Recognition: no theorem link
Flow Matching for Count Data
Pith reviewed 2026-05-11 02:23 UTC · model grok-4.3
The pith
Count-FM generates higher-quality count data samples than baselines while using substantially fewer parameters by modeling dynamics directly with birth-death processes.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Count-FM learns marginal transitions efficiently in count space through simulation-free training of conditional transition rates in a birth-death framework. This enables transport between arbitrary count-distributed populations. In simulation it produces better sample quality than representative baselines while using substantially fewer parameters. Applied to scRNA-seq and neural spike-train data, the same model supports unconditional generation, transport between distributions, and conditional generation, delivering improved sample quality, greater modeling efficiency, and interpretable transport paths.
What carries the argument
continuous-time birth-death process with local unit jumps, which models each count changing by +1 or -1 at stochastic times and carries the flow-matching objective directly in discrete space
If this is right
- Unconditional generation of realistic count vectors for gene expression or neural activity becomes feasible with reduced parameter counts.
- Transport maps between successive batches or time points of count data can be learned and visualized as sequences of unit jumps.
- Conditional generation tasks, such as predicting target counts given partial observations, inherit the same efficiency and path interpretability.
- The simulation-free training of transition rates scales to high-dimensional count problems where direct simulation of full trajectories would be prohibitive.
Where Pith is reading between the lines
- The unit-jump birth-death structure might combine with existing continuous flow models to handle mixed continuous-discrete observations in a single framework.
- Interpretable paths could be used post hoc to identify which features drive transitions in biological count data, offering a form of mechanistic insight without additional supervision.
- Because the model stays in count space, downstream tasks such as differential expression testing or spike-count statistics can be performed on generated samples without inverse transformations.
