Recognition: unknown
Decoding Alignment without Encoding Alignment: A critique of similarity analysis in neuroscience
Pith reviewed 2026-05-08 03:26 UTC · model grok-4.3
The pith
Decoding similarity metrics do not imply similar neural computation because they can be shaped by tiny neuron subsets
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The paper establishes that similarity in decoding representations, as captured by classic alignment metrics such as RSA and DSA, does not entail similarity in function or computation. This holds because decoding geometry is often dominated by non-representative subpopulations, and these metrics remain insensitive to the topology of encoding manifolds that describe how responses are organized across neurons. A causal MNIST demonstration confirms that manipulating encoding structure via training loss leaves decoding metrics intact, while biological and model comparisons reveal that alignment can arise without matching encoding organization.
What carries the argument
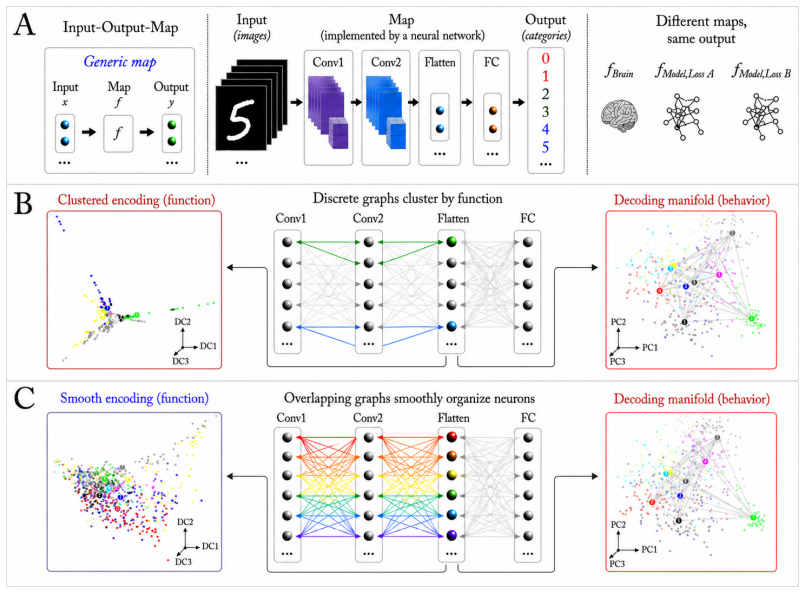
