Recognition: unknown
Genotype specificity and spatial arrangement govern the direction and magnitude of selection in variable environments
Pith reviewed 2026-05-08 16:54 UTC · model grok-4.3
The pith
Genotype specificity determines whether environmental heterogeneity amplifies or suppresses selection, while spatial arrangement controls its magnitude.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The paper claims that across three common classes of genotype-environment interactions and a wide range of spatial arrangements of environmental states, genotype specificity governs the direction of selection change due to heterogeneity while spatial arrangement governs the magnitude, reconciling disparate prior results on whether spatial variation helps or hinders adaptation.
What carries the argument
A lattice graph model in which mutant and resident fitness depend on the local environmental state, yielding the principles that genotype specificity sets effect direction and spatial arrangement sets effect magnitude.
Load-bearing premise
The modeling choices that genotype-environment interactions fall into the three considered classes and that lattice graphs adequately capture spatial structure.
What would settle it
An observation in a biological system or simulation where spatial arrangement reverses the direction of the heterogeneity effect on selection, or where modulating mutant fitness amplifies rather than suppresses selection, would falsify the principles.
Figures
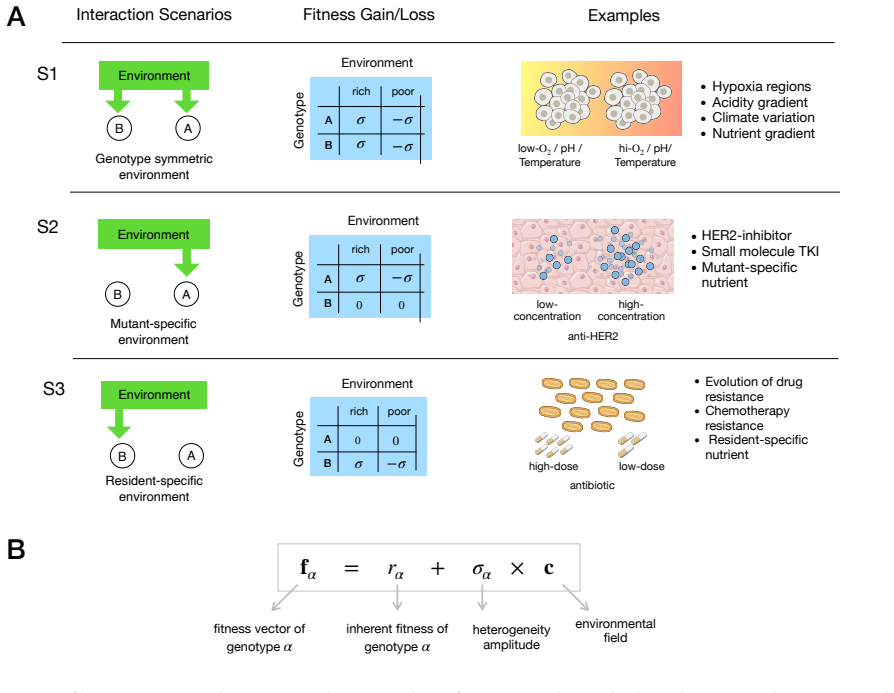
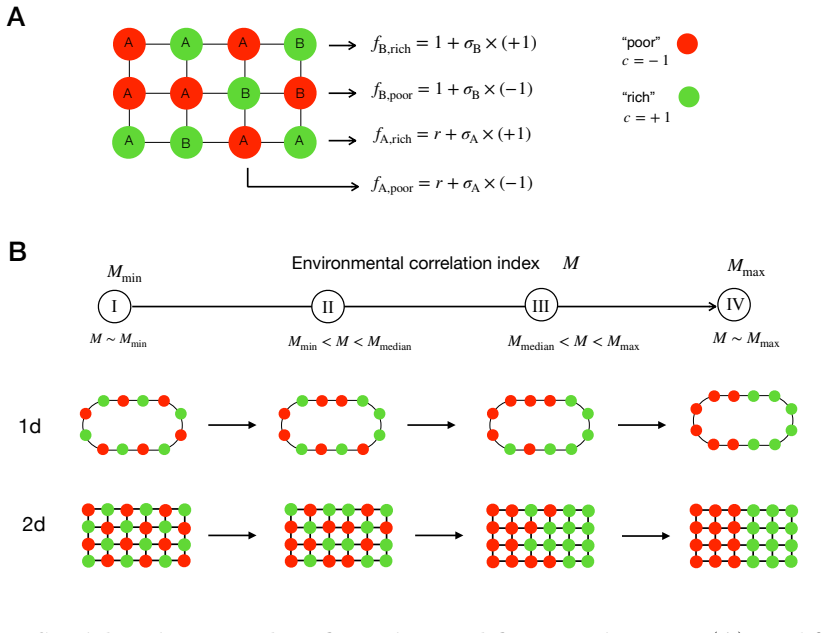



read the original abstract
Spatial environmental variation can either amplify or suppress the fixation of beneficial mutants in structured populations, yet the interplay of ecological factors and spatial structure in determining which outcome occurs remains theoretically unresolved. Here, we develop a unified framework for selection on lattice graphs with environmental heterogeneity, in which mutant and resident fitness depend on the local environmental state. Across three common classes of genotype-environment interactions and a wide range of spatial arrangements of environmental states, we identify two governing principles. Genotype specificity determines the direction of the effect: heterogeneity amplifies selection when it modulates resident fitness, but suppresses selection when it modulates mutant fitness, with genotype-symmetric modulation producing weaker amplification. Spatial arrangement determines the magnitude: intermixed versus clustered environments tune the strength of amplification or suppression without reversing the direction of the effect. Together, these principles reconcile disparate theoretical results and provide predictive criteria for adaptation in heterogeneous landscapes, from microbial communities to somatic evolution and cancer.