Recognition: 3 theorem links
· Lean TheoremGenetic Information as a "Chord" of Chemical Oscillations: Emergence of Catalyst-RNA Systems Driven by Superposed Rhythms
Pith reviewed 2026-05-11 01:17 UTC · model grok-4.3
The pith
Chemical oscillations can bias binary polymer growth to establish catalytic loops and record functional sequences
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
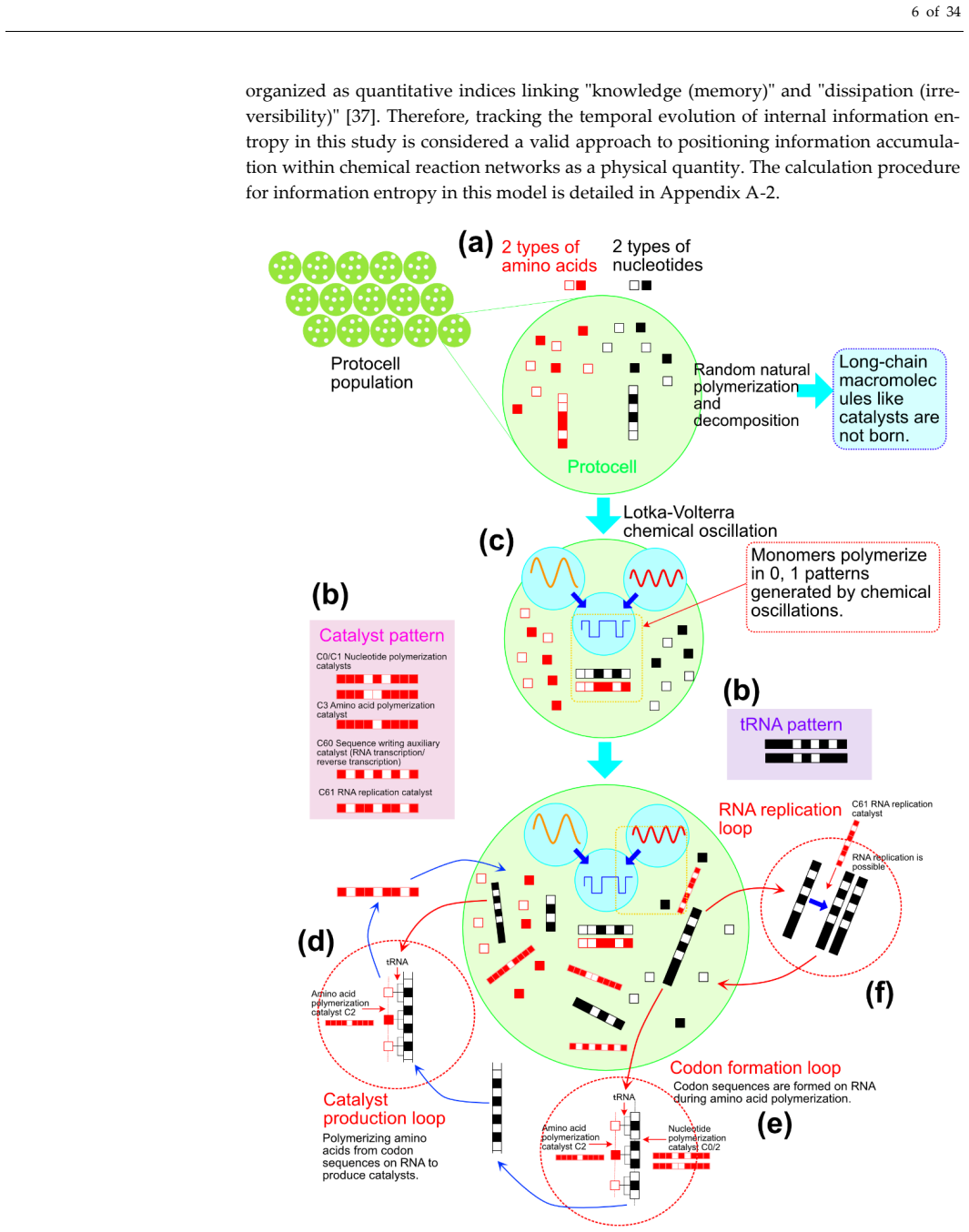
The central discovery is that superposing rhythms from two Lotka-Volterra oscillators during polymer elongation provides a temporal bias for 0/1 selection that enables the formation of interdependent catalyst-RNA systems, including catalytic loops, primordial tRNAs, and nucleic acids that record and amplify them, outperforming random models in loop establishment, functional molecule accumulation, polymer length, and entropy reduction.
