Recognition: unknown
SHIELD: A Diverse Clinical Note Dataset and Distilled Small Language Models for Enterprise-Scale De-identification
Pith reviewed 2026-05-07 16:48 UTC · model grok-4.3
The pith
Small language models distilled from large ones match teacher performance on structured patient identifiers in clinical notes at 0.88 precision and 0.86 recall on standard hardware.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
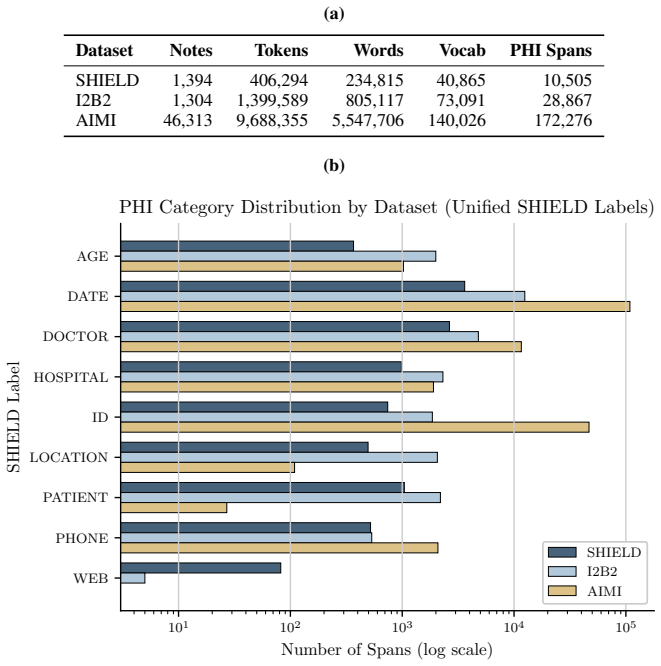
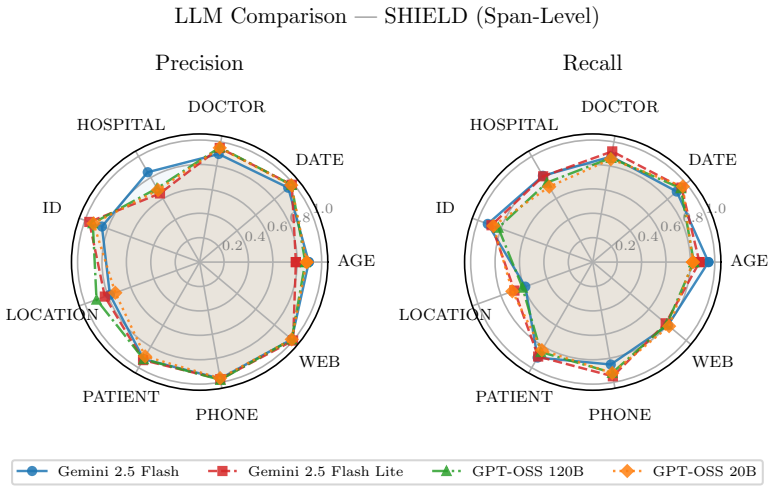
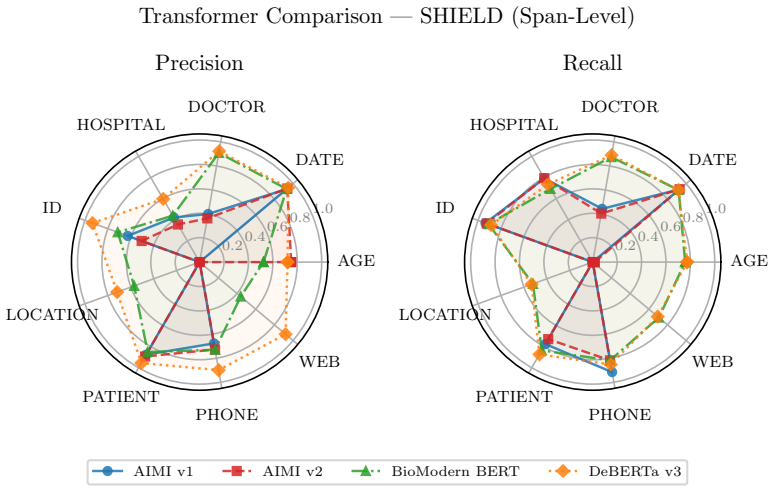
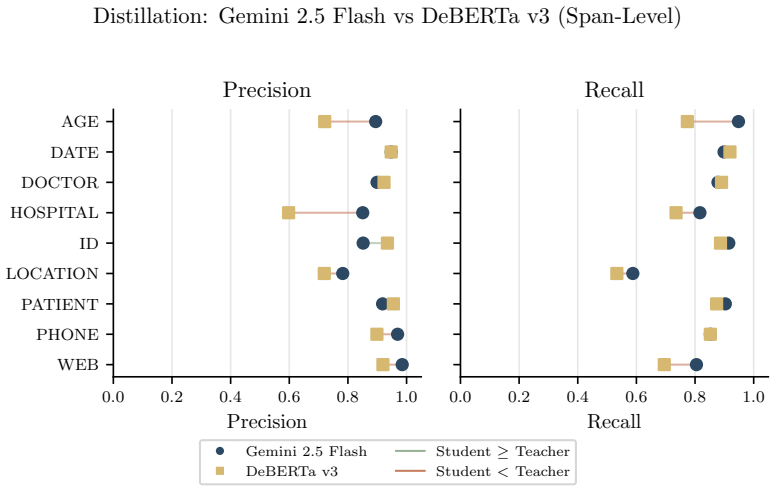
SHIELD supplies 1,394 notes and 10,505 annotated PHI spans; after large language models establish an upper bound, distilled small language models achieve equivalent results to the teacher on structured PHI categories and reach micro-averaged span-level precision of 0.88 with recall of 0.86 on ordinary workstation hardware.
What carries the argument
Knowledge distillation from large language models into small language models trained on the SHIELD dataset, which itself is assembled by set-cover diversity sampling plus human-in-the-loop adjudication.
If this is right
- De-identification can run entirely on local hardware without transmitting protected health information to external APIs.
- Models trained on diverse data generalize reliably across institutions for universal structured PHI categories.
- Institution-specific PHI entities require additional specialized models or adaptation for high-volume notes.
- Public release of the dataset and DeBERTa v3 model enables further fine-tuning and benchmarking by other groups.
Where Pith is reading between the lines
- The same distillation pipeline could be reused for privacy-sensitive text in legal or financial domains.
- If the diversity claim holds, SHIELD could serve as a replacement benchmark for clinical natural language processing tasks.
- Persistent gaps on institution-specific entities suggest that periodic retraining or federated updates will be needed in real deployments.
Load-bearing premise
The set-cover diversity sampling combined with human-in-the-loop adjudication yields a dataset representative of modern clinical narratives that supports generalization beyond the sampled notes and institutions.
What would settle it
Testing the distilled model on a large collection of notes drawn from institutions excluded from the original sampling and observing a drop in micro-averaged precision or recall below 0.80 would falsify the generalization claim.
