Recognition: 2 theorem links
· Lean TheoremModel-based Dynamic 3D MRI Reconstructions using Neural Fields and Tensor Product Expansions
Pith reviewed 2026-05-12 01:02 UTC · model grok-4.3
The pith
A tensor-product neural-field model reconstructs dynamic 3D MRI from highly undersampled data while preserving structure and motion.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Conventional MRI reconstruction methods treat images and coil sensitivities as discrete objects, leading to high memory demands and limited structural awareness that hamper effective regularization. These limitations hinder accurate reconstruction in highly undersampled scenarios, such as dynamic 3D cardiac magnetic resonance (CMR). We introduce a discretization-free, memory-efficient, model-based framework for dynamic 2D and 3D MRI reconstruction from highly undersampled data. We represent magnetization and coil sensitivities as continuous objects -- differentiable functions -- using tensor products of univariate neural fields. This tensor product structure enables scalable optimization in高
What carries the argument
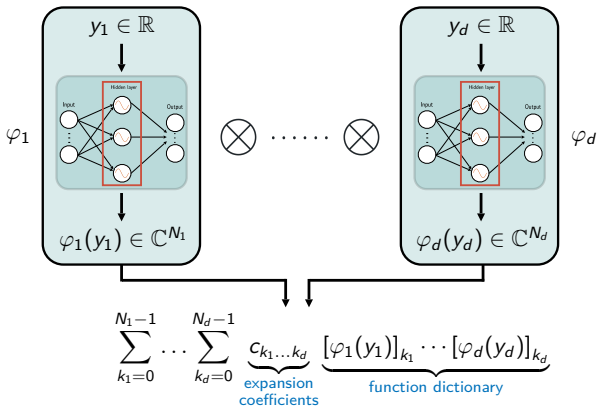
Tensor products of univariate neural fields representing magnetization and coil sensitivities as continuous differentiable functions for scalable optimization in spatiotemporal MRI.
If this is right
- The method reduces memory usage by avoiding discrete grids.
- It enables effective regularization through the continuous representation.
- Reconstructions maintain structure and motion at high undersampling rates like 16x acceleration.
- Optimization scales to dynamic 3D and 2D settings without instability.
- It outperforms current model-based methods in quality for undersampled dynamic MRI.
Where Pith is reading between the lines
- This continuous neural-field approach could be applied to other tomographic inverse problems where memory and regularization are bottlenecks.
- The tensor product expansion might allow extension to 4D flow imaging or higher temporal resolutions.
- Combining with other implicit representations could further improve efficiency in clinical workflows.
- The built-in differentiability may facilitate integration with downstream tasks like motion correction or segmentation.
Load-bearing premise
Representing magnetization and coil sensitivities as continuous differentiable functions via tensor products of univariate neural fields will automatically provide effective regularization and scalable optimization in high-dimensional spatiotemporal settings without introducing new artifacts or optimization instabilities.
What would settle it
A head-to-head test on dynamic 3D cardiac MRI data at 16-fold acceleration showing whether the neural-field method uses less memory and achieves lower reconstruction error than discrete SOTA model-based methods while avoiding blurring or motion artifacts.
Figures








read the original abstract
Conventional MRI reconstruction methods treat images and coil sensitivities as discrete objects, leading to high memory demands and limited structural awareness that hamper effective regularization. These limitations hinder accurate reconstruction in highly undersampled scenarios, such as dynamic 3D cardiac magnetic resonance (CMR). We introduce a discretization-free, memory-efficient, model-based framework for dynamic 2D and 3D MRI reconstruction from highly undersampled data. We represent magnetization and coil sensitivities as continuous objects -- differentiable functions -- using tensor products of univariate neural fields. This tensor product structure enables scalable optimization in high-dimensional spatiotemporal settings. Our method outperforms state-of-the-art model-based reconstructions in dynamic 2D and 3D MR settings, preserving structure and motion even under aggressive undersampling (e.g., acceleration factor 16).
