Recognition: 2 theorem links
· Lean TheoremLearning Cross-Atlas Consistent Brain Disorder Representations via Disentangled Multi-Atlas Functional Connectivity Learning
Pith reviewed 2026-05-11 01:11 UTC · model grok-4.3
The pith
MADCLE disentangles functional connectivity to learn cross-atlas consistent representations of brain disorders.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
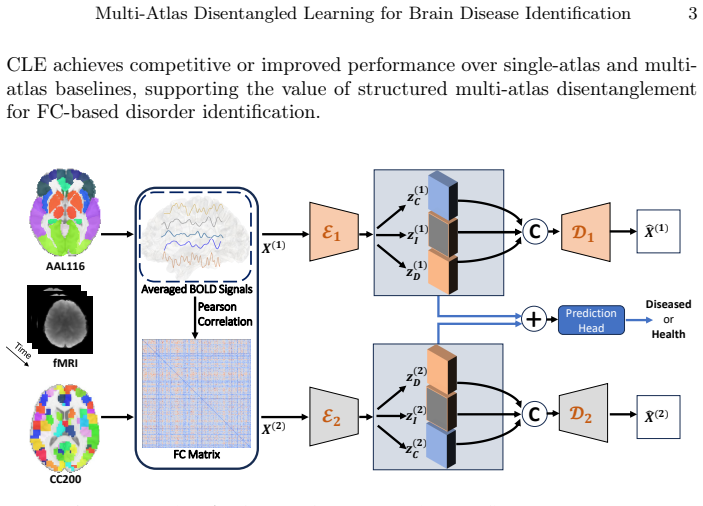
MADCLE jointly encodes FC matrices derived from different brain atlases in parallel branches. Atlas-wise disease-related representations are learned and encouraged to be cross-atlas consistent through distributional alignment. Covariate-related factors receive similarity supervision, atlas-dependent residuals are modeled via atlas-specific reconstruction and decorrelation constraints, and these steps reduce leakage of non-disease or parcellation-specific information into the disease embeddings.
What carries the argument
Multi-branch disentangled representation learning that applies distributional alignment to disease factors while separating covariate and atlas-specific residuals through supervision, reconstruction, and decorrelation.
If this is right
- MADCLE produces competitive or improved performance on disorder identification tasks compared with single-atlas baselines and other multi-atlas GNN or Transformer models.
- Disease-related embeddings become less contaminated by parcellation-specific features.
- The framework supports more stable FC-based disorder classification when multiple heterogeneous atlases are available.
- Structured disentanglement offers an alternative to shallow feature fusion across atlases.
Where Pith is reading between the lines
- The method could be applied to other neuroimaging modalities or additional clinical cohorts to test broader robustness.
- In applied settings it might reduce the practical impact of choosing one particular brain atlas over another.
- Direct measurement of alignment quality between disease representations from different atlases on new datasets would provide further validation.
Load-bearing premise
The combination of distributional alignment, covariate similarity supervision, atlas-specific reconstruction, and decorrelation successfully isolates disease signals without losing useful information or introducing alignment artifacts.
What would settle it
Training the model without the distributional alignment or decorrelation terms and observing whether classification accuracy on ADNI or ADHD-200 drops or cross-atlas consistency metrics fail to improve.
Figures
read the original abstract
Functional connectivity (FC) derived from resting-state fMRI is widely used to characterize large-scale brain network alterations in neurological and psychiatric disorders. However, FC construction critically depends on the choice of brain atlas, and different parcellations may emphasize distinct organizational features, leading to heterogeneous and sometimes inconsistent representations. Existing multi-atlas approaches partially alleviate this issue but often fuse atlas-derived features or predictions at a relatively shallow level, while single-atlas disentanglement methods do not explicitly address cross-atlas heterogeneity. We propose Multi-Atlas Disentangled Connectivity LEarning (MADCLE), a multi-branch representation learning framework that jointly encodes FC matrices derived from different brain atlases. Rather than introducing a single explicitly shared latent variable across parcellations, MADCLE learns atlas-wise disease-related representations and encourages them to be cross-atlas consistent through distributional alignment. Meanwhile, covariate-related and atlas-dependent residual factors are modeled separately using covariate similarity supervision, atlas-specific reconstruction, and decorrelation constraints, thereby reducing the leakage of non-disease and parcellation-dependent information into the disease-related embeddings. Experiments on the ADNI and ADHD-200 datasets suggest that MADCLE achieves competitive or improved performance compared with single-atlas baselines, multi-atlas GNN/Transformer models, and recent multi-atlas consistency frameworks. These results support the potential value of structured disentanglement for FC-based disorder identification under heterogeneous parcellation schemes.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces MADCLE, a multi-branch representation learning framework for functional connectivity (FC) matrices from multiple brain atlases in neurological and psychiatric disorder identification. It learns atlas-wise disease-related embeddings encouraged to be cross-atlas consistent via distributional alignment, while separately modeling covariate-related factors (via similarity supervision) and atlas-dependent residuals (via atlas-specific reconstruction and decorrelation constraints) to reduce non-disease and parcellation leakage. Experiments on the ADNI and ADHD-200 datasets are reported to show competitive or improved performance relative to single-atlas baselines, multi-atlas GNN/Transformer models, and recent multi-atlas consistency methods.