Figures






read the original abstract
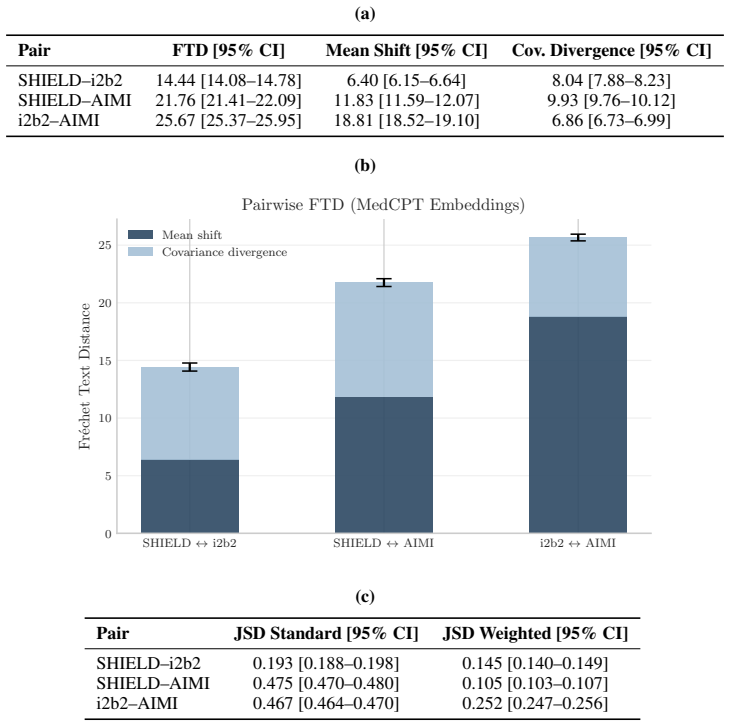
De-identification of clinical text remains essential for secondary use of electronic health records (EHRs), yet public benchmarks such as i2b2 2006/2014 are over a decade old and lack the semantic and demographic diversity of modern narratives. While Large Language Models (LLMs) achieve state-of-the-art zero-shot extraction, enterprise deployment is hindered by compute costs and governance restricting Protected Health Information (PHI) from cloud APIs. We introduce SHIELD (Synthetic Human-annotated Identifier-replaced Entries for Learning and De-identification), a diverse dataset of 1,394 notes with 10,505 gold-standard PHI spans across 9 categories, built via set-cover diversity sampling with human-in-the-loop adjudication. We evaluate four LLMs (two proprietary, two open-weight) to establish a performance ceiling, then distill these capabilities into locally deployable Small Language Models (SLMs). Distributional analysis using Frechet Text Distance and Jensen-Shannon Divergence confirms SHIELD occupies a distinct region of biomedical embedding and vocabulary space versus legacy benchmarks. Our best distilled model matches its teacher on structured PHI categories (DATE, DOCTOR, ID, PATIENT, PHONE) and achieves micro-averaged span-level precision of 0.88 and recall of 0.86 on standard workstation hardware. Cross-dataset evaluation shows diversity-trained models generalize well on universal structured PHI, while institution-specific entities remain hard to transfer, suggesting optimal deployment combines broad-coverage models with specialized models for high-volume notes. We publicly release the SHIELD dataset and the distilled DeBERTa v3 model.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript presents SHIELD, a dataset of 1,394 clinical notes with 10,505 gold-standard PHI spans across 9 categories, constructed via set-cover diversity sampling and human-in-the-loop adjudication to overcome limitations of legacy benchmarks such as i2b2. It evaluates four LLMs (proprietary and open-weight) to establish performance ceilings, distills these into locally deployable small language models, and reports that the best distilled model matches its teacher on structured PHI categories (DATE, DOCTOR, ID, PATIENT, PHONE) while achieving micro-averaged span-level precision of 0.88 and recall of 0.86 on standard hardware. Distributional metrics (Fréchet Text Distance, Jensen-Shannon Divergence) position SHIELD apart from prior data, and cross-dataset tests indicate strong transfer on universal structured PHI but weaker results on institution-specific entities, leading to a recommendation for hybrid deployment; the dataset and a distilled DeBERTa v3 model are publicly released.
Significance. If the representativeness claims hold, the work is significant for providing a modern, diverse clinical de-identification resource that addresses the semantic and demographic gaps in decade-old public benchmarks. The distillation approach yielding competitive SLM performance on workstation hardware is practically relevant for privacy-sensitive enterprise settings where cloud LLMs are restricted. Public release of the dataset and model supports reproducibility and community follow-up, while the concrete metrics and distributional comparisons strengthen the contribution to applied NLP for healthcare.
major comments (2)
- [Dataset construction and distributional analysis] The central claim that SHIELD supports generalization beyond sampled notes and institutions for enterprise deployment rests on set-cover diversity sampling yielding representativeness, yet the manuscript provides no direct evidence that the chosen cover features (embeddings, metadata, vocabulary) span demographic, semantic, and stylistic variation across institutions; the Fréchet Text Distance and Jensen-Shannon Divergence results only show separation from i2b2 rather than positive coverage or absence of source-specific bias (dataset construction and distributional analysis sections).