What carries the argument
The synthesis of internal oscillations that supplies a temporal bias for monomer selection in polymer elongation, combined with mechanisms for protecting, recording, and re-amplifying functional sequences.
Load-bearing premise
That the internal oscillations generate a dependable temporal signal for biasing sequence choices and that the simplified rules suffice to protect and replicate functional sequences without degradation.
What would settle it
A direct comparison in which the oscillation bias is turned off but all other mechanisms like sequence protection and re-amplification are retained; if the catalytic loop formation rate then equals that of the random model, the role of the rhythms would be falsified.
Figures
read the original abstract
A central challenge in the origin of life is understanding how catalytic peptide-like polymers and information-bearing nucleic acid-like polymers emerged as an interde-pendent system. This study constructs a primordial cognitive model incorporating two internal Lotka-Volterra chemical oscillators to investigate, through simulation, whether a catalytic loop, primordial tRNAs, and nucleic acids that record and amplify them, can form through the interaction of polymers represented by binary (0/1) sequences. In this model, a mechanism was introduced where the synthesis of internal oscillations pro-vides a temporal bias for 0/1 selection during polymer elongation, while generated functional sequences are protected, recorded, and re-amplified. Simulation results demonstrated that the proposed cognitive model significantly outperformed a contrast model based on random 0/1 selection in terms of the establishment rate of catalytic loops, the accumulation of functional molecules, polymer elongation, and the reduction of Shannon entropy in sequence distribution. Furthermore, this superiority was generally maintained across sensitivity analyses, including batch calculations with different ran-dom seeds. While this study is a computational model based on abstract binary se-quences and simplified translation/replication rules rather than a direct reconstruction of life's origin, it provides a working hypothesis for the interdependent emergence of catalytic function and information retention by demonstrating that internal oscillations can bias sequence exploration within a framework linking autocatalytic networks, re-cording, and group selection. Future research must verify the generality and empirical validity of this framework by expanding monomer types, evolving into multi-oscillator systems, and establishing correspondences with compartmentalized experimental sys-tems.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proposes a primordial cognitive model incorporating two internal Lotka-Volterra chemical oscillators that supply a temporal bias for 0/1 selection during elongation of binary-sequence polymers. Functional sequences receive explicit protection, recording, and re-amplification under simplified translation/replication rules. Simulations show the model outperforms a random 0/1-selection contrast model in catalytic-loop establishment rate, functional-molecule accumulation, polymer elongation, and Shannon-entropy reduction in sequence distributions; the advantage persists across sensitivity runs with varied random seeds. The work offers a working hypothesis for the co-emergence of catalytic function and information retention via superposed rhythms.
Significance. If the reported simulation outcomes prove robust, the paper supplies a concrete computational framework that couples autocatalytic networks, internal oscillations, and explicit recording/amplification steps, thereby generating a falsifiable hypothesis for how informational and catalytic polymers could become interdependent. The systematic sensitivity analyses with multiple random seeds constitute a methodological strength that supports reproducibility within the model's abstract setting.
major comments (2)
- Abstract and Results sections: the central claim of statistically significant outperformance is presented without quantitative parameter values (Lotka-Volterra rates, bias strength, protection/amplification rates), without reported effect sizes, and without description of the statistical tests used to compare the oscillator model against the random baseline. Because the headline result rests entirely on these simulation outcomes, the absence of these details prevents independent verification of robustness.
- Model Description and Contrast Model paragraph: it is not stated whether the random 0/1-selection contrast model receives the same protection, recording, and re-amplification rules applied to functional sequences in the oscillator model. If protection is withheld from the contrast, the reported gains in loop formation and entropy reduction cannot be attributed specifically to the temporal bias supplied by the superposed Lotka-Volterra oscillators rather than to the protection mechanism itself.
minor comments (2)
- Abstract: typographical line-break artifacts appear (interde-pendent, pro-vides, se-quences, re-cording); these should be removed.
