Recognition: no theorem link
Measuring and Decomposing Mode Separation via the Canonical Diffusion
Pith reviewed 2026-05-12 01:19 UTC · model grok-4.3
The pith
A canonical reversible diffusion with constant scalar coefficient extracts mode separation from a density's autocovariance matrix.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
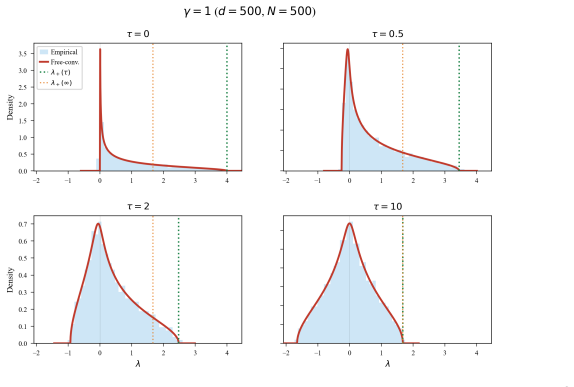
We measure mode separation through a single stochastic process intrinsic to the density: a unique reversible diffusion with f as its stationary distribution and constant scalar diffusion coefficient. We extract two readouts from its autocovariance matrix: SSA (Sum of Squared Autocorrelations), a scalar barrier-sensitive measure; and DA (Dominant Autocorrelation directions), linear projections ordered by metastability rather than variance. Under an isotropic-Gaussian null, we derive a closed-form spectrum for the empirical autocovariance that generalizes Marchenko-Pastur, with an analytic upper edge that selects the lag at which DA is read off. Both readouts use only samples and a score, and,
What carries the argument
The canonical diffusion: the unique reversible diffusion with given stationary density f and constant scalar diffusion coefficient, whose autocovariance matrix supplies SSA and DA.
If this is right
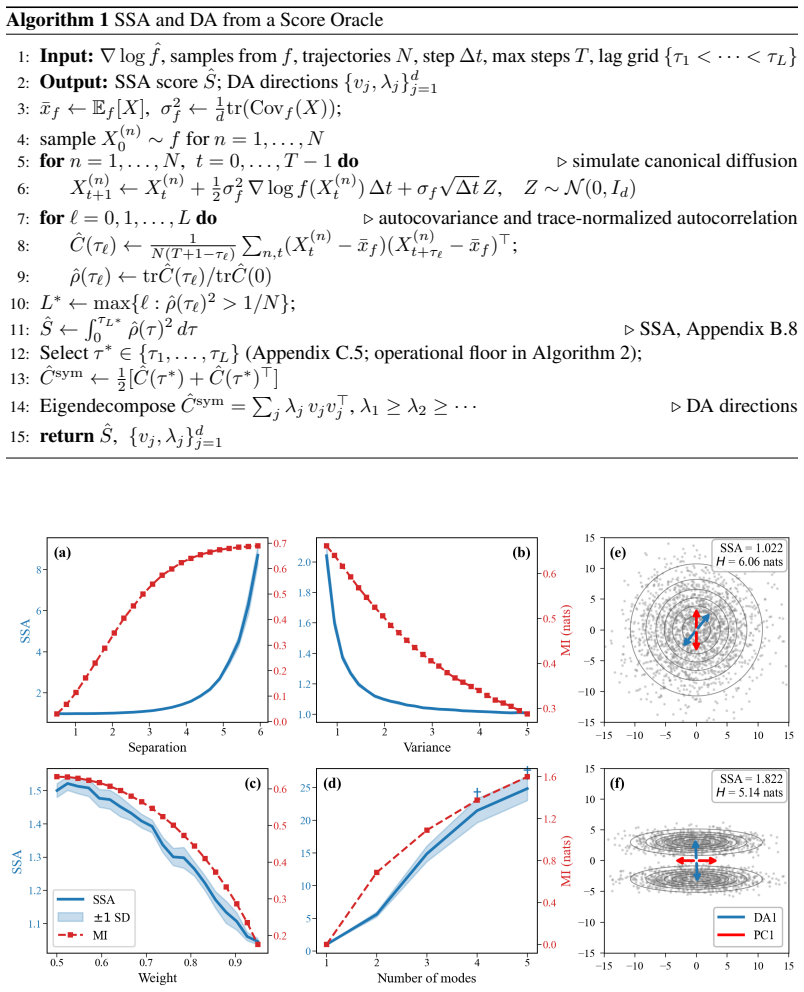
- In Gaussian mixtures SSA tracks mutual information between the latent mode variable and the observed samples.
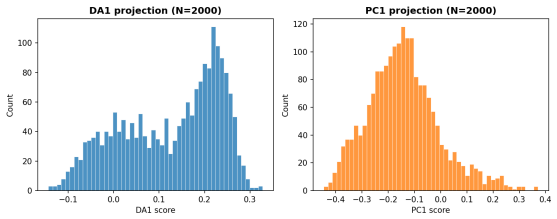
- In SDXL text-to-image outputs the readouts detect compositional structure that differential entropy and PCA both miss.
- For static samples of alanine dipeptide the DA directions recover the known slow backbone dihedrals.