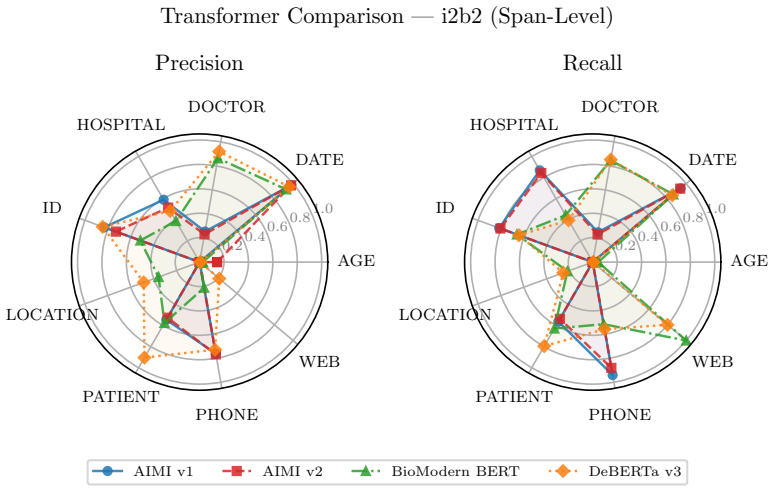
- [Cross-dataset evaluation] Cross-dataset evaluation results (strong on universal structured PHI, poor on institution-specific) are presented as supporting a hybrid deployment strategy, but this pattern is the expected outcome if the sampling pool is narrow and directly weakens the abstract's claim that the distilled model is ready for general workstation deployment (cross-dataset evaluation section).
minor comments (2)
- [Abstract] The abstract states '9 categories' without enumerating them; listing the PHI categories explicitly would improve immediate clarity.
- [Results] Ensure uniform terminology for 'micro-averaged span-level precision and recall' across results tables and text to prevent minor ambiguity in metric reporting.
Simulated Author's Rebuttal
We thank the referee for the constructive and detailed feedback. We address each major comment point by point below, providing clarifications on the scope of our claims and proposing targeted revisions to improve precision without overstating the results.
read point-by-point responses
-
Referee: The central claim that SHIELD supports generalization beyond sampled notes and institutions for enterprise deployment rests on set-cover diversity sampling yielding representativeness, yet the manuscript provides no direct evidence that the chosen cover features (embeddings, metadata, vocabulary) span demographic, semantic, and stylistic variation across institutions; the Fréchet Text Distance and Jensen-Shannon Divergence results only show separation from i2b2 rather than positive coverage or absence of source-specific bias (dataset construction and distributional analysis sections).
Authors: The set-cover diversity sampling was applied within our single-institution note pool to maximize coverage of the observed embedding, metadata, and vocabulary features, producing a subset more representative of that pool than random sampling. The FTD and JSD metrics establish that SHIELD occupies a distinct region relative to i2b2, supporting its utility as a modern benchmark. We acknowledge that these steps do not constitute direct evidence of spanning all demographic, semantic, or stylistic variations across institutions, as the source data is institution-specific. We will revise the dataset construction and distributional analysis sections to explicitly limit the representativeness claim to the sampled pool and to note that broader generalization requires further validation or adaptation. revision: partial
-
Referee: Cross-dataset evaluation results (strong on universal structured PHI, poor on institution-specific) are presented as supporting a hybrid deployment strategy, but this pattern is the expected outcome if the sampling pool is narrow and directly weakens the abstract's claim that the distilled model is ready for general workstation deployment (cross-dataset evaluation section).
Authors: The cross-dataset results indeed reflect the single-institution sampling pool, with strong transfer on universal structured categories and weaker performance on institution-specific ones. This pattern directly motivates the hybrid deployment recommendation already present in the abstract and discussion. The manuscript reports workstation performance while explicitly suggesting combination with specialized models rather than claiming the distilled model is ready for unrestricted general deployment. We will revise the abstract to sharpen this distinction and foreground the hybrid strategy as a core practical takeaway. revision: yes
Circularity Check
No circularity: empirical results rest on independent gold-standard annotations and standard sampling
full rationale
The paper's derivation chain consists of dataset construction via set-cover sampling plus human adjudication, LLM evaluation to set a ceiling, distillation to SLMs, and direct span-level P/R measurement against the human-created gold labels. These steps do not reduce by construction to fitted parameters or self-referential definitions. Distributional metrics (Frechet Text Distance, Jensen-Shannon) are computed independently on embeddings and vocabulary. No load-bearing self-citations, no uniqueness theorems imported from prior author work, and no equations that make the reported 0.88/0.86 scores tautological. Performance claims are falsifiable against external benchmarks and the released dataset.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Human-in-the-loop adjudication produces reliable gold-standard PHI annotations.