- Methods/Implementation: the exact discrete-time update rules for the two Lotka-Volterra oscillators and the precise mapping from oscillator phase to 0/1 selection probability are not supplied; adding the governing equations or pseudocode would improve reproducibility.
Simulated Author's Rebuttal
We thank the referee for the constructive and detailed comments, which highlight important areas for improving the clarity, completeness, and verifiability of our manuscript. We address each major comment point by point below and will make the necessary revisions to strengthen the presentation of our simulation results and model description.
read point-by-point responses
-
Referee: Abstract and Results sections: the central claim of statistically significant outperformance is presented without quantitative parameter values (Lotka-Volterra rates, bias strength, protection/amplification rates), without reported effect sizes, and without description of the statistical tests used to compare the oscillator model against the random baseline. Because the headline result rests entirely on these simulation outcomes, the absence of these details prevents independent verification of robustness.
Authors: We agree that the abstract and results sections would benefit from greater quantitative detail to support independent verification. The specific Lotka-Volterra rates, bias strength, and protection/amplification rates are defined in the Methods section, along with the simulation parameters. We will revise the abstract and results to explicitly summarize these values, report effect sizes for the key performance metrics (catalytic loop establishment, functional molecule accumulation, polymer length, and entropy reduction), and describe the statistical tests employed (including the type of test, sample sizes from the sensitivity runs, and any multiple-comparison corrections). These additions will be incorporated without altering the core findings. revision: yes
-
Referee: Model Description and Contrast Model paragraph: it is not stated whether the random 0/1-selection contrast model receives the same protection, recording, and re-amplification rules applied to functional sequences in the oscillator model. If protection is withheld from the contrast, the reported gains in loop formation and entropy reduction cannot be attributed specifically to the temporal bias supplied by the superposed Lotka-Volterra oscillators rather than to the protection mechanism itself.
Authors: We apologize for the lack of explicit statement on this point. The random 0/1-selection contrast model applies identical protection, recording, and re-amplification rules to functional sequences; the sole difference is the absence of the temporal bias from the superposed Lotka-Volterra oscillators during the 0/1 selection step in polymer elongation. We will revise the Model Description and Contrast Model paragraph to state this equivalence clearly, thereby ensuring that the reported advantages are attributable specifically to the oscillatory bias rather than the protection mechanism. revision: yes
Circularity Check
No circularity: simulation results compared to independent random control
full rationale
The paper describes a simulation in which two Lotka-Volterra oscillators supply a temporal bias for 0/1 monomer selection during polymer elongation, while separately specifying protection, recording, and re-amplification rules for functional sequences. Outcomes are evaluated against an explicit contrast model that performs random 0/1 selection. No equations, fitted parameters, or self-citations are invoked that would make the reported superiority (catalytic-loop establishment, entropy reduction, etc.) equivalent to the model's own inputs by construction. The central claim therefore rests on an external benchmark rather than on any definitional or self-referential reduction.
Axiom & Free-Parameter Ledger
free parameters (3)
- Lotka-Volterra oscillator parameters
- selection bias strength
- protection and amplification rates
axioms (2)
- domain assumption Binary (0/1) sequences can represent primordial polymers with catalytic and informational properties under simplified translation/replication rules.
- domain assumption Internal chemical oscillations can be treated as independent Lotka-Volterra systems whose phases provide a temporal selection bias.
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel uncleartwo internal Lotka-Volterra chemical oscillators... temporal bias for 0/1 selection during polymer elongation, while generated functional sequences are protected, recorded, and re-amplified
-
IndisputableMonolith/Foundation/ArithmeticFromLogic.leanLogicNat induction and embed_strictMono unclearShannon entropy of 9-bit sub-sequences... reduction of Shannon entropy in sequence distribution
-
IndisputableMonolith/Foundation/AbsoluteFloorClosure.leanreality_from_one_distinction unclearsuccess rate of catalytic loop emergence... across random seeds and target sequences
Reference graph
Works this paper leans on
-
[1]
Gilbert, W. Origin of life: The RNA world. Nature 1986, 319, 618 . https://doi.org/10.1038/319618a0 https://doi.org/10.1038/319618a0
-
[2]
The RNA World: molecular cooperation at the origins of life
Higgs, P.; Lehman, N. The RNA World: molecular cooperation at the origins of life. Nat Rev Genet 2015, 16, 7 –17. doi:10.1038/nrg3841. https://doi.org/10.1038/nrg3841 tRNA Hamming Distance Weights 0.005 0.01 (Ref) 0.02 0.05 Cognitive Model 93.3 % 83.3 % 73.3 % 70.0 % Number of Protocells 10 20 50 (Ref) 80 120 Cognitive Model 66.7 % 93.3 % 83.3 % 83.3 % 70...