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proposes a discretization-free, model-based framework for dynamic 2D and 3D MRI reconstruction from highly undersampled k-space data. Magnetization M(x,t) and coil sensitivities C(x) are represented as continuous differentiable functions via tensor products of univariate neural fields; this structure is claimed to enable memory-efficient, scalable optimization while providing implicit regularization that preserves structure and motion, outperforming state-of-the-art model-based methods even at acceleration factor 16 in dynamic cardiac settings.
Significance. If the performance claims hold under rigorous validation, the work could meaningfully advance high-dimensional dynamic MRI by mitigating memory bottlenecks and discretization artifacts that limit conventional model-based approaches. The continuous neural-field representation with tensor-product factorization is a timely idea for spatiotemporal scalability, and explicit credit is due for targeting the non-convex joint optimization of magnetization and sensitivities in 3D+t regimes.
major comments (2)
- [Abstract] Abstract: the central claim that tensor products of univariate neural fields 'automatically' yield effective implicit regularization and artifact-free optimization under R=16 undersampling is load-bearing yet unsupported by any derivation, complexity analysis, or ablation isolating this mechanism from standard model-based penalties; the non-convex MRI forward model is known to require explicit regularization to avoid noise fitting, and the degree-of-freedom reduction alone does not guarantee motion/structure preservation.
- [Results] Results/Experiments section: no quantitative metrics, baseline comparisons, or error maps are referenced to substantiate the outperformance statement; without reported NRMSE, SSIM, or temporal fidelity values on public dynamic 3D datasets, the claim that the method 'preserves structure and motion' remains unverified and cannot be assessed for statistical significance.
minor comments (2)
- Define all acronyms at first use (e.g., CMR) and ensure consistent notation for the tensor-product operator across text and any equations.
- [Abstract] The abstract would be strengthened by including one or two concrete quantitative results (e.g., acceleration factor and a key metric) rather than qualitative statements alone.
Simulated Author's Rebuttal
We thank the referee for the constructive and insightful comments on our manuscript. We value the recognition of the potential for our tensor-product neural field approach to address memory and scalability challenges in dynamic 3D MRI. We address each major comment below with specific plans for revision.
read point-by-point responses
-
Referee: [Abstract] Abstract: the central claim that tensor products of univariate neural fields 'automatically' yield effective implicit regularization and artifact-free optimization under R=16 undersampling is load-bearing yet unsupported by any derivation, complexity analysis, or ablation isolating this mechanism from standard model-based penalties; the non-convex MRI forward model is known to require explicit regularization to avoid noise fitting, and the degree-of-freedom reduction alone does not guarantee motion/structure preservation.
Authors: We agree that the abstract's phrasing regarding automatic implicit regularization requires stronger substantiation. The manuscript motivates the tensor-product factorization primarily through memory efficiency and differentiability, but does not provide an explicit derivation or ablation. In revision, we will add a dedicated paragraph in the Methods section deriving the effective regularization from the separable univariate fields (including a parameter-count analysis showing the reduction in degrees of freedom relative to a full spatiotemporal grid), and we will include a new ablation experiment comparing tensor-product versus non-factorized neural-field baselines under identical optimization settings to isolate the contribution to structure and motion preservation. revision: yes
-
Referee: [Results] Results/Experiments section: no quantitative metrics, baseline comparisons, or error maps are referenced to substantiate the outperformance statement; without reported NRMSE, SSIM, or temporal fidelity values on public dynamic 3D datasets, the claim that the method 'preserves structure and motion' remains unverified and cannot be assessed for statistical significance.