- domain assumption Distillation from large LLMs to small models preserves performance on structured extraction tasks.
Reference graph
Works this paper leans on
-
[1]
ModernBERT or DeBERTaV3 ? Examining architecture and data influence on transformer encoder models performance
Wissam Antoun, Beno \^i t Sagot, and Djam \'e Seddah. ModernBERT or DeBERTaV3 ? Examining architecture and data influence on transformer encoder models performance. In Proceedings of the 14th International Joint Conference on Natural Language Processing and the 4th Conference of the Asia-Pacific Chapter of the Association for Computational Linguistics (IJ...
2025
-
[2]
Fr\' e chet distance for offline evaluation of information retrieval systems with sparse labels
Negar Arabzadeh and Charles Clarke. Fr\' e chet distance for offline evaluation of information retrieval systems with sparse labels. In Proceedings of the 18th Conference of the European Chapter of the Association for Computational Linguistics (Volume 1: Long Papers), pages 420--431, St. Julian's, Malta, 2024. Association for Computational Linguistics. UR...
2024
-
[3]
Peter Belcak, Greg Heinrich, Shizhe Diao, Yonggan Fu, Xin Dong, Saurav Muralidharan, Yingyan Celine Lin, and Pavlo Molchanov. Small Language Models are the Future of Agentic AI , June 2025. URL http://arxiv.org/abs/2506.02153. arXiv:2506.02153 [cs]
-
[4]
The stanford medicine data science ecosystem for clinical and translational research
Alison Callahan, Euan Ashley, Somalee Datta, Priyamvada Desai, Todd A Ferris, Jason A Fries, Michael Halaas, Curtis P Langlotz, Sean Mackey, José D Posada, Michael A Pfeffer, and Nigam H Shah. The stanford medicine data science ecosystem for clinical and translational research. JAMIA Open, 6 0 (3): 0 ooad054, 08 2023. ISSN 2574-2531. doi:10.1093/jamiaopen...
-
[5]
Pierre J Chambon, Christopher Wu, Jackson M Steinkamp, Jason Adleberg, Tessa S Cook, and Curtis P Langlotz. Automated deidentification of radiology reports combining transformer and ``hide in plain sight'' rule-based methods. Journal of the American Medical Informatics Association, 30 0 (2): 0 318--328, January 2023. ISSN 1067-5027, 1527-974X. doi:10.1093...
-
[6]
Min Jin Chong and David Forsyth. Effectively unbiased FID and inception score and where to find them. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pages 6070--6079, 2020. doi:10.1109/CVPR42600.2020.00611
-
[7]
Exploiting vocabulary frequency imbalance in language model pre-training, 2025
Woojin Chung and Jeonghoon Kim. Exploiting vocabulary frequency imbalance in language model pre-training, 2025. URL https://arxiv.org/abs/2508.15390. NeurIPS 2025
-
[8]
A new paradigm for accelerating clinical data science at stanford medicine, 2020
Somalee Datta, Jose Posada, Garrick Olson, Wencheng Li, Ciaran O'Reilly, Deepa Balraj, Joseph Mesterhazy, Joseph Pallas, Priyamvada Desai, and Nigam Shah. A new paradigm for accelerating clinical data science at stanford medicine, 2020. URL https://arxiv.org/abs/2003.10534
-
[9]
D. C. Dowson and B. V. Landau. The Fr\' e chet distance between multivariate normal distributions. Journal of Multivariate Analysis, 12 0 (3): 0 450--455, 1982. doi:10.1016/0047-259X(82)90077-X
-
[10]