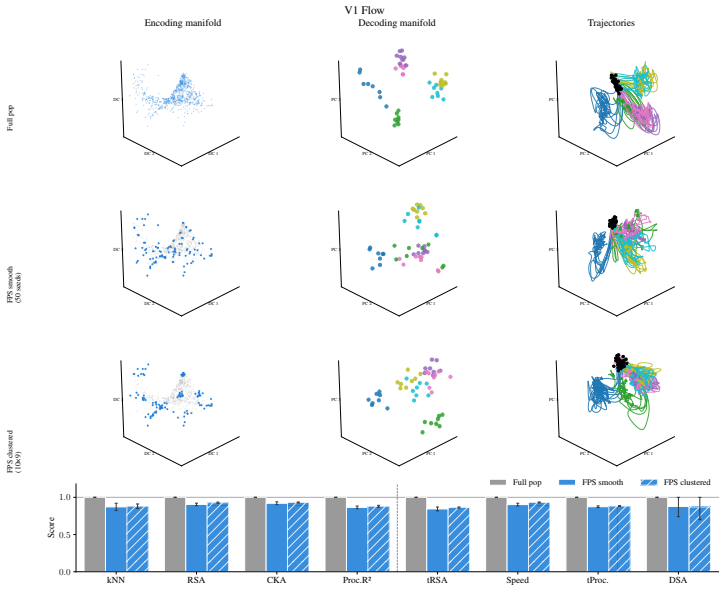
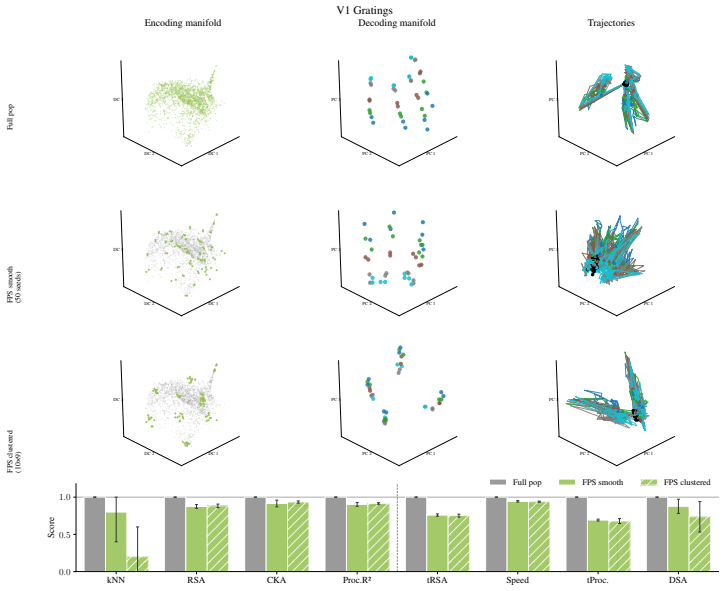
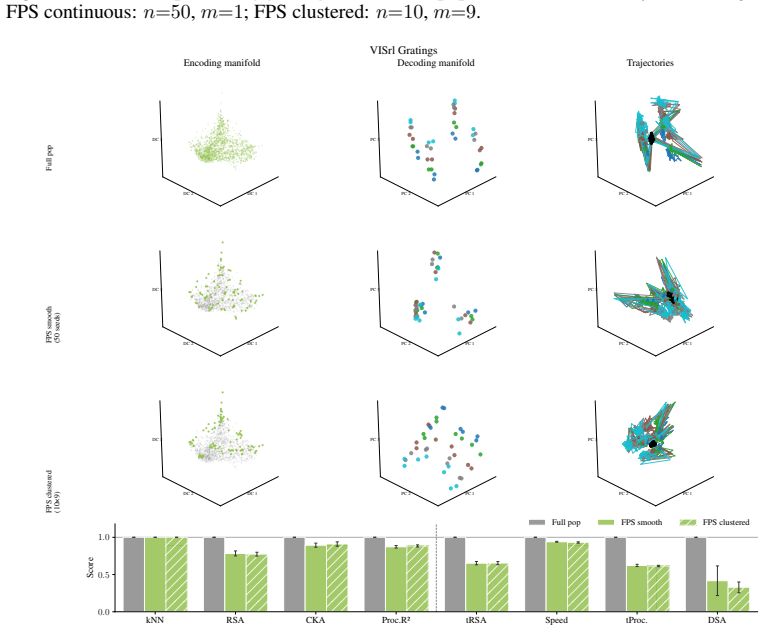
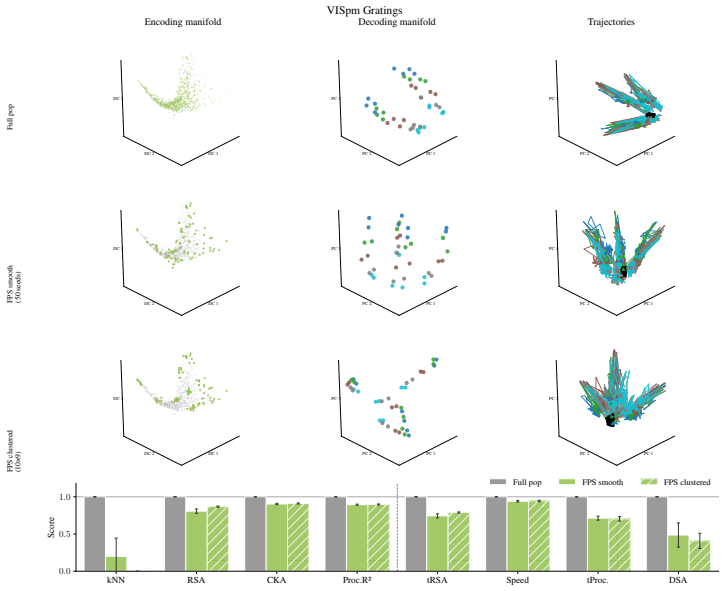
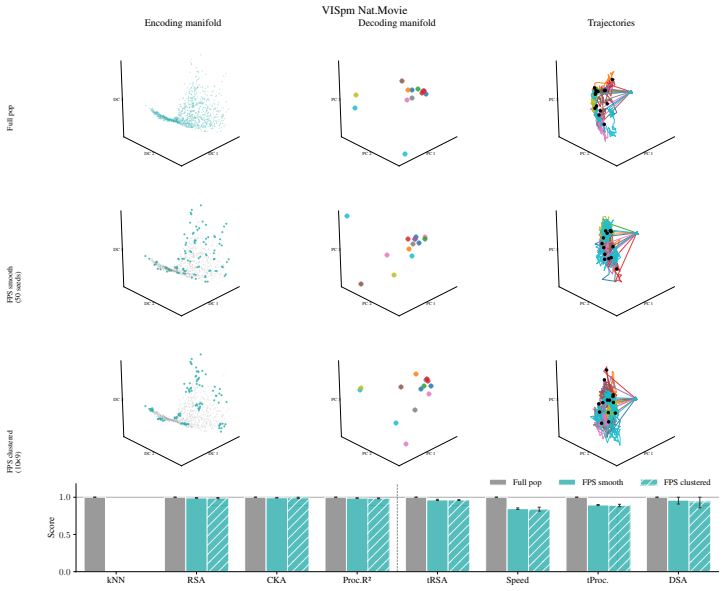
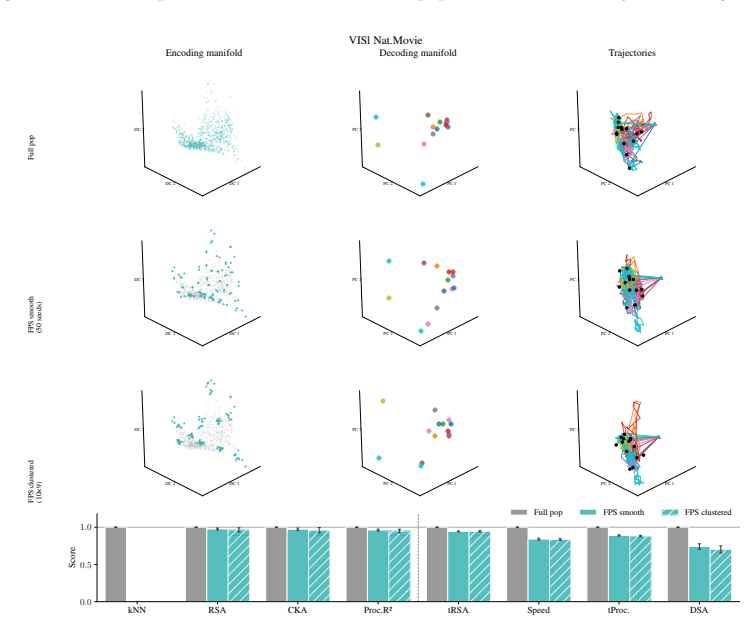
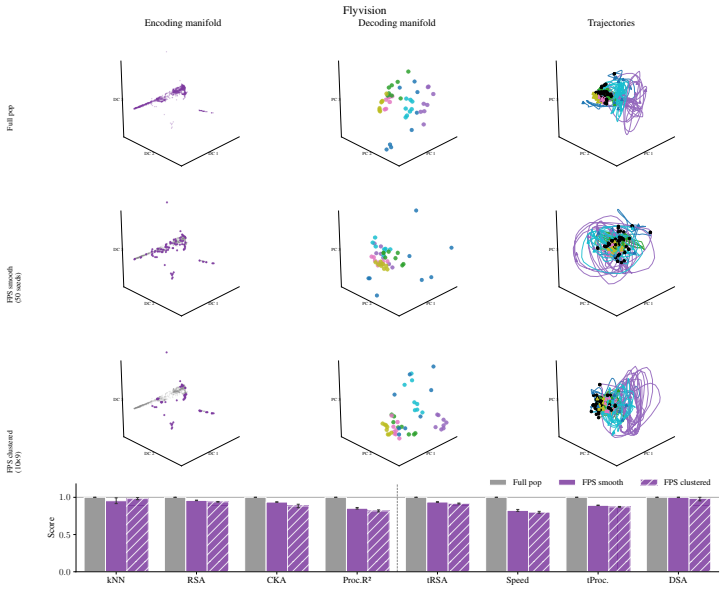
Encoding manifolds, which describe the global organization of neuron responses to stimuli, contrasted with decoding manifolds that capture stimulus geometry independent of population structure.
If this is right
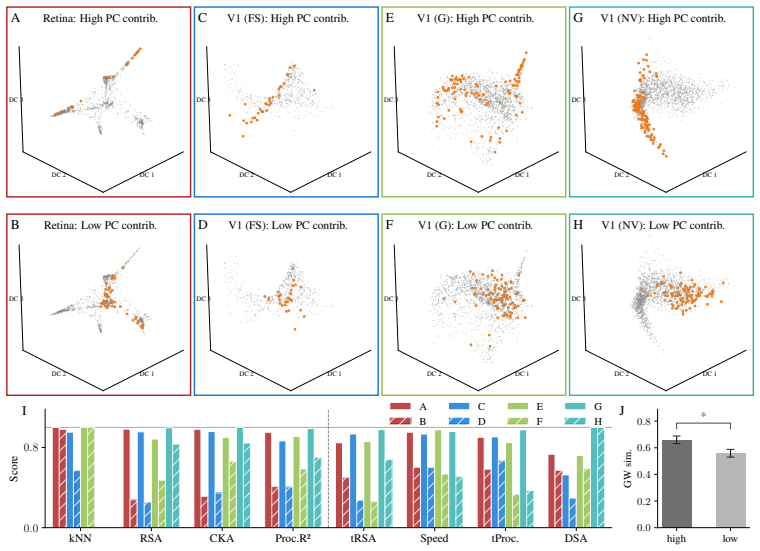
- High alignment scores from RSA or DSA can occur between systems whose neurons are organized differently to encode information.
- Encoding topology serves as a distinguishing signature across biological neural systems that decoding metrics overlook.
- Decoding-based comparisons between brain regions or between brains and models can miss computational differences.
- Encoding manifolds provide a direct way to characterize how a population implements its representations.
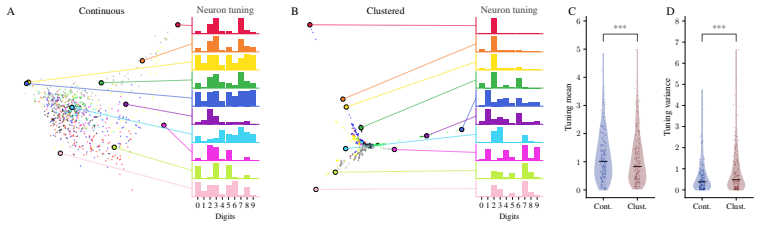
- Causal interventions on encoding structure, such as loss function changes, can leave decoding metrics unchanged.
Where Pith is reading between the lines
- Comparisons between artificial networks and biological circuits may need to incorporate explicit checks on encoding organization rather than relying solely on output geometry.
- Methods that sample or ablate neurons could test whether alignment persists after removing suspected small subpopulations.
- This distinction may help explain cases where neural systems align in decoding yet fail to match in behavioral predictions.
- Future work could develop metrics that jointly assess both decoding geometry and encoding topology.
Load-bearing premise
Decoding geometry reflects the properties of the full neuronal population rather than being controlled by a small subset of neurons.
What would settle it
An experiment in which two systems show identical decoding alignment scores yet differ in measured encoding manifold topology and also differ in a direct functional test, such as generalization behavior or perturbation sensitivity.