-
[3]
Kruger, K.; Grabowski, PJ.; Zaug AJ.; Sands J.; Gottschling DE.; Cech TR.; Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell, 1982,31(1):147-157. doi:10.1016/0092-8674(82)90414-7. https://doi.org/10.1016/0092-8674(82)90414-7
-
[4]
Cell, 1983, 35(3 Pt 2), 849-857
Guerrier-Takada C.; Gardiner K.; Marsh T.; Pace N.; Altman S., The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell, 1983, 35(3 Pt 2), 849-857. doi:10.1016/0092-8674(83)90117-4. https://doi.org/10.1016/0092-8674(83)90117-4
-
[5]
Development, Growth and Differentiation, 2023, 65(3), 167-174
Tagami S.; Li P.; The origin of life: RNA and protein co -evolution on the ancient Earth. Development, Growth and Differentiation, 2023, 65(3), 167-174. doi:10.1111/dgd.12845. https://doi.org/10.1111/dgd.12845
-
[6]
Journal of Systematics and Evolution
Saad NY., A ribonucleopeptide world at the origin of life. Journal of Systematics and Evolution. 2018, 56(1), 1-13. doi:10.1111/jse.12287 https://doi.org/10.1111/jse.12287
-
[7]
Ikehara, K. Evolutionary Steps in the Emergence of Life Deduced from the Bottom -Up Approach and GADV Hypothesis (Top - Down Approach). Life 2016, 6, 6. https://doi.org/10.3390/life6010006
-
[8]
Ikehara, K., Towards Revealing the Origin of Life: Presenting the GADV Hypothesis, Springer, 2021, ISBN : 978-3030710866
work page 2021
-
[9]
Proceedings of the National Academy of Sciences of the United States of America
Schimmel P.; Giegé R.; Moras D.; Yokoyama S., An operational RNA code for amino acids and possible relationship to genetic code. Proceedings of the National Academy of Sciences of the United States of America . 1993 , 90(19), 8763-8768. doi:10.1073/pnas.90.19.8763. https://doi.org/10.1073/pnas.90.19.8763
-
[10]
Journal of Molecular Evo- lution
Yarus M.; Widmann JJ.; Knight R., RNA-amino acid binding: a stereochemical era for the genetic code. Journal of Molecular Evo- lution. 2009, 69, 406-429. doi:10.1007/s00239-009-9270-1. https://doi.org/10.1007/s00239-009-9270-1
-
[11]
, The origin of the genetic code: theories and their relationships, a review
Di Giulio M. , The origin of the genetic code: theories and their relationships, a review. Biosystems, 2005, 80(2), 175-184. doi:10.1016/j.biosystems.2004.11.005. https://doi.org/10.1016/j.biosystems.2004.11.005
-
[12]
Philosophical Transactions of the Royal Society B: Biological Sciences
Suga H.; Hayashi G.; Terasaka N., The RNA origin of transfer RNA aminoacylation and beyond. Philosophical Transactions of the Royal Society B: Biological Sciences. 2011, 366(1580), 2959-2964. doi:10.1098/rstb.2011.0137. https://doi.org/10.1098/rstb.2011.0137
-
[13]
Ishida S.; Terasaka N.; Katoh T.; Suga H., An aminoacylation ribozyme evolved from a natural tRNA-sensing T-box riboswitch. Nature Chemical Biology. 2020, 16(6), 702-709. doi:10.1038/s41589-020-0500-6. https://doi.org/10.1038/s41589-020-0500-6
-
[14]
Naturwissenschaften 1971, 58(10), 465–523
Eigen, M., Selforganization of matter and the evolution of biological macromolecules. Naturwissenschaften 1971, 58(10), 465–523. https://doi.org/10.1007/BF00623322
-
[15]
Nicolis, G.,; Prigogine, I., Self-organization in non-equilibrium systems. Wiley. 1977. ISBN 978-0471024019
work page 1977
-
[16]
A principle of natural self -organization