- Efficiency gains from fewer parameters could allow deployment on edge devices for real-time neural decoding from spike trains.
Load-bearing premise
A continuous-time birth-death process with only local unit jumps supplies a sufficient model for the dynamics of high-dimensional count data.
What would settle it
If count-FM applied to held-out scRNA-seq or spike-train datasets produces samples whose empirical distributions match the true data worse than a standard continuous diffusion baseline when both are given equal training budget, the claimed superiority collapses.
Figures
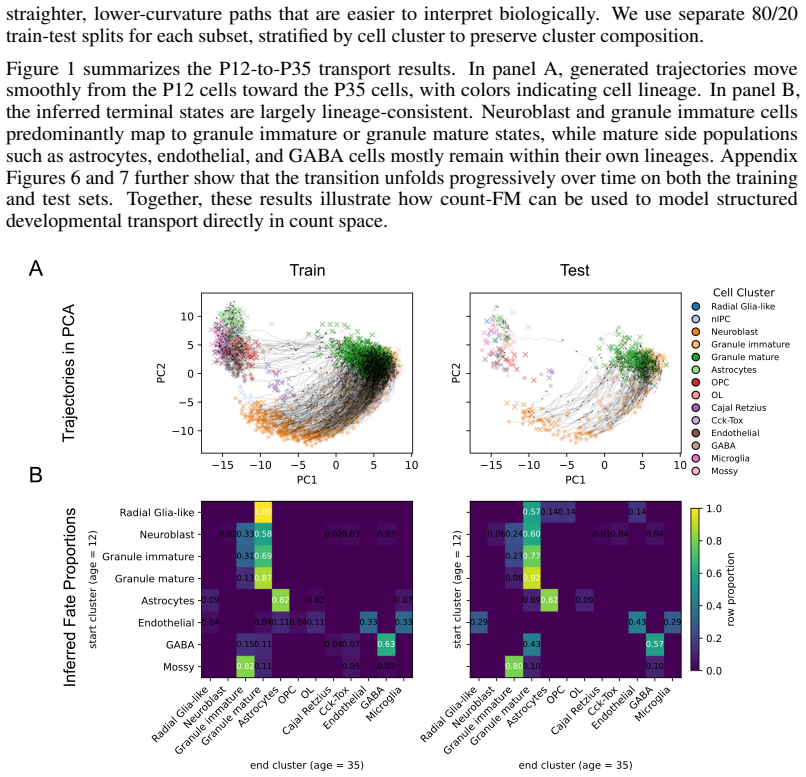


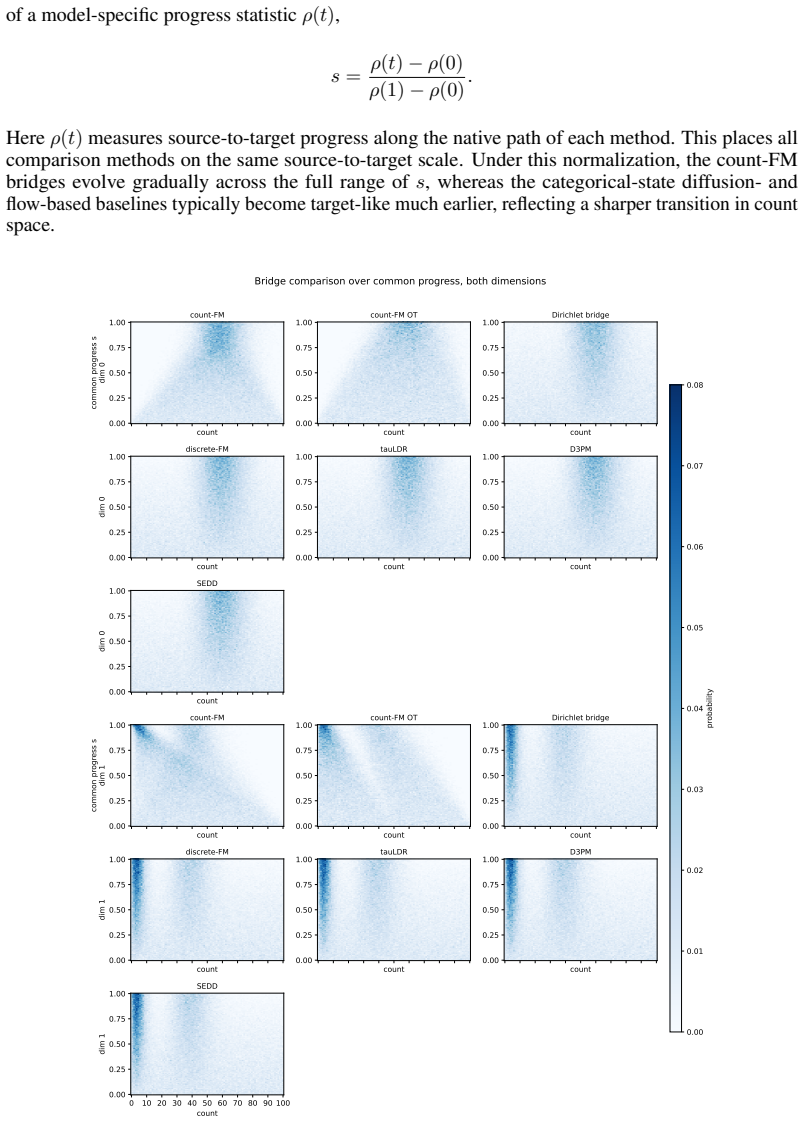

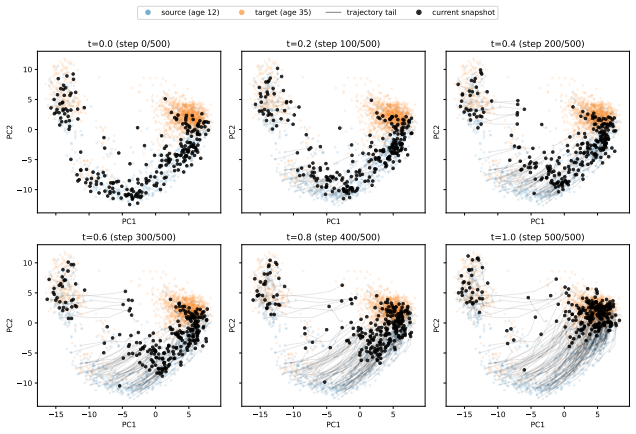

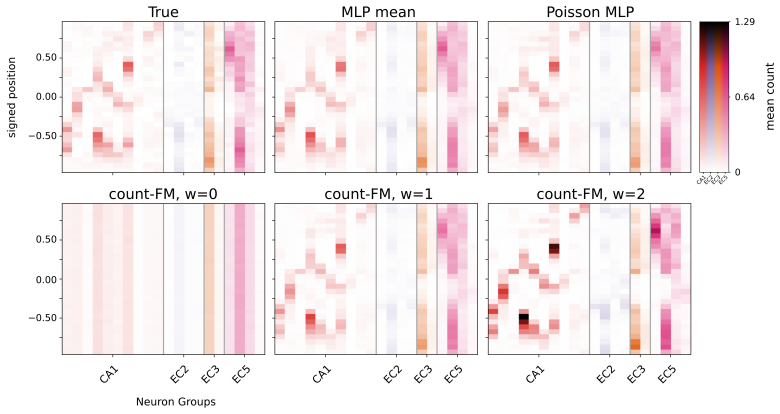

read the original abstract
High-dimensional count data arise in applications such as single-cell RNA sequencing and neural spike trains, where mapping between distributions across successive batches or time points form critical components of data analysis. The recent success of diffusion- and flow-based deep generative models for images, video, and text motivates extending these ideas to count-valued settings, but many existing methods either treat each count as a categorical state or transform counts into a continuous space, neither of which is natural or efficient when the count range is large. We propose count-FM, a flow-matching framework for count data based on a continuous-time birth-death process with local unit jumps. Count-FM learns marginal transitions efficiently in count space through simulation-free training of conditional transition rates, allowing transport between arbitrary count-distributed source and target populations. In simulation, count-FM achieves better sample quality than representative baselines while using substantially fewer parameters. We further apply count-FM to scRNA-seq and neural spike-train data for unconditional generation, transport, and conditional generation. Across these tasks, count-FM yields improved sample quality, greater modeling efficiency, and interpretable transport paths.