Recognition: unknown
Mathematical Modeling of Early Embryonic Cell Cycles of Drosophila melanogaster
Pith reviewed 2026-05-08 03:08 UTC · model grok-4.3
The pith
A mathematical model shows that gradually slowing cyclin B synthesis drives the lengthening of the first 14 cell cycles in Drosophila embryos.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
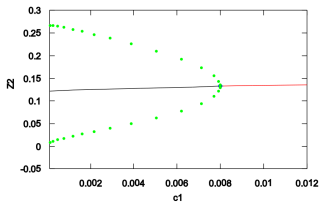
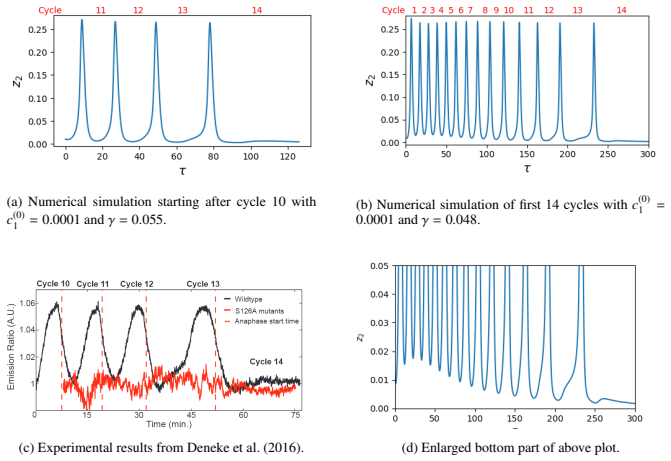
The reduced model exhibits oscillations in a suitable parameter region. When the synthesis rate of cyclin B is made explicitly time-dependent and decreased over successive cycles, the simulated periods lengthen in a manner that qualitatively matches data from Drosophila embryos.
What carries the argument
The reduced ordinary differential equation system for CDK-CycB dynamics, driven by a time-dependent CycB synthesis rate that decreases to produce longer oscillation periods.
If this is right
- The reduced model produces sustained oscillations inside a defined region of parameter space.
- Changes to the activation coefficient of APC alter the oscillatory behavior.
- Imposing a time-dependent decrease in CycB synthesis rate produces progressive period lengthening that matches experimental observations.
- The hypothesis stands only if the chosen time-dependence approximates real biological dynamics.
Where Pith is reading between the lines
- If synthesis slows because maternal stores are depleted, the mechanism would naturally limit the number of rapid divisions before zygotic transcription begins.
- Comparable time-dependent synthesis could be tested in other embryos that undergo rapid cleavage cycles.
- Perturbing cyclin production rates experimentally should shift the timing of period lengthening in predictable ways.
Load-bearing premise
The mathematical function chosen to make the cyclin B synthesis rate decrease over time must reflect actual biological changes rather than being selected only to match the observed period increase.
What would settle it
Direct experimental measurement of cyclin B production rates across the first 14 cycles showing no systematic slowdown would disprove the central hypothesis.