Figures















read the original abstract
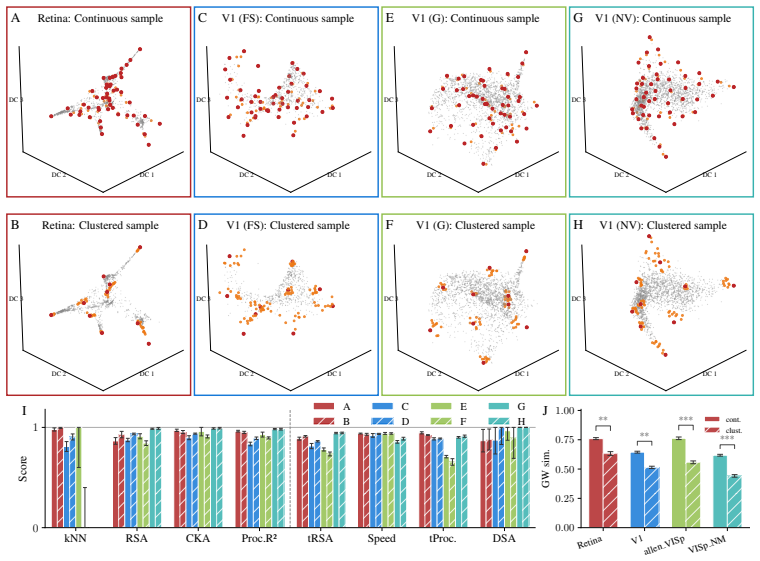
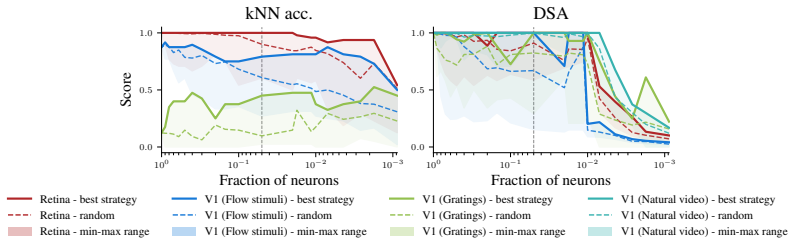
Decoding approaches are widely used in neuroscience and machine learning to compare stimulus representations across neural systems, such as different brain regions, organisms, and deep learning models. Popular methods include decoding (perceptual) manifolds and alignment metrics such as Representational Similarity Analysis (RSA) and Dynamic Similarity Analysis (DSA), where similarity in decoding representations is interpreted as evidence for similar computation. This paper demonstrates a fundamental weakness behind this approach: it is misleading to assume that representational geometry is representative of a neuronal population as a whole, when such representations may actually be shaped by a very small subset of neurons. We show that the complementary encoding paradigm addresses this issue directly: it characterizes how neurons are organized globally in terms of their responses to a set of data, providing insight into how the decoding representation is implemented by neurons within a population. We demonstrate across experiments in biological systems and deep learning models that (i) surprisingly, similar decoding behavior and high representational alignment can arise from small, non-representative subpopulations of neurons; and critically, (ii) alignment metrics are insensitive to encoding manifold topology (how function is distributed across neurons), despite this being a key signature of differentiation across biological systems. A controlled MNIST experiment provides causal evidence: decoding metrics remain unchanged even when encoding topology is causally manipulated via the training loss. Overall, similarity in decoding behavior, as measured by classic alignment metrics, does not imply similarity in function or computation, motivating the use of encoding manifolds as a complementary tool for comparing neural systems.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper claims that decoding-based alignment metrics such as RSA and DSA, widely used to compare representations across neural systems, can produce high similarity scores even when the underlying encoding manifolds differ in how function is distributed across neurons. This occurs because decoding geometry can be dominated by small, non-representative subpopulations. The authors support this via experiments in biological systems, deep learning models, and a causal MNIST manipulation where training loss alters encoding topology without changing decoding alignment scores, concluding that decoding similarity does not imply similar computation and motivating encoding manifolds as a complementary tool.
Significance. If the central claim holds, the work has clear significance for computational neuroscience and ML interpretability by identifying a systematic limitation in popular alignment methods that assume representational geometry reflects the full population. The causal MNIST experiment is a particular strength, as it directly tests insensitivity to encoding topology changes rather than relying on correlational counterexamples. This could shift practice toward hybrid decoding-encoding analyses when comparing systems.
major comments (2)
- [Abstract / Results] The abstract and introduction state that experiments demonstrate claims (i) and (ii), but the manuscript provides no methods section details on datasets, model architectures, neuron selection criteria, statistical tests, or data availability. Without these, the quantitative support for high alignment arising from small subpopulations cannot be evaluated (e.g., what fraction of neurons drives the RSA/DSA scores, and what are the effect sizes?).
- [MNIST experiment] § on the MNIST causal experiment: while the manipulation via training loss is described as leaving decoding metrics unchanged, the specific loss terms, the resulting changes in encoding manifold topology (e.g., via some distance or clustering metric), and the invariance of alignment scores need explicit quantification and controls to confirm causality rather than incidental invariance.
minor comments (2)
- [Introduction] Notation for 'encoding manifold' and 'decoding manifold' should be defined more precisely early on, including how they are computed from population responses, to avoid ambiguity with standard manifold learning terms.
- [Results] The paper would benefit from a table or figure summarizing alignment scores (RSA/DSA values) across the biological, DL, and MNIST cases before/after manipulations for direct comparison.