Eigen M.; Schuster P., The hypercycle. A principle of natural self -organization. Part A: Emergence of the hypercycle. Naturwis- senschaften, 1977, 64(11), 541-565. doi:10.1007/BF00450633. https://doi.org/10.1007/BF00450633
-
[17]
https://www.mdpi.com/2075-1729/12/10/1580
Ishida,T., Emergence Simulation of Biological Cell -like Shapes Satisfying the Conditions of Life Using a Lattice -Type Multiset Chemical Model, LIFE, 2022, 12(10), 1580. https://www.mdpi.com/2075-1729/12/10/1580
work page 2022
-
[18]
Ishida,T., A constructive chemical oscillator model demonstrates the emergence of homeostasis before genetic information through active inference, Discov Life 56, 2 (2026) https://doi.org/10.1007/s11084-026-09723-x
-
[19]
J R Soc Interface, 2013; 10 (86): 20130475
Friston K., Life as we know it. J R Soc Interface, 2013; 10 (86): 20130475. https://doi.org/10.1098/rsif.2013.0475
-
[20]
Hordijk, W., Autocatalytic Sets: From the Origin of Life to the Economy, BioScience, 2013, vol. 63, no. 11, pp. 877 –881, doi: 10.1525/bio.2013.63.11.6. https://academic.oup.com/bioscience/article-lookup/doi/10.1525/bio.2013.63.11.6
-
[21]
Systematic and Applied Microbiology
Wächtershäuser G., Pyrite Formation, the First Energy Source for Life: a Hypothesis. Systematic and Applied Microbiology. 1988, 10(3), 207-210. doi:10.1016/S0723-2020(88)80001-8. https://doi.org/10.1016/S0723-2020(88)80001-8
-
[22]
Huber C.; Wächtershäuser G., Peptides by Activation of Amino Acids with CO on (Ni,Fe)S Surfaces: Implications for the Origin of Life. Science. 1998, 281(5377), 670-672. doi:10.1126/science.281.5377.670. https://doi.org/10.1126/science.281.5377.670
-
[23]
Chen, C.; Li, J., Recent advances in coacervate protocells from passive catalysts to chemically programmable systems, Commu- nications Chemistry, 2026, vol. 9, art. no. 76, doi: 10.1038/s42004-026-01937-4. 33 of 34 https://www.nature.com/articles/s42004-026-01937-4
-
[24]
Matsuo, M.; Kurihara, K., Proliferating coacervate droplets as the missing link between chemistry and biology in the origin s of life, Nature Communications, 2021, vol. 12, no. 1, art. no. 5487, doi: 10.1038/s41467-021-25530-6. https://doi.org/10.1038/s41467-021-25530-6
-
[25]
Kauffman, S. A., Autocatalytic sets of proteins, Journal of Theoretical Biology, 1986,Vol.119, Issue 1, , Pages 1 -24, https://doi.org/10.1016/S0022-5193(86)80047-9 https://www.nature.com/articles/s41467-021-25530-6
-
[26]
A.; Roli, A., Is the emergence of life and of agency expected?, Phil
Kauffman, S. A.; Roli, A., Is the emergence of life and of agency expected?, Phil. Trans. R. Soc. B, 2025, vol. 380, no. 1 936, p. 20240283, doi: 10.1098/rstb.2024.0283. https://royalsocietypublishing.org/doi/10.1098/rstb.2024.0283
-
[27]
Hordijk, W.; Steel, M., Autocatalytic Networks at the Basis of Life’s Origin and Organization, Life, 2018, vol. 8, no. 4, p . 62, doi: 10.3390/life8040062. https://www.mdpi.com/2075-1729/8/4/62
-
[28]
Hordijk, W.; Steel, M., Detecting autocatalytic, self -sustaining sets in chemical reaction systems. J. Theor. Biol. 2004, 227, 451 –
work page 2004
-
[29]
(doi:10.1016/j.jtbi.2003.11.020) https://doi.org/ doi:10.1016/j.jtbi.2003.11.020
-
[30]
doi: 10.26434/chemrxiv.7895891.v1