Figures





read the original abstract
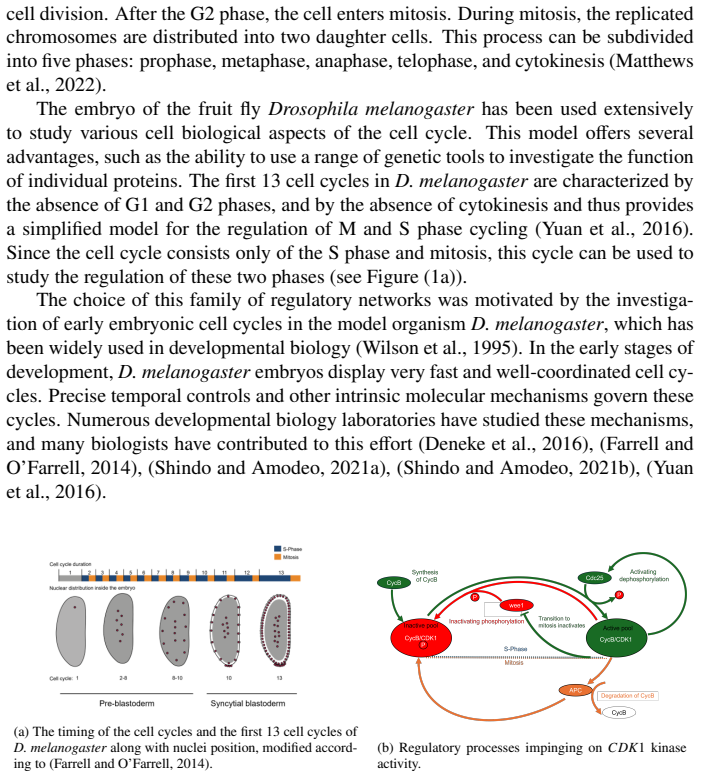
In the early stages of development, Drosophila melanogaster embryos possess very fast and well-coordinated cell cycles. In the cell cycle, CDK activity is essentially regulated by binding CDK and CycB to form an active complex and by phosphorylating CDK via CDC25 and dephosphorylating it via Wee1. We develop a mathematical model for the embryonic cell cycle which is biochemically sound and which can be rigorously analysed after a model reduction. We show that there exists a region in the parameter space where the model describes oscillations. We then focus on the role of two parameters: the CycB synthesis and the activation coefficient of APC. Our main biological hypothesis is that the first one is responsible for the period lengthening over the first 14 cycles which can be experimentally observed and this hypothesis is supported by numerical simulations of our model: if the CycB synthesis is made time-dependent with a prescribed dynamics, then our simulations show qualitatively a very similar behavior to experimental data reported in the literature.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript develops a biochemical model of the early embryonic cell cycles in Drosophila melanogaster, centered on CDK-CycB complex formation and regulation by CDC25 and Wee1. After a model reduction, the authors identify a region of parameter space supporting oscillations. They focus on two parameters—the CycB synthesis rate and the APC activation coefficient—and hypothesize that time-dependent CycB synthesis drives the experimentally observed period lengthening over cycles 1–14. This hypothesis is supported by numerical simulations in which a prescribed time-dependent form for CycB synthesis produces qualitatively similar period progression to literature data.
Significance. If the time-dependent synthesis rate can be placed on independent biological footing, the work would offer a mechanistic account of how synthesis-rate changes contribute to the progressive slowing of early embryonic cycles. The reduction step that permits rigorous identification of oscillatory regimes is a methodological strength, as is the explicit focus on two load-bearing parameters.
major comments (3)
- [numerical simulations of time-dependent synthesis] The central claim that time-dependent CycB synthesis is responsible for period lengthening rests on numerical simulations that impose a specific functional form chosen to reproduce the observed progression. No derivation from measured maternal mRNA levels, transcription timing, or other independent data is supplied, so the agreement is consistent with the hypothesis but does not discriminate it from other tunable mechanisms.
- [model reduction] The abstract states that a model reduction preserves the existence and qualitative properties of oscillations, yet no explicit reduction steps, conserved quantities, or verification that the reduced system retains the same bifurcation structure are provided. This omission is load-bearing for the subsequent parameter-space analysis.
- [comparison with experimental data] The match between simulations and experimental period data is described only as 'qualitatively similar' with no error bars, quantitative distance metrics, or statistical comparison to the literature curves. This weakens the evidential support for the biological hypothesis.
minor comments (2)
- [model equations] Notation for the APC activation coefficient should be defined once and used consistently; its appearance in both the full and reduced models is not cross-referenced.
- [parameter table] The manuscript would benefit from a brief table listing the free parameters retained after reduction and their biological interpretations.