Simulated Author's Rebuttal
We thank the referee for their positive assessment of the work's significance and for highlighting the strength of the causal MNIST experiment. We agree that additional methodological transparency and quantification are needed to fully support the claims. Below we respond point-by-point to the major comments and commit to a revised manuscript that incorporates the requested details.
read point-by-point responses
-
Referee: [Abstract / Results] The abstract and introduction state that experiments demonstrate claims (i) and (ii), but the manuscript provides no methods section details on datasets, model architectures, neuron selection criteria, statistical tests, or data availability. Without these, the quantitative support for high alignment arising from small subpopulations cannot be evaluated (e.g., what fraction of neurons drives the RSA/DSA scores, and what are the effect sizes?).
Authors: We agree that the current version lacks a dedicated Methods section with the requested details. In the revision we will add a comprehensive Methods section specifying: (1) all datasets (biological recordings with sources, DL model architectures and training procedures), (2) neuron selection criteria and subpopulation identification methods, (3) exact statistical tests and effect-size calculations, and (4) data and code availability. We will also include new supplementary figures/tables that quantify the fraction of neurons driving RSA/DSA scores and report effect sizes for the reported alignment values. These additions will make the quantitative support for claims (i) and (ii) directly evaluable. revision: yes
-
Referee: [MNIST experiment] § on the MNIST causal experiment: while the manipulation via training loss is described as leaving decoding metrics unchanged, the specific loss terms, the resulting changes in encoding manifold topology (e.g., via some distance or clustering metric), and the invariance of alignment scores need explicit quantification and controls to confirm causality rather than incidental invariance.
Authors: We accept that the current description of the MNIST experiment requires more explicit quantification. In the revised manuscript we will: (1) state the precise loss terms and hyperparameters used for the causal manipulation, (2) report quantitative changes in encoding manifold topology using explicit metrics (e.g., clustering coefficients, pairwise distance distributions, or manifold dimensionality estimates), (3) provide statistical tests confirming invariance of RSA/DSA scores across conditions, and (4) add control analyses (e.g., varying manipulation strength and showing that topology changes occur without corresponding decoding shifts). These additions will strengthen the causal interpretation. revision: yes
Circularity Check
No significant circularity
full rationale
The paper's argument is advanced entirely through empirical counterexamples and a controlled causal manipulation (MNIST training-loss intervention that redistributes function across neurons while leaving decoding metrics unchanged). No equations, fitted parameters, or self-referential definitions appear in the provided text; the central claim that high RSA/DSA scores can arise from non-representative subpopulations is demonstrated directly by the reported experiments rather than derived from prior self-citations or ansatzes. The derivation chain is therefore self-contained against external benchmarks and contains no load-bearing steps that reduce to the inputs by construction.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Representational geometry from decoding is assumed to be representative of the entire neuronal population
Reference graph
Works this paper leans on
-
[1]
Ahlert, Jannis and Klein, Thomas and Wichmann, Felix and Geirhos, Robert , year =. How. doi:10.48550/arXiv.2407.07530 , url =. 2407.07530 , eprinttype =
-
[2]
Almudévar, Antonio and Ortega, Alfonso , year =. Bridging. doi:10.48550/arXiv.2601.21568 , url =. 2601.21568 , eprinttype =
-
[3]
The Functional Diversity of Retinal Ganglion Cells in the Mouse , author =. 2016 , date =. doi:10.1038/nature16468 , url =
-
[4]
Learning to Cluster Neuronal Function , author =. 2025 , date =. doi:10.48550/arXiv.2506.03293 , url =. 2506.03293 , eprinttype =
-
[5]
2025 , date =
Fast Dynamical Similarity Analysis , author =. 2025 , date =
2025
-
[6]
Bertram, Johannes and Dyballa, Luciano and Keller, Anderson and Kinger, Savik and Zucker, Steven W. , year =. How '. doi:10.48550/arXiv.2601.21508 , url =. 2601.21508 , eprinttype =
-
[7]
Anderson and Kinger, Savik and Zucker, Steven W