Authors: The referee is correct that the current Results section emphasizes qualitative visual comparisons and does not tabulate quantitative metrics or reference error maps in the text. While the figures illustrate preservation of fine structure and temporal dynamics at R=16, this is insufficient for rigorous evaluation. We will revise the Results section to include tables of NRMSE, SSIM, and temporal fidelity (e.g., temporal gradient error) for both 2D and 3D experiments, with direct numerical comparisons against the cited state-of-the-art model-based baselines. Error maps will be added as supplementary figures and explicitly referenced. Our primary dataset is clinical dynamic cardiac MRI; we will clearly state this and, where feasible, add a supplementary experiment on an available public dynamic dataset to allow broader assessment. revision: partial
Circularity Check
No circularity: independent modeling choice for continuous representation
full rationale
The paper proposes representing magnetization and coil sensitivities as tensor products of univariate neural fields as a discretization-free modeling decision. This choice is presented directly in the abstract as enabling scalable optimization in high-dimensional settings, without any derivation that reduces the claimed benefits to fitted parameters, self-referential equations, or load-bearing self-citations. No equations or steps in the provided text exhibit self-definition, renaming of known results as new predictions, or ansatz smuggling. The outperformance claims rest on experimental comparisons rather than tautological reductions, making the framework self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
free parameters (1)
- Neural field network parameters
axioms (2)
- domain assumption Magnetization and coil sensitivities can be accurately represented as continuous differentiable functions
- domain assumption Tensor product structure enables scalable optimization in high-dimensional settings
Reference graph
Works this paper leans on
- [1]
-
[2]
Syed M Arshad, Lee C Potter, Chong Chen, Yingmin Liu, Preethi Chandrasekaran, Christopher Crabtree, Matthew S Tong, Orlando P Simonetti, Yuchi Han, and Rizwan Ahmad,Motion-robust free-running volumetric cardiovascular mri, Magnetic Resonance in Medicine92(2024), no. 3, 1248–1262
work page 2024
-
[3]
Sampurna Biswas, Hemant K Aggarwal, and Mathews Jacob,Dynamic mri using model-based deep learning and storm priors: Modl-storm, Magnetic resonance in medicine82(2019), no. 1, 485–494
work page 2019
-
[4]
Chong Chen, Yingmin Liu, Philip Schniter, Matthew Tong, Karolina Zareba, Orlando Simonetti, Lee Potter, and Rizwan Ahmad,Ocmr (v1.0)–open-access multi-coil k-space dataset for cardiovascular magnetic resonance imaging, 2020
work page 2020
-
[5]
Cheney,Introduction to approximation theory, AMS Chelsea Publishing Series, AMS Chelsea Pub., 1998
E.W. Cheney,Introduction to approximation theory, AMS Chelsea Publishing Series, AMS Chelsea Pub., 1998
work page 1998
-
[6]
Junwoo Cho, Seungtae Nam, Hyunmo Yang, Seok-Bae Yun, Youngjoon Hong, and Eunbyung Park,Separable physics-informed neural networks, Advances in Neural Information Processing Systems (2023)
work page 2023
-
[7]
Carl De Boor,A practical guide to splines, Vol. 27, Springer, New York, 2001
work page 2001
-
[8]