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes count-FM, a flow-matching framework for high-dimensional count data that models dynamics via a continuous-time birth-death process restricted to local unit jumps per coordinate. It introduces simulation-free training of conditional transition rates to enable transport between arbitrary count distributions, and reports superior sample quality and parameter efficiency versus baselines in simulations, plus applications to unconditional/conditional generation and transport on scRNA-seq and neural spike-train data.
Significance. If the central modeling assumptions hold, count-FM offers a natural extension of flow matching to count-valued data without continuous transformations or categorical treatments, which is valuable for single-cell and neuroscience applications. The simulation-free objective and reported gains in modeling efficiency and interpretable paths represent concrete strengths over diffusion-style alternatives that require simulation.
major comments (2)
- [§2] §2 (Birth-death process definition): the framework restricts dynamics to independent per-coordinate unit jumps. For applications like scRNA-seq (co-regulated genes) and spike trains (population coding), where strong cross-dimensional correlations are expected, no analysis or theorem is given showing that learned marginal rates compose to accurate joint distributions; this assumption is load-bearing for the sample-quality and transport claims.
- [Experiments] Experiments section (simulation and real-data results): the abstract and main claims assert better sample quality with fewer parameters, but no quantitative metrics (e.g., MMD, log-likelihood, or Wasserstein distances), baseline implementation details, error bars, or data-exclusion criteria are referenced in the provided text; without these, the performance advantage cannot be verified as load-bearing evidence.
minor comments (2)
- [§3] Notation for the conditional transition rates could be clarified with an explicit equation linking the simulation-free objective to the birth-death generator.
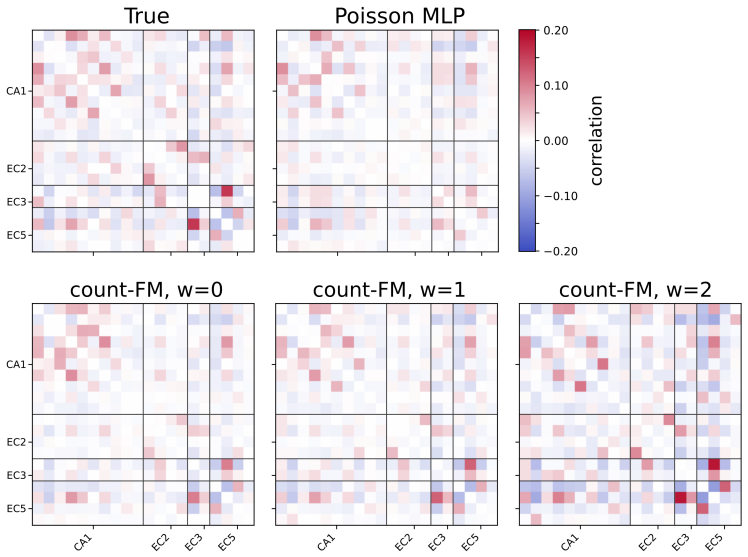
- [Figures] Figure captions for transport paths should specify how interpretability is quantified or visualized (e.g., via marginal trajectories or correlation preservation).