Bertram, Johannes and Dyballa, Luciano and Keller, T. Anderson and Kinger, Savik and Zucker, Steven W. , year =. Manifolds and. doi:10.48550/arXiv.2512.07869 , url =. 2512.07869 , eprinttype =
-
[8]
doi:10.48550/arXiv.2602.16756 , url =
Bertram, Johannes and Geiping, Jonas , year =. doi:10.48550/arXiv.2602.16756 , url =. 2602.16756 , eprinttype =
-
[9]
arXiv preprint arXiv:2411.14633 (2024)
Bo, Yiqing and Soni, Ansh and Srivastava, Sudhanshu and Khosla, Meenakshi , year =. Evaluating. doi:10.48550/arXiv.2411.14633 , url =. 2411.14633 , eprinttype =
-
[10]
and Hong, Ha and Yamins, Daniel L
Cadieu, Charles F. and Hong, Ha and Yamins, Daniel L. K. and Pinto, Nicolas and Ardila, Diego and Solomon, Ethan A. and Majaj, Najib J. and DiCarlo, James J. , year =. Deep. doi:10.1371/journal.pcbi.1003963 , url =
-
[11]
Cain, Jericho , year =. Gauge. doi:10.48550/arXiv.2603.06774 , url =. 2603.06774 , eprinttype =
-
[12]
Current opinion in neurobiology , volume=
Neural population geometry: An approach for understanding biological and artificial neural networks , author=. Current opinion in neurobiology , volume=. 2021 , publisher=
2021
-
[13]
and Sompolinsky, Haim , year =
Chung, SueYeon and Lee, Daniel D. and Sompolinsky, Haim , year =. Classification and. doi:10.1103/PhysRevX.8.031003 , url =. 1710.06487 , eprinttype =
-
[14]
Cloos, Nathan and Li, Moufan and Siegel, Markus and Brincat, Scott L. and Miller, Earl K. and Yang, Guangyu Robert and Cueva, Christopher J. , year =. Differentiable. doi:10.48550/arXiv.2407.07059 , url =. 2407.07059 , eprinttype =
-
[15]
Only Brains Align with Brains: Cross-Region Alignment Patterns Expose Limits of Normative Models
Only Brains Align with Brains: Cross-Region Alignment Patterns Expose Limits of Normative Models , author=. arXiv preprint arXiv:2604.21780 , year=
work page internal anchor Pith review Pith/arXiv arXiv
-
[16]
arXiv preprint arXiv:2307.10246 , year=
Deep neural networks and brain alignment: Brain encoding and decoding (survey) , author=. arXiv preprint arXiv:2307.10246 , year=
-
[17]
Proceedings of the national academy of sciences , volume=
Geometric diffusions as a tool for harmonic analysis and structure definition of data: Diffusion maps , author=. Proceedings of the national academy of sciences , volume=. 2005 , publisher=
2005
-
[18]
Diffusion Maps , author =. 2006 , date =. doi:10.1016/j.acha.2006.04.006 , url =
-
[19]
Reliability of cka as a similarity measure in deep learning.arXiv preprint arXiv:2210.16156(2022)
Davari, MohammadReza and Horoi, Stefan and Natik, Amine and Lajoie, Guillaume and Wolf, Guy and Belilovsky, Eugene , year =. Reliability of. doi:10.48550/arXiv.2210.16156 , url =. 2210.16156 , eprinttype =
-
[20]
A Large-Scale Standardized Physiological Survey Reveals Functional Organization of the Mouse Visual Cortex , author =. 2020 , date =. doi:10.1038/s41593-019-0550-9 , url =
-
[21]
Ding, Frances and Denain, Jean-Stanislas and Steinhardt, Jacob , year =. Grounding. doi:10.48550/arXiv.2108.01661 , url =. 2108.01661 , eprinttype =
-
[22]
Dyballa, Luciano and Zucker, Steven W. , year =. doi:10.1162/neco_a_01566 , url =. 2208.09123 , eprinttype =
-
[23]
Population Encoding of Stimulus Features along the Visual Hierarchy , author =. 2024 , date =. doi:10.1073/pnas.2317773121 , url =
-
[24]
Dynamic Mode Decomposition of Numerical and Experimental Data |
-
[25]
Feghhi, Ebrahim and Hadidi, Nima and Song, Bryan and Blank, Idan A. and Kao, Jonathan C. , year =. What. doi:10.48550/arXiv.2406.01538 , url =. 2406.01538 , eprinttype =
-
[26]
Ghosh, Arna and Chorghay, Zahraa and Bakhtiari, Shahab and Richards, Blake A. , year =. Why All Roads Don't Lead to. doi:10.48550/arXiv.2509.13459 , url =. 2509.13459 , eprinttype =
-
[27]
Harvey, David Lipshutz, and Alex H
Harvey, Sarah E. and Lipshutz, David and Williams, Alex H. , year =. What. doi:10.48550/arXiv.2411.08197 , url =. 2411.08197 , eprinttype =
-
[28]
Houry, Guillaume and Feydy, Jean and Vialard, François-Xavier , year =. Gromov-. doi:10.48550/arXiv.2602.06658 , url =. 2602.06658 , eprinttype =
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2602.06658
-
[29]
Khaligh-Razavi, Seyed-Mahdi and Kriegeskorte, Nikolaus , year =. Deep. doi:10.1371/journal.pcbi.1003915 , url =
-
[30]
Kornblith, Simon and Norouzi, Mohammad and Lee, Honglak and Hinton, Geoffrey , year =. Similarity of. doi:10.48550/arXiv.1905.00414 , url =. 1905.00414 , eprinttype =
-
[31]
Representational Similarity Analysis - Connecting the Branches of Systems Neuroscience , author =. 2008 , date =. doi:10.3389/neuro.06.004.2008 , url =
-
[32]