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript develops a unified framework for selection on lattice graphs with environmental heterogeneity, where mutant and resident fitness depend on the local environmental state. Across three common classes of genotype-environment interactions and a range of spatial arrangements, it identifies two governing principles: genotype specificity determines the direction of the heterogeneity effect on selection (amplification when modulating resident fitness, suppression when modulating mutant fitness, with symmetric modulation yielding weaker effects), while spatial arrangement determines only the magnitude (intermixed vs. clustered environments tune strength without reversing direction). These principles are presented as reconciling prior theoretical results and providing predictive criteria for adaptation in heterogeneous landscapes.
Significance. If the derivations and simulations hold, the work is significant for providing a clear, predictive separation of direction and magnitude in how ecological heterogeneity and spatial structure interact to shape fixation probabilities. This unifies disparate findings in spatial evolutionary dynamics and offers testable criteria applicable to microbial communities, somatic evolution, and cancer. The systematic coverage of interaction classes and arrangements on lattices is a strength, enabling precise identification of the principles within the modeled systems.
minor comments (3)
- In the model section, clarify whether the local fitness function includes any implicit averaging over neighbors or is strictly focal-individual only; this would help readers assess the scope of the directionality principle.
- Figure legends for results on amplification/suppression factors should explicitly label the three GxE classes and the spatial configurations (intermixed/clustered) to improve readability.
- The discussion could briefly note potential empirical tests, such as in microbial metapopulations with controlled environmental patches, to strengthen the predictive claims.
Simulated Author's Rebuttal
We thank the referee for their positive and constructive review of our manuscript, including the recognition of its significance in unifying results on spatial evolutionary dynamics under environmental heterogeneity. We appreciate the recommendation for minor revision and will incorporate any editorial suggestions in the revised version.