Simulated Author's Rebuttal
We thank the referee for the constructive and detailed feedback on our manuscript. We address each major comment below and outline the changes we will make in revision.
read point-by-point responses
-
Referee: [§2] §2 (Birth-death process definition): the framework restricts dynamics to independent per-coordinate unit jumps. For applications like scRNA-seq (co-regulated genes) and spike trains (population coding), where strong cross-dimensional correlations are expected, no analysis or theorem is given showing that learned marginal rates compose to accurate joint distributions; this assumption is load-bearing for the sample-quality and transport claims.
Authors: We thank the referee for highlighting this modeling choice. The birth-death process indeed restricts transitions to unit jumps in a single coordinate at a time, but the transition rates for each such jump are explicitly functions of the full current state vector. This state dependence allows the rates to encode cross-coordinate correlations (e.g., co-regulation in genes or population coding in spikes). The simulation-free training objective learns conditional transition rates that are consistent with the joint transport between source and target distributions, analogous to conditional flow matching in the continuous case. While we do not supply a formal theorem proving exact marginal-to-joint composition, the framework is constructed so that the learned rates define a valid joint Markov process. We will add a clarifying paragraph in §2 explaining the role of state-dependent rates and their empirical validation on correlated real-world data. revision: partial
-
Referee: [Experiments] Experiments section (simulation and real-data results): the abstract and main claims assert better sample quality with fewer parameters, but no quantitative metrics (e.g., MMD, log-likelihood, or Wasserstein distances), baseline implementation details, error bars, or data-exclusion criteria are referenced in the provided text; without these, the performance advantage cannot be verified as load-bearing evidence.
Authors: We agree that the current text would be strengthened by explicit quantitative metrics and experimental details. Although comparative figures and supplementary results support the claims of improved sample quality and parameter efficiency, we acknowledge that MMD, Wasserstein distances, log-likelihood values, error bars from multiple runs, baseline implementation specifics, and data-exclusion criteria are not directly referenced in the main body. In the revised manuscript we will add these quantitative evaluations, report means and standard deviations across repeated experiments, expand the baseline descriptions, and include a clear data-processing section. revision: yes
Circularity Check
No significant circularity; derivation is self-contained from birth-death process definition
full rationale
The paper introduces count-FM by defining a continuous-time birth-death process with local unit jumps as the underlying dynamics for count data, then derives a simulation-free training objective for conditional transition rates directly from this process. No step reduces a claimed prediction or result to a fitted parameter or self-citation by construction; the marginal transition learning and transport paths follow from the process equations without tautological equivalence to inputs. The framework remains independent of the target data distributions and does not invoke prior author work to force uniqueness or ansatz choices.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption A continuous-time birth-death process with local unit jumps is a suitable model for transitions between count distributions.