Significance. If the disentanglement successfully isolates consistent disease signals without discarding useful variance or introducing alignment artifacts, the framework could meaningfully improve robustness of FC-based disorder classification under heterogeneous parcellation schemes, a persistent issue in rs-fMRI analysis. The structured separation of factors is a principled design choice that goes beyond shallow fusion approaches.
major comments (2)
- [Abstract] Abstract and Experiments: The headline claim of competitive or improved performance is presented without any reported statistical testing, error bars, cross-validation details, or ablation studies isolating the contributions of distributional alignment, covariate supervision, reconstruction, and decorrelation. This leaves the attribution of gains to the proposed disentanglement unsupported by verifiable evidence.
- [Method] Method and Experiments: Distributional alignment of atlas-wise disease representations is central to the consistency claim, yet no direct checks (e.g., cross-atlas agreement on disease predictions, embedding similarity metrics for the disease factors, or analysis of retained disease variance) are described. Downstream accuracy alone cannot confirm that alignment captures shared disease mechanisms rather than marginal distributions, particularly when atlases emphasize different scales.
minor comments (1)
- The abstract and method description would benefit from explicit notation for the alignment loss (e.g., MMD or adversarial) and the weighting hyperparameters of the combined objective to aid reproducibility.
Simulated Author's Rebuttal
We thank the referee for the constructive feedback. We address each major comment point by point below and have revised the manuscript to incorporate additional analyses and details where the concerns are valid.
read point-by-point responses
-
Referee: [Abstract] Abstract and Experiments: The headline claim of competitive or improved performance is presented without any reported statistical testing, error bars, cross-validation details, or ablation studies isolating the contributions of distributional alignment, covariate supervision, reconstruction, and decorrelation. This leaves the attribution of gains to the proposed disentanglement unsupported by verifiable evidence.
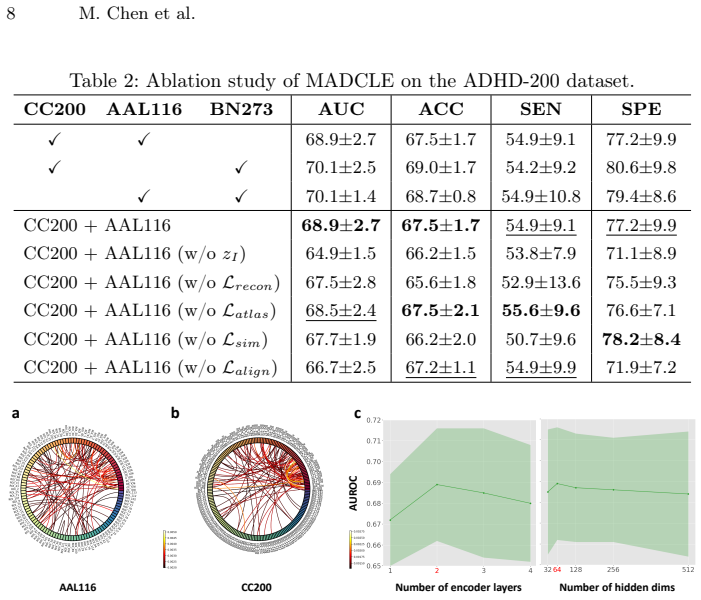
Authors: We agree that the original manuscript lacked explicit statistical testing, error bars, cross-validation details, and component-wise ablations, which weakens the support for attributing performance gains specifically to the disentanglement components. In the revised version, we have added 5-fold cross-validation results with standard error bars across all reported metrics, paired t-tests comparing MADCLE against baselines (with p-values), and ablation studies that isolate the effect of each term (distributional alignment, covariate similarity supervision, atlas-specific reconstruction, and decorrelation constraints). These are now included in the Experiments section with updated tables and figures, and the abstract has been revised to reference the cross-validation and statistical evaluation. This provides the requested verifiable evidence. revision: yes
-
Referee: [Method] Method and Experiments: Distributional alignment of atlas-wise disease representations is central to the consistency claim, yet no direct checks (e.g., cross-atlas agreement on disease predictions, embedding similarity metrics for the disease factors, or analysis of retained disease variance) are described. Downstream accuracy alone cannot confirm that alignment captures shared disease mechanisms rather than marginal distributions, particularly when atlases emphasize different scales.