Jie Feng, Ruimin Feng, Qing Wu, Xin Shen, Lixuan Chen, Xin Li, Li Feng, Jingjia Chen, Zhiyong Zhang, Chunlei Liu, Yuyao Zhang, and Hongjiang Wei,Spatiotemporal implicit neural representation for unsupervised dynamic mri reconstruction, IEEE Transactions on Medical Imaging44(2025), no. 5, 2143–2156
work page 2025
-
[9]
Giannakopoulos, Matthew Muckley, Jesi Kim, Matthew Breen, Patricia M Johnson, Yvonne W
Ilias I. Giannakopoulos, Matthew Muckley, Jesi Kim, Matthew Breen, Patricia M Johnson, Yvonne W. Lui, and Riccardo Lattanzi,Accelerated mri reconstructions via variational network and feature domain learning, Scientific Reports14(2024)
work page 2024
-
[10]
Mark A. Griswold, Peter M. Jakob, Robin M. Heidemann, Mathias Nittka, Vladimír Jellús, Jianmin Wang, Berthold Kiefer, and Axel Haase,Generalized autocalibrating partially parallel acquisitions (grappa), Magnetic Resonance in Medicine47(2002)
work page 2002
-
[11]
Mihir Joshi, Aaron Pruitt, Chong Chen, Yingmin Liu, and Rizwan Ahmad,Technical report (v1.0)–pseudo- random cartesian sampling for dynamic mri, 2022
work page 2022
-
[12]
Rui jun Feng, Qing Wu, Jie Feng, Huajun She, Chunlei Liu, Yuyao Zhang, and Hongjiang Wei,Imjense: Scan- specific implicit representation for joint coil sensitivity and image estimation in parallel mri,IEEETransactions on Medical Imaging43(2023), 1539–1553
work page 2023
- [13]
-
[14]
Diederik Kinga, Jimmy Ba Adam, et al.,A method for stochastic optimization, International conference on learning representations (iclr), 2015
work page 2015
-
[15]
Stefan Klein, Marius Staring, Keelin Murphy, Max A Viergever, and Josien PW Pluim,Elastix: a toolbox for intensity-based medical image registration, IEEE transactions on medical imaging29(2009), no. 1, 196–205
work page 2009
-
[16]
Thomas Küstner, Niccolo Fuin, Kerstin Hammernik, Aurélien Bustin, Haikun Qi, Reza Hajhosseiny, Pier Gior- gio Masci, Radhouène Neji, Daniel Rueckert, René M. Botnar, and Claudia Prieto,Cinenet: deep learning-based 3d cardiac cine mri reconstruction with multi-coil complex-valued 4d spatio-temporal convolutions, Scientific Reports10(2020)
work page 2020
-
[17]
Jun Lyu, Chen Qin, Shuo Wang, Fanwen Wang, Yan Li, Zi Wang, Kunyuan Guo, Ouyang Cheng, Michael Tanzer, Meng Liu, Longyu Sun, Meng qi Sun, Qin Li, Zhang Shi, Sha Hua, Hao Li, Zhensen Chen, Zhenlin Zhang, Bingyu Xin, Dimitris N. Metaxas, George Yiasemis, Jonas Teuwen, Liping Zhang, Weitian Chen, Yanwei Pang, Xiaohan Liu, Artem Razumov, Dmitry V. Dylov, Quan...
work page 2023
-
[18]
Amirali Molaei, Amirhossein Aminimehr, Armin Tavakoli, Amirhossein Kazerouni, Bobby Azad, Reza Azad, and Dorit Merhof,Implicit neural representation in medical imaging: A comparative survey, Proceedings of the ieee/cvf international conference on computer vision, 2023, pp. 2381–2391. 15
work page 2023
-
[19]
Thomas Müller, Alex Evans, Christoph Schied, and Alexander Keller,Instant neural graphics primitives with a multiresolution hash encoding41(July 2022), no. 4
work page 2022
-
[20]
Thomas E Olausson, Maarten L Terpstra, Niek RF Huttinga, Casper Beijst, Niels Blanken, Dominika Suchá, Teresa Correia, Birgitta K Velthuis, Cornelis AT van den Berg, and Alessandro Sbrizzi,Free-running time- resolved first-pass myocardial perfusion using a multi-scale dynamics decomposition: Cmr-motus, Magnetic Resonance Materials in Physics, Biology and ...