Hannah Eyre, Qiwei Gan, Mengke Hu, Annie Bowles, Johnathan Stanley, Jianlin Shi, Scott L. DuVall, and Patrick R. Alba. Evaluating Clinical Note Deidentification Tools and Transformer Transferability between Public and Private Data from the US Department of Veterans Affairs , June 2025. URL https://www.medrxiv.org/content/10.1101/2025.03.21.25323520v2
-
[11]
Vertex AI — Generative AI Pricing
Google Cloud . Vertex AI — Generative AI Pricing . https://cloud.google.com/vertex-ai/generative-ai/pricing, 2025. Gemini 2.5 Flash Flex/Batch pricing: \ 0.15 per 1M input tokens, \ 1.25 per 1M output tokens. Accessed March 2025
2025
-
[12]
Pengcheng He, Jianfeng Gao, and Weizhu Chen
Pengcheng He, Jianfeng Gao, and Weizhu Chen. Debertav3: Improving deberta using electra-style pre-training with gradient-disentangled embedding sharing. arXiv preprint arXiv:2111.09543, 2021
-
[13]
Paul M Heider and St \'e phane M Meystre. An Extensible Evaluation Framework Applied to Clinical Text Deidentification Natural Language Processing Tools : Multisystem and Multicorpus Study . Journal of Medical Internet Research, 26: 0 e55676, May 2024. ISSN 1439-4456. doi:10.2196/55676. URL https://pmc.ncbi.nlm.nih.gov/articles/PMC11167315/
-
[14]
GANs trained by a two time-scale update rule converge to a local Nash equilibrium
Martin Heusel, Hubert Ramsauer, Thomas Unterthiner, Bernhard Nessler, and Sepp Hochreiter. GANs trained by a two time-scale update rule converge to a local Nash equilibrium. In Advances in Neural Information Processing Systems, volume 30, 2017
2017
-
[15]
Qiao Jin, Won Kim, Qingyu Chen, Donald C. Comeau, Lana Yeganova, W. John Wilbur, and Zhiyong Lu. MedCPT : Contrastive Pre -trained Transformers with large-scale PubMed search logs for zero-shot biomedical information retrieval. Bioinformatics, 39 0 (11), 2023. doi:10.1093/bioinformatics/btad651
-
[16]
Aman Kansal, Emma Chen, Boyang Tom Jin, Pranav Rajpurkar, and David A. Kim. MC-MED , multimodal clinical monitoring in the emergency department. Scientific Data, 12 0 (1): 0 1094, 2025. doi:10.1038/s41597-025-05419-5. URL https://doi.org/10.1038/s41597-025-05419-5
-
[17]
Wiest, Stephen Gilbert, and Daniel Truhn
Jakob Nikolas Kather, Dyke Ferber, Isabelle C. Wiest, Stephen Gilbert, and Daniel Truhn. Large language models could make natural language again the universal interface of healthcare. Nature Medicine, 30: 0 2708--2710, 2024. doi:10.1038/s41591-024-03259-5
-
[18]
Veysel Kocaman, Hasham Ul Haq, and David Talby. Beyond Accuracy : Automated De - Identification of Large Real - World Clinical Text Datasets , December 2023. URL http://arxiv.org/abs/2312.08495. arXiv:2312.08495 [cs]
-
[19]
Automated De - Identification , Consistent Obfuscation , and Regulatory Grade Validation of 2 Billion Patient Notes , September 2025
Veysel Kocaman, Lindsay Mico, Mustafa Aytug Kaya, Nadaa Taiyab, David Talby, Tae Surh, Yuqing Guo, Vivek Tomer, and Robert Kramer. Automated De - Identification , Consistent Obfuscation , and Regulatory Grade Validation of 2 Billion Patient Notes , September 2025. URL https://www.researchsquare.com/article/rs-6867162/v1
2025
-
[20]
Aleksandar Kovačević, Bojana Bašaragin, Nikola Milošević, and Goran Nenadić. De-identification of clinical free text using natural language processing: A systematic review of current approaches. Artificial Intelligence in Medicine, 151: 0 102845, May 2024. ISSN 09333657. doi:10.1016/j.artmed.2024.102845. URL https://linkinghub.elsevier.com/retrieve/pii/S0...