Reference graph
Works this paper leans on
-
[1]
Sanjukta Bhattacharya, Christian Gensbigler, Shaamil Karim, and Jon Lees
URLhttps://openreview.net/forum?id=h7-XixPCAL. Sanjukta Bhattacharya, Christian Gensbigler, Shaamil Karim, and Jon Lees. Discrete diffusion for single-cell gene expression modeling. InICLR 2026 Workshop on Machine Learning for Genomics Explorations,
work page 2026
-
[2]
ISSN 2050-084X. doi: 10.7554/eLife.22630. URL https://doi.org/10.7554/eLife.22630. Kevin A Bolding, Shivathmihai Nagappan, Bao-Xia Han, Fan Wang, and Kevin M Franks. Recurrent circuitry is required to stabilize piriform cortex odor representations across brain states.eLife, 9: e53125, jul
-
[3]
ISSN 2050-084X. doi: 10.7554/eLife.53125. URL https://doi.org/10. 7554/eLife.53125. Andrew Campbell, Joe Benton, Valentin De Bortoli, Tom Rainforth, George Deligiannidis, and Arnaud Doucet. A continuous time framework for discrete denoising models. InAdvances in Neural Information Processing Systems,
-
[4]
URL https://doi.org/ 10.1186/s13059-023-03148-9
doi: 10.1186/s13059-023-03148-9. URL https://doi.org/ 10.1186/s13059-023-03148-9. Jérôme Epsztein, Michael Brecht, and Albert K. Lee. Intracellular determinants of hippocampal ca1 place and silent cell activity in a novel environment.Neuron, 70(1):109–120,
-
[5]
URLhttps://doi.org/10.1016/j.neuron.2011.03.006
doi: 10.1016/j.neuron.2011.03.006. URLhttps://doi.org/10.1016/j.neuron.2011.03.006. Kevin Frans, Danijar Hafner, Sergey Levine, and Pieter Abbeel. One step diffusion via shortcut models. InThe Thirteenth International Conference on Learning Representations,
-
[6]
URL https://proceedings.neurips.cc/paper_files/paper/2014/file/ f033ed80deb0234979a61f95710dbe25-Paper.pdf. Arthur Gretton, Karsten M. Borgwardt, Malte J. Rasch, Bernhard Schölkopf, and Alexander Smola. A kernel two-sample test.Journal of Machine Learning Research, 13(25):723–773,
work page 2014
-
[7]
doi: 10.1113/jphysiol. 2010.194274. URLhttps://doi.org/10.1113/jphysiol.2010.194274. Jonathan Ho and Tim Salimans. Classifier-free diffusion guidance. InNeurIPS 2021 Workshop on Deep Generative Models and Downstream Applications,
-
[8]
URL https://proceedings.neurips.cc/paper_files/paper/2020/ file/4c5bcfec8584af0d967f1ab10179ca4b-Paper.pdf. Asger Hobolth and Eric A. Stone. Simulation from endpoint-conditioned, continuous-time Markov chains on a finite state space, with applications to molecular evolution.The Annals of Applied Statistics, 3(3):1204 – 1231,
work page 2020
-
[9]
doi: 10.1214/09-AOAS247. URL https://doi.org/10. 1214/09-AOAS247. Hannah Hochgerner, Amit Zeisel, Peter Lönnerberg, and Sten Linnarsson. Conserved properties of dentate gyrus neurogenesis across postnatal development revealed by single-cell RNA sequencing. Nature Neuroscience, 21(2):290–299, February
-
[10]
doi: 10.1038/s41593-017-0056-2. Peter Holderrieth, Marton Havasi, Jason Yim, Neta Shaul, Itai Gat, Tommi Jaakkola, Brian Karrer, Ricky T. Q. Chen, and Yaron Lipman. Generator matching: Generative modeling with arbitrary markov processes. InThe Thirteenth International Conference on Learning Representations,
-
[11]
Emiel Hoogeboom, Taco Cohen, and Jakub Mikolaj Tomczak
URL https://proceedings.neurips.cc/paper_files/paper/ 2019/file/9e9a30b74c49d07d8150c8c83b1ccf07-Paper.pdf. Emiel Hoogeboom, Taco Cohen, and Jakub Mikolaj Tomczak. Learning discrete distributions by dequantization. InThird Symposium on Advances in Approximate Bayesian Inference,
work page 2019
-
[12]
Auto-Encoding Variational Bayes
URL https: //arxiv.org/abs/1312.6114. Durk P Kingma and Prafulla Dhariwal. Glow: Generative flow with invertible 1x1 convolu- tions. InAdvances in Neural Information Processing Systems, volume
work page internal anchor Pith review Pith/arXiv arXiv
-
[13]
URL https://proceedings.neurips.cc/paper_files/paper/2018/file/ d139db6a236200b21cc7f752979132d0-Paper.pdf. Marius Lange, V olker Bergen, Michal Klein, Manu Setty, Bernhard Reuter, Mostafa Bakhti, Heiko Lickert, Meshal Ansari, Janine Schniering, Herbert B. Schiller, Dana Pe’er, and Fabian J. Theis. Cellrank for directed single-cell fate mapping.Nature Met...