Simulated Author's Rebuttal
We thank the referee for the constructive and detailed review. The comments identify important points that will improve the clarity and rigor of the manuscript. We address each major comment below and indicate the revisions we will make.
read point-by-point responses
-
Referee: [numerical simulations of time-dependent synthesis] The central claim that time-dependent CycB synthesis is responsible for period lengthening rests on numerical simulations that impose a specific functional form chosen to reproduce the observed progression. No derivation from measured maternal mRNA levels, transcription timing, or other independent data is supplied, so the agreement is consistent with the hypothesis but does not discriminate it from other tunable mechanisms.
Authors: We agree that the functional form is a modeling choice selected to test whether a decreasing CycB synthesis rate can produce the observed period lengthening. The simulation serves as a proof-of-principle for the hypothesis rather than a data-driven fit. In the revised manuscript we will state this limitation explicitly, give the exact functional form used, demonstrate robustness across a family of monotonically decreasing forms, and outline feasible experiments (e.g., quantitative maternal mRNA time courses) that could independently constrain the synthesis schedule. revision: partial
-
Referee: [model reduction] The abstract states that a model reduction preserves the existence and qualitative properties of oscillations, yet no explicit reduction steps, conserved quantities, or verification that the reduced system retains the same bifurcation structure are provided. This omission is load-bearing for the subsequent parameter-space analysis.
Authors: The referee correctly identifies that the reduction procedure was not documented in sufficient detail. The reduction exploits two conservation laws (total CDK and total CycB) together with quasi-steady-state approximations for the phosphorylated forms. We will add an appendix that (i) lists the conserved quantities, (ii) derives the reduced ODEs step by step, and (iii) compares the bifurcation diagrams of the full and reduced systems in the two-parameter plane of interest, confirming that the oscillatory region is preserved both qualitatively and quantitatively. revision: yes
-
Referee: [comparison with experimental data] The match between simulations and experimental period data is described only as 'qualitatively similar' with no error bars, quantitative distance metrics, or statistical comparison to the literature curves. This weakens the evidential support for the biological hypothesis.
Authors: We accept that a purely qualitative statement is insufficient. In the revision we will extract the experimental cycle periods from the cited literature, compute the corresponding periods from our simulations, and report quantitative measures including root-mean-square deviation and Pearson correlation. Where the source data report variability we will include error bars; otherwise we will note the limitation while emphasizing the trend agreement. revision: yes
- Derivation of the precise time-dependent functional form for CycB synthesis from independent measurements of maternal mRNA levels or transcription timing.
Circularity Check
Prescribed time-dependent CycB synthesis reproduces observed period lengthening by construction
specific steps
-
fitted input called prediction
[Abstract]
"Our main biological hypothesis is that the first one is responsible for the period lengthening over the first 14 cycles which can be experimentally observed and this hypothesis is supported by numerical simulations of our model: if the CycB synthesis is made time-dependent with a prescribed dynamics, then our simulations show qualitatively a very similar behavior to experimental data reported in the literature."
The period-lengthening behavior is reproduced by prescribing (i.e., fitting) the time-dependence of the CycB synthesis rate; the resulting match to experimental data is therefore a direct consequence of the chosen input rather than an independent prediction or test of the model.
full rationale
The paper develops a biochemically motivated model, reduces it, and identifies a parameter region supporting oscillations. The load-bearing biological claim—that time-dependent CycB synthesis drives the experimentally observed period increase across cycles 1–14—is justified exclusively by numerical simulations in which a specific functional form for the synthesis rate is imposed to produce qualitative agreement with period-progression data. No independent derivation from maternal mRNA levels, transcription timing, or other measurements is supplied; the input is chosen post-hoc to match the target behavior. This matches the 'fitted input called prediction' pattern and yields partial circularity (score 6) while leaving the oscillatory analysis and parameter identification non-circular.
Axiom & Free-Parameter Ledger
free parameters (2)
- CycB synthesis rate function
- activation coefficient of APC
axioms (2)
- domain assumption The biochemical interactions among CDK, CycB, CDC25, Wee1 and APC can be captured by mass-action kinetics.