Connectome-Constrained Networks Predict Neural Activity across the Fly Visual System , author =. 2024 , date =. doi:10.1038/s41586-024-07939-3 , url =
-
[33]
Li, Zeyu Michael and Vu, Hung Anh and Awofisayo, Damilola and Wenger, Emily , year =. Causes and. doi:10.48550/arXiv.2505.13899 , url =. 2505.13899 , eprinttype =
-
[34]
Representation
Li, Xuhong and Grandvalet, Yves and Flamary, Rémi and Courty, Nicolas and Dou, Dejing , abstract =. Representation
-
[35]
Lu, Zitong and Wang, Yile and Golomb, Julie D. , year =. Achieving. doi:10.48550/arXiv.2401.17231 , url =. 2401.17231 , eprinttype =
-
[36]
Mémoli, Facundo , year =. Gromov–. doi:10.1007/s10208-011-9093-5 , url =
-
[37]
Murphy, Alex and Zylberberg, Joel and Fyshe, Alona , year =. Correcting. doi:10.48550/arXiv.2405.01012 , url =. 2405.01012 , eprinttype =
-
[38]
Comparing Noisy Neural Population Dynamics Using Optimal Transport Distances , author =. 2024 , date =. doi:10.48550/arXiv.2412.14421 , url =. 2412.14421 , eprinttype =
-
[39]
Ostrow, Mitchell and Eisen, Adam and Kozachkov, Leo and Fiete, Ila , year =. Beyond. doi:10.48550/arXiv.2306.10168 , url =. 2306.10168 , eprinttype =
-
[40]
Ricardo Rei, Craig Stewart, Ana C Farinha, and Alon Lavie
Peyré, Gabriel and Cuturi, Marco , year =. Computational. doi:10.48550/arXiv.1803.00567 , url =. 1803.00567 , eprinttype =
-
[41]
Sasaki, Masaru and Takeda, Ken and Abe, Kota and Oizumi, Masafumi , year =. Unsupervised. doi:10.1101/2023.09.15.558038 , url =
-
[42]
Majaj, Rishi Rajalingham, Elias B
Schrimpf, Martin and Kubilius, Jonas and Hong, Ha and Majaj, Najib J. and Rajalingham, Rishi and Issa, Elias B. and Kar, Kohitij and Bashivan, Pouya and Prescott-Roy, Jonathan and Geiger, Franziska and Schmidt, Kailyn and Yamins, Daniel L. K. and DiCarlo, James J. , year =. Brain-. doi:10.1101/407007 , url =
-
[43]
Neuron , volume=
How does the brain solve visual object recognition? , author=. Neuron , volume=. 2012 , publisher=
2012
-
[44]
Nature , volume=
Identifying natural images from human brain activity , author=. Nature , volume=. 2008 , publisher=
2008
-
[45]
biorxiv , pages=
Rarely categorical, always high-dimensional: how the neural code changes along the cortical hierarchy , author=. biorxiv , pages=
-
[46]
Current opinion in neurobiology , volume=
On simplicity and complexity in the brave new world of large-scale neuroscience , author=. Current opinion in neurobiology , volume=. 2015 , publisher=
2015
-
[47]
Annual review of neuroscience , volume=
Compressed sensing, sparsity, and dimensionality in neuronal information processing and data analysis , author=. Annual review of neuroscience , volume=. 2012 , publisher=
2012
-
[48]
Cell , volume=
Decoding the brain: From neural representations to mechanistic models , author=. Cell , volume=. 2024 , publisher=
2024
-
[49]
Schrimpf, Martin and Kubilius, Jonas and Lee, Michael J. and Ratan Murty, N. Apurva and Ajemian, Robert and DiCarlo, James J. , year =. Integrative. doi:10.1016/j.neuron.2020.07.040 , url =
-
[50]
Representational Alignment Across Model Layers and Brain Regions with Multi-Level Optimal Transport
Shah, Shaan and Khosla, Meenakshi , year =. Representational. doi:10.48550/arXiv.2510.01706 , url =. 2510.01706 , eprinttype =
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2510.01706
-
[51]
Takahashi, Soh and Sasaki, Masaru and Takeda, Ken and Oizumi, Masafumi , year =. Investigating. doi:10.1016/j.neunet.2025.108222 , url =. 2505.16419 , eprinttype =
-
[52]
Unsupervised Alignment in Neuroscience:
Takeda, Ken and Sasaki, Masaru and Abe, Kota and Oizumi, Masafumi , year =. Unsupervised Alignment in Neuroscience:. doi:10.1016/j.jneumeth.2025.110443 , url =
-
[53]
Aligning Individual Brains with
Thual, Alexis and Tran, Huy and Zemskova, Tatiana and Courty, Nicolas and Flamary, Rémi and Dehaene, Stanislas and Thirion, Bertrand , year =. Aligning Individual Brains with. doi:10.48550/arXiv.2206.09398 , url =. 2206.09398 , eprinttype =
-
[54]
Tran, Du and Wang, Heng and Torresani, Lorenzo and Ray, Jamie and LeCun, Yann and Paluri, Manohar , year =. A. doi:10.48550/arXiv.1711.11248 , url =. 1711.11248 , eprinttype =
-
[55]
Reproducibility of Predictive Networks for Mouse Visual Cortex , author =. 2024 , date =. doi:10.48550/arXiv.2406.12625 , url =. 2406.12625 , eprinttype =
-
[56]
Vaswani, Ashish and Shazeer, Noam and Parmar, Niki and Uszkoreit, Jakob and Jones, Llion and Gomez, Aidan N. and Kaiser, Lukasz and Polosukhin, Illia , year =. Attention. doi:10.48550/arXiv.1706.03762 , url =. 1706.03762 , eprinttype =
work page internal anchor Pith review doi:10.48550/arxiv.1706.03762
-
[57]
Transformer Circuits Thread , year =