work page 2018
-
[14]
URL https://doi.org/10.1038/s41592-021-01346-6
doi: 10.1038/s41592-021-01346-6. URL https://doi.org/10.1038/s41592-021-01346-6 . Yaron Lipman, Ricky T. Q. Chen, Heli Ben-Hamu, Maximilian Nickel, and Matthew Le. Flow matching for generative modeling. InThe Eleventh International Conference on Learning Repre- sentations,
-
[15]
Aaron Lou, Chenlin Meng, and Stefano Ermon
doi: 10.1038/s41592-018-0229-2. Aaron Lou, Chenlin Meng, and Stefano Ermon. Discrete diffusion modeling by estimating the ratios of the data distribution. InForty-first International Conference on Machine Learning,
-
[16]
doi: 10.1093/bioinformatics/btae518
ISSN 1367-4811. doi: 10.1093/bioinformatics/btae518. URL https://doi.org/10.1093/ bioinformatics/btae518. Youzhi Luo, Keqiang Yan, and Shuiwang Ji. Graphdf: A discrete flow model for molecular graph generation. InProceedings of the 38th International Conference on Machine Learning, volume 139 ofProceedings of Machine Learning Research, pages 7192–7203. PM...
-
[18]
URLhttps://doi.org/10.1007/BF00237147
doi: 10.1007/BF00237147. URLhttps://doi.org/10.1007/BF00237147. Keiji Miura, Zachary F. Mainen, and Naoshige Uchida. Odor representations in olfactory cortex: Distributed rate coding and decorrelated population activity.Neuron, 74(6):1087–1098, June
-
[19]
URL https://doi.org/10.1016/j.neuron.2012.04
doi: 10.1016/j.neuron.2012.04.021. URL https://doi.org/10.1016/j.neuron.2012.04
-
[20]
doi: 10.12688/f1000research.3895.2. Giovanni Palla, Sudarshan Babu, Payam Dibaeinia, Donghui Li, Aly A Khan, Theofanis Karaletsos, and Jakub M. Tomczak. A scalable latent diffusion model for single-cell gene expression data. In NeurIPS 2025 Workshop on AI Virtual Cells and Instruments: A New Era in Drug Discovery and Development,
-
[21]
URLhttps://arxiv.org/abs/2604.09784. Geoffrey Schiebinger, Jian Shu, Marcin Tabaka, Brian Cleary, Vidya Subramanian, Aryeh Solomon, Joshua Gould, Siyan Liu, Stacie Lin, Peter Berube, Lia Lee, Jenny Chen, Justin Brumbaugh, Philippe Rigollet, Konrad Hochedlinger, Rudolf Jaenisch, Aviv Regev, and Eric S. Lander. Optimal-transport analysis of single-cell gene...
work page internal anchor Pith review Pith/arXiv arXiv
-
[22]
ISSN 0092-8674. doi: https://doi.org/10. 1016/j.cell.2019.01.006. URL https://www.sciencedirect.com/science/article/pii/ S009286741930039X. Marta Skreta, Tara Akhound-Sadegh, Viktor Ohanesian, Roberto Bondesan, Alan Aspuru-Guzik, Ar- naud Doucet, Rob Brekelmans, Alexander Tong, and Kirill Neklyudov. Feynman-kac correctors in diffusion: Annealing, guidance...
work page 2019
-
[23]
doi: 10.1523/JNEUROSCI. 09-07-02382.1989. URLhttps://www.jneurosci.org/content/9/7/2382. Alexander Tong, Kilian FATRAS, Nikolay Malkin, Guillaume Huguet, Yanlei Zhang, Jarrid Rector- Brooks, Guy Wolf, and Yoshua Bengio. Improving and generalizing flow-based generative models with minibatch optimal transport.Transactions on Machine Learning Research,
-
[24]
We also describe the discretization used for sampling in Section 2.2
We first derive the conditional bridge rates, then derive the local rate-matching objective and show how the local KL terms integrate to the path-space KL objective. We also describe the discretization used for sampling in Section 2.2. A.1 Derivation of conditional bridge rates This subsection derives the conditional birth and death rates used in Section ...
work page 2022
-
[25]
trajectory tail current snapshot Figure 7:Transition snapshots for testing.The same progressive transport pattern is observed on held-out cells, with trajectories moving smoothly from the P12 manifold toward the P35 manifold. D.3 Conditional generation of piriform cortex spike trains We also evaluate count-FM on conditional generation of multichannel piri...
work page 2017
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.