-
[21]
Shah, Richard Dobson, and James Teo
Zeljko Kraljevic, Anthony Shek, Joshua Au Yeung, Ewart Jonathan Sheldon, Mohammad Al-Agil, Haris Shuaib, Xi Bai, Kawsar Noor, Anoop D. Shah, Richard Dobson, and James Teo. Validating transformers for redaction of text from electronic health records in real-world healthcare, October 2023. URL http://arxiv.org/abs/2310.04468. arXiv:2310.04468 [cs]
-
[22]
Rachel Kuo, Andrew A. S. Soltan, Ciaran O'Hanlon, Alan Hasanic, David A. Clifton, Collins Gary, Dominic Furniss, and David W. Eyre. Benchmarking transformer-based models for medical record deidentification: A single centre, multi-specialty evaluation, May 2025. URL https://www.medrxiv.org/content/10.1101/2025.05.05.25326979v1
-
[23]
Hee-Jin Lee, Zhen Guo, Luchao Jin, and Morteza Moazami Goudarzi. Targeted Error Correction in Knowledge Distillation : Small Language Models Surpass GPT , November 2025. URL http://arxiv.org/abs/2511.03005. arXiv:2511.03005 [cs]
-
[24]
RAD : Towards Trustworthy Retrieval - Augmented Multi -modal Clinical Diagnosis , 2025
Haolin Li, Tianjie Dai, Zhe Chen, Siyuan Du, Jiangchao Yao, Ya Zhang, and Yanfeng Wang. RAD : Towards Trustworthy Retrieval - Augmented Multi -modal Clinical Diagnosis , 2025. URL https://arxiv.org/abs/2509.19980. NeurIPS 2025
-
[25]
Jianhua Lin. Divergence measures based on the Shannon entropy. IEEE Transactions on Information Theory, 37 0 (1): 0 145--151, 1991. doi:10.1109/18.61115
-
[26]
DeID - GPT : Zero -shot Medical Text De - Identification by GPT -4, December 2023
Zhengliang Liu, Yue Huang, Xiaowei Yu, Lu Zhang, Zihao Wu, Chao Cao, Haixing Dai, Lin Zhao, Yiwei Li, Peng Shu, Fang Zeng, Lichao Sun, Wei Liu, Dinggang Shen, Quanzheng Li, Tianming Liu, Dajiang Zhu, and Xiang Li. DeID - GPT : Zero -shot Medical Text De - Identification by GPT -4, December 2023. URL http://arxiv.org/abs/2303.11032. arXiv:2303.11032 [cs]
-
[27]
Evaluation metrics for headline generation using deep pre-trained embeddings
Abdul Moeed, Yang An, Gerhard Hagerer, and Georg Groh. Evaluation metrics for headline generation using deep pre-trained embeddings. In Proceedings of the Twelfth Language Resources and Evaluation Conference, pages 1796--1802, Marseille, France, 2020. European Language Resources Association. URL https://aclanthology.org/2020.lrec-1.222/
2020
-
[28]
Neilson, Moniruzzaman Moni, Marcello Nesca, Alexander Singer, and Jennifer E
Bekelu Negash, Alan Katz, Christine J. Neilson, Moniruzzaman Moni, Marcello Nesca, Alexander Singer, and Jennifer E. Enns. De-identification of Free Text Data containing Personal Health Information : A Scoping Review of Reviews . International Journal of Population Data Science, 8 0 (1), December 2023. ISSN 2399-4908. doi:10.23889/ijpds.v8i1.2153. URL htt...
-
[29]
gpt-oss-120b & gpt-oss-20b Model Card
OpenAI . GPT-OSS : Open -weight models for reasoning, agentic tasks, and versatile developer use cases. arXiv preprint arXiv:2508.10925, 2025. URL https://huggingface.co/openai/gpt-oss-120b. Mixture-of-Experts architecture; GPT-OSS 120B (117B total, 5.1B active parameters) and GPT-OSS 20B (21B total, 3.6B active parameters). Apache 2.0 license
work page internal anchor Pith review arXiv 2025
-
[30]
Osborne, Andrew Trotter, Tobias O'Leary, Chris Coffee, Micah D
John D. Osborne, Andrew Trotter, Tobias O'Leary, Chris Coffee, Micah D. Cochran, Luis Mansilla-Gonzalez, Akhil Nadimpalli, Alex McAnnally, Abdulateef I. Almudaifer, Jeffrey R. Curtis, Salma M. Aly, and Richard E. Kennedy. A Markov Chain Replacement Strategy for Surrogate Identifiers : Minimizing Re - Identification Risk While Preserving Text Reuse . Elect...