Hou, L.; Dueñas-Diez, M.; Srivastava, R.; Perez-Mercader, J., Flow Chemistry Controls Both Self -Assembly and the Entrapped Oscillatory Cargo in Belousov -Zhabotinsky Driven Polymerization -Induced Self -Assembly, ChemRxiv, 2019. doi: 10.26434/chemrxiv.7895891.v1. https://chemrxiv.org/doi/full/10.26434/chemrxiv.7895891.v1
-
[31]
Leira-Iglesias, J.; Tassoni, A.; Adachi, T.; Stich, M.; Hermans, T. M., Oscillations, travelling fronts and patterns in a supramo- lecular system, Nature Nanotech, 2018, vol. 13, no. 11, pp. 1021–1027, doi: 10.1038/s41565-018-0270-4. https://www.nature.com/articles/s41565-018-0270-4
-
[32]
-F.; Zhang, X., Dissipative Supramolecular Polymerization Powered by Light, CCS Chem, 2019, vol
Yin, Z.; Song, G.; Jiao, Y.; Zheng, P.; Xu, J. -F.; Zhang, X., Dissipative Supramolecular Polymerization Powered by Light, CCS Chem, 2019, vol. 1, no. 4, pp. 335–342,doi: 10.31635/ccschem.019.20190013. http://www.chinesechemsoc.org/doi/10.31635/ccschem.019.20190013
-
[33]
Ter Harmsel, M.; Maguire, O. R.; Runikhina, S. A.; Wong, A. S. Y.; Huck, W. T. S.; Harutyunyan, S. R., A catalytically ac tive oscillator made from small organic molecules, Nature, 2023, vol. 621, no. 7977, pp. 87–93, doi: 10.1038/s41586-023-06310-2. https://www.nature.com/articles/s41586-023-06310-2
-
[34]
A.; Rossi, F., Transport -driven chemical oscillations: a review,Phys
Budroni, M. A.; Rossi, F., Transport -driven chemical oscillations: a review,Phys. Chem. Chem. Phys.,2024, vol. 26, no. 47, pp. 29185–29226, doi: 10.1039/D4CP03466J. https://xlink.rsc.org/?DOI=D4CP03466J
-
[35]
Damer, B.; Deamer, D., The Hot Spring Hypothesis for an Origin of Life, Astrobiology, 2020, vol. 20, no. 4, pp. 429 –452, doi: 10.1089/ast.2019.2045. https://journals.sagepub.com/doi/full/10.1089/ast.2019.2045
-
[36]
N., Wet-dry cycles cause nucleic acid monomers to polymerize into long chains, Proc
Song, X.; Šimonis, P.; Deamer, D.; Zare, R. N., Wet-dry cycles cause nucleic acid monomers to polymerize into long chains, Proc. Natl. Acad. Sci. U.S.A., 2024, vol. 121, no. 49, p. e2412784121, doi: 10.1073/pnas.2412784121. https://pnas.org/doi/10.1073/pnas.2412784121
-
[37]
Segré, D.; Ben-Eli, D.; Lancet, D., Compositional genomes: prebiotic information transfer in mutually catalytic noncovalent as- semblies, Proc. Natl. Acad. Sci. U.S.A., 2000, vol. 97, no. 8, pp. 4112–4117, doi: 10.1073/pnas.97.8.4112. https://pnas.org/doi/full/10.1073/pnas.97.8.4112
-
[38]
Parrondo, J. M. R. ; Horowitz, J. M.; Sagawa, T., Thermodynamics of information, Nature Physics, 2015, vol. 11, pp. 131 –139, 2015, doi: 10.1038/nphys3230. https://www.nature.com/articles/nphys3230
-
[39]
Marsaglia,G., Xorshift RNGs, Journal of Statistical Software, 2003, Vol. 8, Issue 14, pp. 1-6, DOI: 10.18637/jss.v008.i14 https://doi.org/10.18637/jss.v008.i14
-
[40]
J., The Origin and Nature of Life on Earth: The Emergence of the Fourth Geosphere
Smith, E.; Morowitz, H. J., The Origin and Nature of Life on Earth: The Emergence of the Fourth Geosphere. Cambridge Univer- sity Press. 2016, ISBN: 978-1107121881 34 of 34 Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual au- thor(s) and contributor(s) and not of MDPI and/or th...
work page 2016
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.