Circularity Check
No significant circularity; derivation self-contained
full rationale
The paper constructs a modeling framework on lattice graphs, defines fitness via local environmental states for three specified GxE interaction classes, and computes outcomes (e.g., fixation probabilities) across spatial arrangements. The two governing principles are presented as emergent results from systematic variation of those inputs rather than presupposed definitions, fitted parameters renamed as predictions, or load-bearing self-citations. No equation or step reduces by construction to its own inputs; the separation of direction (genotype specificity) and magnitude (spatial arrangement) follows from explicit enumeration of the model classes. External benchmarks such as prior spatial selection literature are not required for the internal logic, and no ansatz or uniqueness theorem is smuggled in via self-reference.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Populations are structured on lattice graphs with local environmental states determining fitness
- domain assumption Three common classes of genotype-environment interactions cover the relevant cases
Reference graph
Works this paper leans on
-
[1]
The moran process on 2-chromatic graphs
Kamran Kaveh, Alex McAvoy, Krishnendu Chatterjee, and Martin A Nowak. The moran process on 2-chromatic graphs. PLOS Computational Biology , 16(11):e1008402, 2020
2020
-
[2]
Alternative to the diffusion equation in population genetics
Bahram Houchmandzadeh and Marcel Vallade. Alternative to the diffusion equation in population genetics. Physical Review E, 82(5):051913, 2010
2010
-
[3]
The fixation probability of a beneficial mutation in a geographically structured population
Bahram Houchmandzadeh and Marcel Vallade. The fixation probability of a beneficial mutation in a geographically structured population. New Journal of Physics , 13(7): 073020, 2011
2011
-
[4]
Evolutionary dynamics
Martin A Nowak. Evolutionary dynamics. Harvard University Press, 2006
2006
-
[5]
Fixation probabilities in a spatially heterogeneous environment
Sergey Gavrilets and Nathan Gibson. Fixation probabilities in a spatially heterogeneous environment. Population Ecology, 44(2):51–58, 2002
2002
-
[6]
Probability of fixation in a heteroge- neous environment
Michael C Whitlock and Richard Gomulkiewicz. Probability of fixation in a heteroge- neous environment. Genetics, 171(3):1407–1417, 2005
2005
-
[7]
Fixation probability in spatially changing envi- ronments
Hidenori Tachida and Masaru Iizuka. Fixation probability in spatially changing envi- ronments. Genetics Research, 58(3):243–251, 1991
1991
-
[8]
Heterogeneity in background fitness acts as a suppressor of selection
Oliver P Hauser, Arne Traulsen, and Martin A Nowak. Heterogeneity in background fitness acts as a suppressor of selection. Journal of theoretical biology , 343:178–185, 2014
2014
-
[9]
Environmental fitness heterogene- ity in the moran process
Kamran Kaveh, Alex McAvoy, and Martin A Nowak. Environmental fitness heterogene- ity in the moran process. Royal Society open science, 6(1):181661, 2019
2019
-
[10]
Genotype by random environmental interactions gives an advantage to non-favored minor alleles
A Mahdipour-Shirayeh, AH Darooneh, AD Long, NL Komarova, and M Kohandel. Genotype by random environmental interactions gives an advantage to non-favored minor alleles. Scientific reports, 7(1):1–8, 2017
2017
-
[11]
The effect of spatial randomness on the average fixation time of mutants
Suzan Farhang-Sardroodi, Amir H Darooneh, Moladad Nikbakht, Natalia L Komarova, and Mohammad Kohandel. The effect of spatial randomness on the average fixation time of mutants. PLoS computational biology, 13(11):e1005864, 2017
2017
-
[12]
Environmental spatial and temporal variability and its role in non-favoured mutant dynamics
Suzan Farhang-Sardroodi, Amir H Darooneh, Mohammad Kohandel, and Natalia L Komarova. Environmental spatial and temporal variability and its role in non-favoured mutant dynamics. Journal of The Royal Society Interface , 16(157):20180781, 2019
2019
-
[13]
Modeling invasion dynamics with spatial random-fitness due to micro-environment
Venkata SK Manem, Kamran Kaveh, Mohammad Kohandel, and Siv Sivaloganathan. Modeling invasion dynamics with spatial random-fitness due to micro-environment. PLoS One, 10(10):e0140234, 2015
2015
-
[14]
Invasion and effective size of graph- structured populations
Stefano Giaimo, Jordi Arranz, and Arne Traulsen. Invasion and effective size of graph- structured populations. PLoS computational biology, 14(11):e1006559, 2018. 20
2018
-
[15]
Environmental evolutionary graph theory
Wes Maciejewski and Gregory J Puleo. Environmental evolutionary graph theory. Jour- nal of theoretical biology, 360:117–128, 2014. 21
2014
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.