- ad hoc to paper Model reduction preserves the existence and qualitative properties of oscillations.
Reference graph
Works this paper leans on
-
[1]
The anaphase-promoting complex/cyclosome: APC/C
Acquaviva, C., Pines, J., 2006. The anaphase-promoting complex/cyclosome: APC/C. Journal of cell science 119, 2401–2404
2006
-
[2]
An Introduction to Systems Biology: Design Principles of Biological Circuits
Alon, U., 2019. An Introduction to Systems Biology: Design Principles of Biological Circuits. 2nd ed., Chapman and Hall/CRC, Boca Raton, FL
2019
-
[3]
Detection of multistability, bifurcations, and hysteresis in a large class of biological positive-feedback systems
Angeli, D., Ferrell, J., Sontag, E., 2004. Detection of multistability, bifurcations, and hysteresis in a large class of biological positive-feedback systems. Proc. Natl. Acad. Sci. USA 101, 1822–1827
2004
-
[4]
Algorithms in Real Algebraic Geometry
Basu, S., Pollack, R., Roy, M., 2006. Algorithms in Real Algebraic Geometry. 2nd ed.,
2006
-
[5]
Multi-timescale systems and fast-slow analysis
Bertram, R., Rubin, J., 2017. Multi-timescale systems and fast-slow analysis. Mathe- matical biosciences 287, 105–121
2017
-
[6]
Cdk activation by phosphorylation: link- ing growth signals to cell cycle control
Blank, H., No, E., Polymenis, M., 2025. Cdk activation by phosphorylation: link- ing growth signals to cell cycle control. Biochemical Society Transactions 53, BST20253004
2025
-
[7]
Metabolic cycles in yeast share features conserved among circadian rhythms
Causton, H.C., Feeney, K.A., Ziegler, C.A., O’Neill, J.S., 2015. Metabolic cycles in yeast share features conserved among circadian rhythms. Current Biology 25, 1056– 1062
2015
-
[8]
Waves of Cdk1 activity in S phase synchronize the cell cycle in drosophila embryos
Deneke, V ., Melbinger, A., Vergassola, M., Di Talia, S., 2016. Waves of Cdk1 activity in S phase synchronize the cell cycle in drosophila embryos. Developmental cell 38, 399–412. 18
2016
-
[9]
From egg to gastrula: how the cell cycle is remodeled during the drosophila mid-blastula transition
Farrell, J., O’Farrell, P., 2014. From egg to gastrula: how the cell cycle is remodeled during the drosophila mid-blastula transition. Annual review of genetics 48, 269– 294
2014
-
[10]
Studies of nuclear and cytoplasmic behaviour during the five mitotic cycles that precede gastrulation in drosophila embryogenesis
Foe, V .E., Alberts, B.M., 1983. Studies of nuclear and cytoplasmic behaviour during the five mitotic cycles that precede gastrulation in drosophila embryogenesis. Journal of cell science 61, 31–70
1983
-
[11]
Scaling invariants and symmetry reduction of dynamical systems
Hubert, E., Labahn, G., 2013. Scaling invariants and symmetry reduction of dynamical systems. Foundations of Computational Mathematics 13, 479–516
2013
-
[12]
Interplay between phosphatases and the anaphase- promoting complex/cyclosome in mitosis
Kataria, M., Yamano, H., 2019. Interplay between phosphatases and the anaphase- promoting complex/cyclosome in mitosis. Cells 8, 814
2019
-
[13]
Multiple Time Scale Dynamics
Kuehn, C., 2015. Multiple Time Scale Dynamics. volume 191 ofApplied Mathematical Sciences. Springer, Cham, Switzerland
2015
-
[14]
Regulatory roles of cyclin dependent kinase phosphory- lation in cell cycle control
Lew, D., Kornbluth, S., 1996. Regulatory roles of cyclin dependent kinase phosphory- lation in cell cycle control. Current opinion in cell biology 8, 795–804