Authors: We acknowledge that relying solely on downstream accuracy is insufficient to confirm the alignment isolates shared disease mechanisms. In the revised manuscript, we have added direct verification analyses in the Experiments section: (1) cross-atlas agreement on disease predictions derived from the disease-related embeddings, (2) cosine similarity and distributional distance metrics between atlas-wise disease factors, and (3) retained disease variance assessment via performance comparison with and without the alignment term. These results indicate improved cross-atlas consistency while preserving task-relevant variance, supporting that the alignment targets shared mechanisms beyond marginal distributions. We have also clarified the design rationale in the Method section to address scale differences across atlases. revision: yes
Circularity Check
No significant circularity; new multi-branch architecture and losses validated externally
full rationale
The paper introduces MADCLE as a novel multi-branch representation learning framework with distributional alignment, covariate supervision, reconstruction, and decorrelation losses to achieve cross-atlas consistent disease representations from FC matrices. The central claims rest on empirical performance comparisons against baselines on the independent ADNI and ADHD-200 datasets rather than any self-referential derivation, fitted parameter renamed as prediction, or load-bearing self-citation chain. No equations or steps in the provided description reduce the reported results to inputs by construction, and the method's outputs are tested on external data without internal tautology.
Axiom & Free-Parameter Ledger
free parameters (2)
- loss weights for alignment, reconstruction, and decorrelation terms
- network architecture hyperparameters (depth, width, learning rate)
axioms (2)
- domain assumption Disease-related signals in functional connectivity are sufficiently shared across different brain atlases to permit distributional alignment.
- domain assumption Covariate and atlas-specific factors can be isolated via similarity supervision and reconstruction without interfering with disease representations.
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearMADCLE learns atlas-wise disease-related representations and encourages them to be cross-atlas consistent through distributional alignment... covariate similarity supervision, atlas-specific reconstruction, and decorrelation constraints
-
IndisputableMonolith/Foundation/RealityFromDistinction.leanreality_from_one_distinction unclearExperiments on the ADNI and ADHD-200 datasets suggest that MADCLE achieves competitive or improved performance
Reference graph
Works this paper leans on
-
[1]
In: International Conference on Medical Image Computing and Computer-Assisted Intervention
Chen, M., Yu, X., Zhang, J., Chen, T., Cao, C., Zhuang, Y., Lyu, Y., Zhang, L., Liu, T., Zhu, D.: Core-periphery principle guided state space model for func- tional connectome classification. In: International Conference on Medical Image Computing and Computer-Assisted Intervention. pp. 236–246. Springer (2025)
2025
-
[2]
Chu, Y., Wang, G., Cao, L., Qiao, L., Liu, M.: Multi-scale graph representation learningforautismidentificationwithfunctionalmri.FrontiersinNeuroinformatics 15, 802305 (2022)
2022
-
[3]
Human brain mapping33(8), 1914–1928 (2012)
Craddock, R.C., James, G.A., Holtzheimer III, P.E., Hu, X.P., Mayberg, H.S.: A whole brain fmri atlas generated via spatially constrained spectral clustering. Human brain mapping33(8), 1914–1928 (2012)
1914
-
[4]
Neuroimage31(3), 968–980 (2006)
Desikan, R.S., Ségonne, F., Fischl, B., Quinn, B.T., Dickerson, B.C., Blacker, D., Buckner, R.L., Dale, A.M., Maguire, R.P., Hyman, B.T., et al.: An automated labeling system for subdividing the human cerebral cortex on mri scans into gyral based regions of interest. Neuroimage31(3), 968–980 (2006)
2006