work page 2025
-
[21]
Ricardo Otazo, Emmanuel Candes, and Daniel K Sodickson,Low-rank plus sparse matrix decomposition for ac- celerated dynamic mri with separation of background and dynamic components, Magnetic resonance in medicine 73(2015), no. 3, 1125–1136
work page 2015
-
[22]
63, American Mathematical Soc., 2012
María Cristina Pereyra and Lesley A Ward,Harmonic analysis: from fourier to wavelets, Vol. 63, American Mathematical Soc., 2012
work page 2012
-
[23]
Claudia Prieto, Mariya Doneva, Muhammad Usman, Markus Henningsson, Gerald Greil, Tobias Schaeffter, and Rene M Botnar,Highly efficient respiratory motion compensated free-breathing coronary mra using golden-step cartesian acquisition, Journal of magnetic resonance imaging41(2015), no. 3, 738–746
work page 2015
-
[24]
Klaas Paul Pruessmann, Markus Weiger, Markus B. Scheidegger, and Peter Boesiger,Sense: Sensitivity encod- ing for fast mri, Magnetic Resonance in Medicine42(1999)
work page 1999
-
[25]
Javier Royuela-del Val, Lucilio Cordero-Grande, Federico Simmross-Wattenberg, Marcos Martín-Fernández, and Carlos Alberola-López,Jacobian weighted temporal total variation for motion compensated compressed sensing reconstruction of dynamic mri, Magnetic resonance in medicine77(2017), no. 3, 1208–1215
work page 2017
-
[26]
Matthias Schloegl, Martin Holler, Andreas Schwarzl, Kristian Bredies, and Rudolf Stollberger,Infimal convo- lution of total generalized variation functionals for dynamic mri, Magnetic resonance in medicine78(2017), no. 1, 142–155
work page 2017
-
[27]
Larry Schumaker,Spline functions: basic theory, Cambridge university press, 2007
work page 2007
-
[28]
Denis P Shamonin, Esther E Bron, Boudewijn PF Lelieveldt, Marion Smits, Stefan Klein, Marius Staring, and Alzheimer’s Disease Neuroimaging Initiative,Fast parallel image registration on cpu and gpu for diagnostic classification of alzheimer’s disease, Frontiers in neuroinformatics7(2014), 50
work page 2014
-
[29]
Liyue Shen, John Pauly, and Lei Xing,Nerp: Implicit neural representation learning with prior embedding for sparsely sampled image reconstruction, IEEE Transactions on Neural Networks and Learning Systems35 (2024), no. 1, 770–782
work page 2024
- [30]
-
[31]
Stelter, Tabita Catalan, Rickmer F
Veronika Spieker, Hannah Eichhorn, Wenqi Huang, Jonathan K. Stelter, Tabita Catalan, Rickmer F. Braren, Daniel Rueckert, Francisco Sahli Costabal, Kerstin Hammernik, Dimitrios C. Karampinos, Claudia Prieto, and Julia A. Schnabel,Pisco: Self-supervised k-space regularization for improved neural implicit k-space represen- tations of dynamic mri, Medical Ima...
work page 2025
-
[32]
Martin Uecker, Peng Lai, Mark Murphy, Patrick Virtue, Michael Elad, John M. Pauly, Shreyas S. Vasanawala, and Michael Lustig,Espirit—an eigenvalue approach to autocalibrating parallel mri: Where sense meets grappa, Magnetic Resonance in Medicine71(2014)
work page 2014
-
[33]
Cheng, Tao Zhang, and Michael Lustig,Berkeley advanced reconstruction toolbox, Proc
Martin Uecker, Frank Ong, Jon Tamir, Deniz Bahri, Patrick Virtue, Jane Y. Cheng, Tao Zhang, and Michael Lustig,Berkeley advanced reconstruction toolbox, Proc. intl. soc. mag. reson. med., 2015, pp. 2486
work page 2015
- [34]
-
[35]
David F Walnut,An introduction to wavelet analysis, Springer Science & Business Media, 2013
work page 2013
-
[36]
Jiawen Yao, Zheng Xu, Xiaolei Huang, and Junzhou Huang,An efficient algorithm for dynamic mri using low-rank and total variation regularizations, Medical image analysis44(2018), 14–27. 16
work page 2018
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.