region): the comparison of CFN (0.960 ± 0.055) vs linear CLR (0.900 ± 0.100) after 'feature filtering' lacks the exact filtering rule, threshold, or selection criterion; without this, the numerical advantage cannot be reproduced or interpreted as evidence of CFN superiority.
- [Methods] Methods and cross-dataset transfer: cell-type harmonization between SCP259 (51 types) and Kong (55–68 types, three regions) is not detailed for the four shared types used in CD→UC transfer (AUC 0.833); any systematic annotation mismatch would affect both CLR and CFN equally and render the performance gap uninterpretable.
minor comments (2)
- [Abstract] Abstract and Results: report the exact statistical test (e.g., paired Wilcoxon or DeLong) and p-values for all inter-method comparisons rather than only stating 'no statistical significance'.
- [Methods] Figure legends or Methods: clarify how compartment labels were assigned and whether they were validated against independent annotations.
Simulated Author's Rebuttal
We thank the referee for their detailed and constructive comments on our manuscript. We address each of the major comments below and have made revisions to the manuscript to incorporate the suggested clarifications.
read point-by-point responses
-
Referee: [Methods] Methods (donor-aware CV implementation): the manuscript does not specify whether feature filtering, normalization, or any global preprocessing steps (explicitly mentioned for the Kong colon results) were performed inside or outside the donor-level folds; any global step would introduce leakage and undermine the central claim that the reported AUROCs (0.956–0.978) are unbiased.
Authors: We agree that explicit specification of the cross-validation procedure is essential to support the unbiased nature of the reported performance metrics. All feature filtering, normalization, and preprocessing steps were conducted strictly within each donor-level training fold, with no information from the test donors used at any stage. We have revised the Methods section to include a detailed description of the donor-aware CV pipeline, including confirmation that global steps were avoided, along with pseudocode illustrating the process. revision: yes
-
Referee: [Results] Results (Kong cohort, colon region): the comparison of CFN (0.960 ± 0.055) vs linear CLR (0.900 ± 0.100) after 'feature filtering' lacks the exact filtering rule, threshold, or selection criterion; without this, the numerical advantage cannot be reproduced or interpreted as evidence of CFN superiority.
Authors: The referee is correct that the precise filtering criteria were not fully specified in the original submission. For the Kong colon results, feature filtering consisted of excluding cell types present in fewer than 10% of donors within the training fold (a threshold selected to ensure sufficient data for reliable composition estimation). This was applied independently per fold. We have updated the Results section with this exact rule and added a supplementary table showing the filtered cell types for transparency. revision: yes
-
Referee: [Methods] Methods and cross-dataset transfer: cell-type harmonization between SCP259 (51 types) and Kong (55–68 types, three regions) is not detailed for the four shared types used in CD→UC transfer (AUC 0.833); any systematic annotation mismatch would affect both CLR and CFN equally and render the performance gap uninterpretable.
Authors: We acknowledge that the cell-type harmonization process for the cross-dataset transfer experiment was insufficiently described. The four shared types were identified by aligning cell-type labels based on shared marker genes and standard nomenclature from the original publications (specifically: T cells, B cells, Enterocytes, and Macrophages). We have added a dedicated subsection in the Methods detailing the harmonization criteria and a mapping table. While we agree that annotation inconsistencies could impact interpretability, the primary conclusions of the paper rely on within-cohort analyses, and the transfer results are presented as exploratory. revision: yes
Circularity Check
No derivation chain present; purely empirical benchmark
full rationale
The manuscript is an empirical benchmark study that reports AUROC values obtained via donor-aware cross-validation on held-out donor data from two IBD cohorts. No equations, first-principles derivations, predictions, or uniqueness theorems are advanced that could reduce to fitted parameters, self-citations, or ansatzes by construction. All reported metrics (e.g., 0.956 AUROC for compartment-stratified CLR, 0.978 for GatedStructuralCFN) are direct measurements on independent test splits; compartment-stratified feature construction and edge-stability analysis are likewise post-hoc empirical observations rather than deductive steps. The paper therefore contains no load-bearing circular reductions.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Donor-aware cross-validation is necessary to avoid pseudoreplication in scRNA-seq classification tasks
Reference graph
Works this paper leans on
-
[1]
Cell , volume=
Intra- and inter-cellular rewiring of the human colon during ulcerative colitis , author=. Cell , volume=. 2019 , publisher=
2019
-
[2]
The landscape of immune dysregulation in
Kong, Lingjia and Pokatayev, Vladislav and Lefkovith, Ariel and Carter, Grace T and Creasey, Elizabeth A and Krishna, Chirag and Subramanian, Sathish and Kochar, Bharati and Ashenberg, Orr and Lau, Helena and Ananthakrishnan, Ashwin N and Graham, Daniel B and Deguine, Jacques and Xavier, Ramnik J , journal=. The landscape of immune dysregulation in. 2023 ...
2023
-
[3]
Nature Methods , volume=
Deep generative modeling for single-cell transcriptomics , author=. Nature Methods , volume=. 2018 , publisher=
2018
-
[4]
2026 , eprint=
Interpretable Functional Compositions for Tabular Discovery , author=. 2026 , eprint=
2026
-
[5]
Journal of the Royal Statistical Society: Series B (Methodological) , volume=
The statistical analysis of compositional data , author=. Journal of the Royal Statistical Society: Series B (Methodological) , volume=. 1982 , publisher=
1982
-
[6]
2016 , doi=
Chen, Tianqi and Guestrin, Carlos , booktitle=. 2016 , doi=
2016
-
[7]
Dorogush, Anna Veronika and Ershov, Vasily and Gulin, Andrey , year=. doi:10.48550/arXiv.1810.11363 , url=. 1810.11363 , archivePrefix=
-
[8]
Journal of the Royal Statistical Society: Series B (Methodological) , author =
John Aitchison. The statistical analysis of compositional data. Journal of the Royal Statistical Society: Series B (Methodological), 44 0 (2): 0 139--160, 1982. doi:10.1111/j.2517-6161.1982.tb01195.x
-
[9]
Lingjia Kong, Vladislav Pokatayev, Ariel Lefkovith, Grace T Carter, Elizabeth A Creasey, Chirag Krishna, Sathish Subramanian, Bharati Kochar, Orr Ashenberg, Helena Lau, Ashwin N Ananthakrishnan, Daniel B Graham, Jacques Deguine, and Ramnik J Xavier. The landscape of immune dysregulation in Crohn's disease revealed through single-cell transcriptomic profil...
-
[10]
Interpretable functional compositions for tabular discovery, 2026
Fang Li. Interpretable functional compositions for tabular discovery, 2026. URL https://arxiv.org/abs/2601.20037. Department of Computer Science, Oklahoma Christian University. Code: https://github.com/fanglioc/StructuralCFN-public
-
[11]
Aaron Lou, Chenlin Meng, and Stefano Ermon
Romain Lopez, Jeffrey Regier, Michael B Cole, Michael I Jordan, and Nir Yosef. Deep generative modeling for single-cell transcriptomics. Nature Methods, 15 0 (12): 0 1053--1058, 2018. doi:10.1038/s41592-018-0229-2
-
[12]
Intra- and inter-cellular rewiring of the human colon during ulcerative colitis
Christopher S Smillie, Moshe Biton, Jose Ordovas-Montanes, Keri M Sullivan, Grace Burgin, Daniel B Graham, Rebecca H Herbst, Noga Rogel, Michal Slyper, Julia Waldman, et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell, 178 0 (3): 0 714--730, 2019. doi:10.1016/j.cell.2019.06.029
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.