-
[5]
Cerebral cortex26(8), 3508–3526 (2016)
Fan, L., Li, H., Zhuo, J., Zhang, Y., Wang, J., Chen, L., Yang, Z., Chu, C., Xie, S., Laird, A.R., et al.: The human brainnetome atlas: a new brain atlas based on connectional architecture. Cerebral cortex26(8), 3508–3526 (2016)
2016
-
[6]
Proceedings of the National Academy of Sciences102(27), 9673– 9678 (2005)
Fox, M.D., Snyder, A.Z., Vincent, J.L., Corbetta, M., Van Essen, D.C., Raichle, M.E.: The human brain is intrinsically organized into dynamic, anticorrelated func- tional networks. Proceedings of the National Academy of Sciences102(27), 9673– 9678 (2005)
2005
-
[7]
In: 2025 IEEE 22nd International Symposium on Biomedical Imaging (ISBI)
Han, K., Hu, D., Cheng, J., Liu, T., Bozoki, A., Zhu, D., Li, G.: Dual multi- atlas representation alignment for brain disorder diagnosis using morphological connectome. In: 2025 IEEE 22nd International Symposium on Biomedical Imaging (ISBI). pp. 1–4. IEEE (2025)
2025
-
[8]
Advances in Neural Information Processing Systems35, 25586–25599 (2022)
Kan, X., Dai, W., Cui, H., Zhang, Z., Guo, Y., Yang, C.: Brain network trans- former. Advances in Neural Information Processing Systems35, 25586–25599 (2022)
2022
-
[9]
NeuroImage146, 1038– 1049 (2017)
Kawahara, J., Brown, C.J., Miller, S.P., Booth, B.G., Chau, V., Grunau, R.E., Zwicker, J.G., Hamarneh, G.: Brainnetcnn: Convolutional neural networks for brain networks; towards predicting neurodevelopment. NeuroImage146, 1038– 1049 (2017)
2017
-
[10]
In: International Workshop on Machine Learning in Clinical Neuroimaging
Mahler, L., Wang, Q., Steiglechner, J., Birk, F., Heczko, S., Scheffler, K., Lohmann, G.: Pretraining is all you need: A multi-atlas enhanced transformer framework for autism spectrum disorder classification. In: International Workshop on Machine Learning in Clinical Neuroimaging. pp. 123–132. Springer (2023)
2023
-
[11]
Human brain mapping 35(10), 5052–5070 (2014)
Min, R., Wu, G., Cheng, J., Wang, Q., Shen, D., Initiative, A.D.N.: Multi-atlas based representations for alzheimer’s disease diagnosis. Human brain mapping 35(10), 5052–5070 (2014)
2014
-
[12]
Neuroimage15(1), 273–289 (2002)
Tzourio-Mazoyer, N., Landeau, B., Papathanassiou, D., Crivello, F., Etard, O., Delcroix, N., Mazoyer, B., Joliot, M.: Automated anatomical labeling of activations in spm using a macroscopic anatomical parcellation of the mni mri single-subject brain. Neuroimage15(1), 273–289 (2002)
2002
-
[13]
Human Brain Mapping46(5), e70206 (2025) Multi-Atlas Disentangled Learning for Brain Disease Identification 11
Wu, X., Liang, C., Bustillo, J., Kochunov, P., Wen, X., Sui, J., Jiang, R., Yang, X., Fu, Z., Zhang, D., et al.: The impact of atlas parcellation on functional connectivity analysis across six psychiatric disorders. Human Brain Mapping46(5), e70206 (2025) Multi-Atlas Disentangled Learning for Brain Disease Identification 11
2025
-
[14]
IEEE Journal of Biomedical and Health Informatics pp
Xu, J., Lan, M., Dong, X., He, K., Zhang, W., Bian, Q., Ke, Y.: Multi-atlas brain network classification through consistency distillation and complementary informa- tion fusion. IEEE Journal of Biomedical and Health Informatics pp. 1–12 (2025)
2025
-
[15]
In: International Conference on Medical Image Computing and Computer-Assisted Intervention
Zhang, S., Jiang, Z., Shen, X., Yu, Z., Chen, X., Zhang, X.Y., Zhou, Y.: Graph disentanglement learning for fmri analysis: Decoupling disease, covariates, and in- dividual variability. In: International Conference on Medical Image Computing and Computer-Assisted Intervention. pp. 352–361. Springer (2025)
2025
-
[16]
Human brain mapping35(7), 2911–2923 (2014)
Zhu, D., Li, K., Terry, D.P., Puente, A.N., Wang, L., Shen, D., Miller, L.S., Liu, T.: Connectome-scale assessments of structural and functional connectivity in mci. Human brain mapping35(7), 2911–2923 (2014)
2014
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.