Ameisen, Emmanuel and Syed, Maheen and Lindsey, Jack and Tamkin, Alex and Templeton, Adly and Marcus, Jonathan and Conerly, Thomas and Batson, Joshua and Olah, Christopher , title =. Transformer Circuits Thread , year =
-
[58]
Transformer Circuits Thread , year =
Lindsey, Jack and Templeton, Adly and Marcus, Jonathan and Conerly, Thomas and Batson, Joshua and Olah, Christopher , title =. Transformer Circuits Thread , year =
-
[59]
Dunefsky, Jacob and Chlenski, Philippe and Nanda, Neel , year = 2024, month = nov, number =. Transcoders. doi:10.48550/arXiv.2406.11944 , urldate =. arXiv , keywords =:2406.11944 , primaryclass =
-
[60]
arXiv preprint arXiv:2004.11362 , year=
Khosla, Prannay and Teterwak, Piotr and Wang, Chen and Sarna, Aaron and Tian, Yonglong and Isola, Phillip and Maschinot, Aaron and Liu, Ce and Krishnan, Dilip , year = 2021, month = mar, number =. Supervised. doi:10.48550/arXiv.2004.11362 , urldate =. arXiv , keywords =:2004.11362 , primaryclass =
-
[61]
Foundation Model of Neural Activity Predicts Response to New Stimulus Types , author =. 2025 , date =. doi:10.1038/s41586-025-08829-y , url =
-
[62]
Wang, Tao and Goldfeld, Ziv , year =. Neural. doi:10.1109/TIT.2026.3661439) , url =. 2312.07397 , eprinttype =
-
[63]
and Kunz, Erin and Kornblith, Simon and Linderman, Scott W
Williams, Alex H. and Kunz, Erin and Kornblith, Simon and Linderman, Scott W. , year =. Generalized. doi:10.48550/arXiv.2110.14739 , url =. 2110.14739 , eprinttype =
-
[64]
and Kim, Tony Hyun and Wang, Forea and Vyas, Saurabh and Ryu, Stephen I
Williams, Alex H. and Kim, Tony Hyun and Wang, Forea and Vyas, Saurabh and Ryu, Stephen I. and Shenoy, Krishna V. and Schnitzer, Mark and Kolda, Tamara G. and Ganguli, Surya , year =. Unsupervised. doi:10.1016/j.neuron.2018.05.015 , abstract =. 29887338 , eprinttype =
-
[65]
Yang, Yifei and Lee, Wonjun and Zou, Dongmian and Lerman, Gilad , year =. Improving. doi:10.48550/arXiv.2407.10495 , url =. 2407.10495 , eprinttype =
-
[66]
Zhang, Wei and Wang, Zihao and Fan, Jie and Wu, Hao and Zhang, Yong , year =. Fast. doi:10.48550/arXiv.2404.08970 , url =. 2404.08970 , eprinttype =
-
[67]
Zhou, Zikai and Shen, Yunhang and Shao, Shitong and Gong, Linrui and Lin, Shaohui , year =. Rethinking. doi:10.48550/arXiv.2401.11824 , url =. 2401.11824 , eprinttype =
-
[68]
Charles R. Harris and K. Jarrod Millman and St. Array programming with. 2020 , month = sep, journal =. doi:10.1038/s41586-020-2649-2 , publisher =
-
[69]
PyTorch: An Imperative Style, High-Performance Deep Learning Library
Adam Paszke and Sam Gross and Francisco Massa and Adam Lerer and James Bradbury and others , title =. CoRR , volume =. 2019 , url =. 1912.01703 , timestamp =
work page internal anchor Pith review arXiv 2019
-
[70]
and Varoquaux, G
Pedregosa, F. and Varoquaux, G. and Gramfort, A. and Michel, V. and Thirion, B. and others , journal=. Scikit-learn: Machine Learning in
-
[71]
Tenelle A. Wilks and Alan R. Harvey and Jennifer Rodger , title =. Functional Brain Mapping and the Endeavor to Understand the Working Brain , publisher =. 2013 , editor =. doi:10.5772/56491 , url =
-
[72]
and Haberland, Matt and others , title =
Virtanen, Pauli and Gommers, Ralf and Oliphant, Travis E. and Haberland, Matt and others , title =. Nature Methods , year =
-
[73]
Hunter, J. D. , Title =. Computing in Science & Engineering , Volume =
-
[74]
, title =
Plotly Technologies Inc. , title =
-
[75]
Avitan, Itamar and Golan, Tal , year =. Model-. doi:10.48550/arXiv.2510.23321 , url =. 2510.23321 , eprinttype =
-
[76]
Nature neuroscience , volume=
Dimensionality reduction for large-scale neural recordings , author=. Nature neuroscience , volume=. 2014 , publisher=
2014
-
[77]
Apurva Ratan Murty, and Aran Nayebi
Feather, Jenelle and Khosla, Meenakshi and Murty, N. Apurva Ratan and Nayebi, Aran , year =. Brain-. doi:10.48550/arXiv.2502.16238 , url =. 2502.16238 , eprinttype =
-
[78]
and Boisbunon, Aurélie and Chambon, Stanislas and Chapel, Laetitia and Corenflos, Adrien and Fatras, Kilian and Fournier, Nemo and Gautheron, Léo and Gayraud, Nathalie T.H
Flamary, Rémi and Courty, Nicolas and Gramfort, Alexandre and Alaya, Mokhtar Z. and Boisbunon, Aurélie and Chambon, Stanislas and Chapel, Laetitia and Corenflos, Adrien and Fatras, Kilian and Fournier, Nemo and Gautheron, Léo and Gayraud, Nathalie T.H. and Janati, Hicham and Rakotomamonjy, Alain and Redko, Ievgen and Rolet, Antoine and Schutz, Antony and ...
-
[79]
Krämer, Nicholas and others , title =
-
[80]
and Kolda, Tamara G
Bader, Brett W. and Kolda, Tamara G. and others , title =
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.