-
[31]
Eva Prakash, Maayane Attias, Pierre Chambon, Justin Xu, Steven Truong, Jean-Benoit Delbrouck, Tessa Cook, and Curtis Langlotz. Improving the Performance of Radiology Report De -identification with Large - Scale Training and Benchmarking Against Cloud Vendor Methods , November 2025. URL http://arxiv.org/abs/2511.04079. arXiv:2511.04079 [cs]
-
[32]
RedactOR : An LLM - Powered Framework for Automatic Clinical Data De - Identification , July 2025
Praphul Singh, Charlotte Dzialo, Jangwon Kim, Sumana Srivatsa, Irfan Bulu, Sri Gadde, and Krishnaram Kenthapadi. RedactOR : An LLM - Powered Framework for Automatic Clinical Data De - Identification , July 2025. URL http://arxiv.org/abs/2505.18380. arXiv:2505.18380 [cs]
-
[33]
Thomas Sounack, Joshua Davis, Brigitte Durieux, Antoine Chaffin, Tom J. Pollard, Eric Lehman, Alistair E. W. Johnson, Matthew McDermott, Tristan Naumann, and Charlotta Lindvall. BioClinical ModernBERT : A State -of-the- Art Long - Context Encoder for Biomedical and Clinical NLP , June 2025. URL http://arxiv.org/abs/2506.10896. arXiv:2506.10896 [cs]
-
[34]
Annotating longitudinal clinical narratives for de-identification: The 2014 i2b2/uthealth corpus
Amber Stubbs and "O zlem Uzuner. Annotating longitudinal clinical narratives for de-identification: The 2014 i2b2/uthealth corpus. Journal of Biomedical Informatics, 58: 0 S20--S29, 2015. doi:10.1016/j.jbi.2015.07.020. PMID: 26319540
-
[35]
Evaluating the State -of-the- Art in Automatic De -identification
\"O zlem Uzuner, Yuan Luo, and Peter Szolovits. Evaluating the State -of-the- Art in Automatic De -identification. Journal of the American Medical Informatics Association, 14 0 (5): 0 550--563, September 2007. ISSN 1067-5027. doi:10.1197/jamia.M2444. URL https://doi.org/10.1197/jamia.M2444
-
[36]
Zuo Wang and Ye Yuan. Jensen- Shannon divergence message-passing for rich-text graph representation learning. arXiv preprint arXiv:2512.20094, 2025. URL https://arxiv.org/abs/2512.20094
-
[37]
Wiest, Marie-Elisabeth Le mann, Fabian Wolf, Dyke Ferber, Marko Van Treeck, Jiefu Zhu, Matthias P
Isabella C. Wiest, Marie-Elisabeth Le mann, Fabian Wolf, Dyke Ferber, Marko Van Treeck, Jiefu Zhu, Matthias P. Ebert, Christoph Benedikt Westphalen, Martin Wermke, and Jakob Nikolas Kather. Deidentifying Medical Documents with Local , Privacy - Preserving Large Language Models : The LLM - Anonymizer . NEJM AI, 2 0 (4): 0 AIdbp2400537, March 2025. doi:10.1...
-
[38]
The design of approximation algorithms
David P Williamson and David B Shmoys. The design of approximation algorithms. Cambridge university press, 2011
2011
-
[39]
Yuli Wu, Fucheng Liu, R \"u veyda Yilmaz, Henning Konermann, Peter Walter, and Johannes Stegmaier. A pragmatic note on evaluating generative models with Fr\' e chet inception distance for retinal image synthesis. In Proceedings of Medical Imaging with Deep Learning (MIDL), 2026. URL https://arxiv.org/abs/2502.17160
-
[40]
Xi Yang, Tianchen Lyu, Qian Li, Chih-Yin Lee, Jiang Bian, William R. Hogan, and Yonghui Wu. A study of deep learning methods for de-identification of clinical notes in cross-institute settings. BMC Medical Informatics and Decision Making, 19 0 (Suppl 5): 0 232, December 2019. ISSN 1472-6947. doi:10.1186/s12911-019-0935-4. URL https://pmc.ncbi.nlm.nih.gov/...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.