1996
-
[15]
Multisite phosphoregulation of Cdc25 activity refines the mitotic entrance and exit switches
Lu, L., Domingo-Sananes, M.R., Huzarska, M., Novak, B., Gould, K., 2012. Multisite phosphoregulation of Cdc25 activity refines the mitotic entrance and exit switches. Proceedings of the National Academy of Sciences 109, 9899–9904
2012
-
[16]
Pseudo-nullclines enable the analysis and prediction of signaling model dynamics
Marrone, J., Sepulchre, J., Ventura, A., 2023. Pseudo-nullclines enable the analysis and prediction of signaling model dynamics. Front. Cell Dev. Biol. 11, 1209589
2023
-
[17]
The Hopf bifurcation and its applications
Marsden, J.E., McCracken, M., 2012. The Hopf bifurcation and its applications. vol- ume 19. Springer Science & Business Media, New York
2012
-
[18]
Cell cycle control in cancer
Matthews, H.K., Bertoli, C., De Bruin, R.A., 2022. Cell cycle control in cancer. Nature reviews Molecular cell biology 23, 74–88
2022
-
[19]
Numerical Methods for Roots of Polynomials
McNamee, J., 2007. Numerical Methods for Roots of Polynomials. volume 2. Elsevier, Amsterdam
2007
-
[20]
Creative blocks: cell-cycle checkpoints and feedback controls
Murray, A., 1992. Creative blocks: cell-cycle checkpoints and feedback controls. Na- ture 359, 599–601
1992
-
[21]
Numerical analysis of a comprehensive model of m-phase control in xenopus oocyte extracts and intact embryos
Novak, B., Tyson, J., 1993. Numerical analysis of a comprehensive model of m-phase control in xenopus oocyte extracts and intact embryos. Journal of cell science 106, 1153–1168. 19
1993
-
[22]
Cyclin-dependent kinases: Masters of the eukaryotic universe
Pluta, A.J., Studniarek, C., Murphy, S., Norbury, C., 2024. Cyclin-dependent kinases: Masters of the eukaryotic universe. Wiley Interdisciplinary Reviews: RNA 15, e1816
2024
-
[23]
Systems-level dissection of the cell-cycle oscillator: bypassing positive feedback produces damped oscillations
Pomerening, J., Kim, S., Ferrell, J., 2005. Systems-level dissection of the cell-cycle oscillator: bypassing positive feedback produces damped oscillations. Cell 122, 565–578
2005
-
[24]
Building a cell cycle oscillator: hys- teresis and bistability in the activation of cdc2
Pomerening, J., Sontag, E., Ferrell Jr., J., 2003. Building a cell cycle oscillator: hys- teresis and bistability in the activation of cdc2. Nature cell biology 5, 346–351
2003
-
[25]
Hopf bifurcations in dynamical systems
Rionero, S., 2019. Hopf bifurcations in dynamical systems. Ricerche di Matematica 68, 811–840
2019
-
[26]
Drosophila wee1 kinase regulates cdk1 and mitotic entry during embryogenesis
Stumpff, J., Duncan, T., Homola, E., Campbell, S., Su, T., 2004. Drosophila wee1 kinase regulates cdk1 and mitotic entry during embryogenesis. Current Biology 14, 2143–2148
2004
-
[27]
Drosophila melanogaster: practical uses in cell and molecular biology
Wilson, L., Matsudaira, P., Goldstein, L., Fyrberg, E., 1995. Drosophila melanogaster: practical uses in cell and molecular biology. Academic Press, San Diego
1995
-
[28]
Timing the Drosophila mid- blastula transition: a cell cycle-centered view
Yuan, K., Seller, C., Shermoen, A., O’Farrell, P., 2016. Timing the Drosophila mid- blastula transition: a cell cycle-centered view. Trends in Genetics 32, 496–507. 20
2016
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.