- The Marchenko-Pastur-type edge for the null spectrum gives an automatic rule for choosing the lag at which DA is evaluated.
- Everything runs from samples plus a score function, allowing direct use inside pretrained score-based generative models.
Where Pith is reading between the lines
- The same construction could be applied to any density for which a score estimator exists, including those learned from unlabeled data in other scientific domains.
- Because DA orders directions by decorrelation time rather than variance, it may serve as a drop-in replacement for PCA in tasks where slow mixing matters more than spread.
- The closed-form null spectrum suggests a direct test for whether an empirical density deviates from Gaussianity in its barrier structure.
Load-bearing premise
The diffusion defined by reversibility plus constant scalar coefficient is the right intrinsic process whose autocovariance isolates barrier crossing rather than other geometric features.
What would settle it
Apply the method to a known two-mode Gaussian mixture whose mutual information is independently computable; if SSA fails to increase monotonically with the separation parameter while other measures do, the isolation claim is falsified.
Figures
















read the original abstract
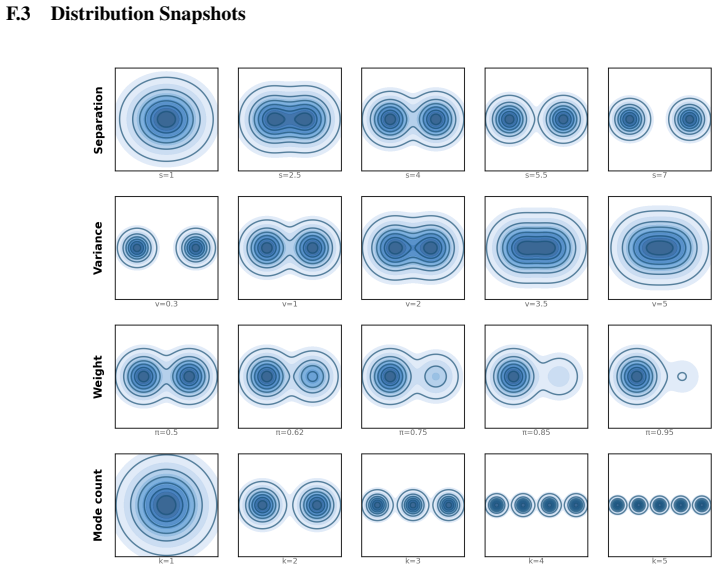
Mode separation, namely how sharply a distribution fragments into barrier-separated clusters, is a fundamental geometric property of densities, difficult to quantify in high dimensions. It is structurally distinct from dispersion, yet existing tools fall short: differential entropy rises with spread regardless of fragmentation, PCA orders directions by variance regardless of barriers, and mutual information requires a mixture decomposition one usually does not have. We measure mode separation through a single stochastic process intrinsic to the density: a unique reversible diffusion with $f$ as its stationary distribution and constant scalar diffusion coefficient. We extract two readouts from its autocovariance matrix: SSA (Sum of Squared Autocorrelations), a scalar barrier-sensitive measure; and DA (Dominant Autocorrelation directions), linear projections ordered by metastability rather than variance. Under an isotropic-Gaussian null, we derive a closed-form spectrum for the empirical autocovariance that generalizes Marchenko--Pastur, with an analytic upper edge that selects the lag at which DA is read off. Both readouts use only samples and a score function, scaling to high dimensions through pretrained score-based generative models via Tweedie's identity. We apply our framework to three settings: (i) synthetic Gaussian mixtures, where SSA tracks mutual information; (ii) SDXL text-to-image generations, where SSA and DA capture structure that entropy and PCA miss; and (iii) molecular dynamics of alanine dipeptide, where DA recovers the known slow backbone dihedrals from static samples alone.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper claims that mode separation (barrier-induced fragmentation of a density) can be quantified and decomposed via the autocovariance operator of the unique reversible diffusion having the target density f as stationary measure and constant scalar diffusion coefficient. From this operator it extracts SSA (sum of squared autocorrelations), a scalar barrier-sensitive statistic, and DA (dominant autocorrelation directions), linear projections ordered by metastability. Under an isotropic-Gaussian null it derives a closed-form spectrum for the empirical autocovariance that generalizes the Marchenko-Pastur law, yielding an analytic edge for lag selection. Both readouts require only samples and a score function and are demonstrated on Gaussian mixtures (where SSA tracks mutual information), SDXL text-to-image outputs, and alanine-dipeptide MD trajectories (where DA recovers known slow dihedrals from static samples).
Significance. If the central invariance claim holds, the framework supplies a new, intrinsically defined, parameter-free diagnostic for high-dimensional fragmentation that is orthogonal to variance-based (PCA) and entropy-based measures. The analytic null spectrum, the scaling route through pretrained score models via Tweedie’s identity, and the concrete recovery of known slow modes in alanine dipeptide are notable strengths that would make the method immediately usable for generative-model evaluation and molecular-dynamics analysis.
major comments (3)
- [§2.1–2.2] The manuscript asserts that the autocovariance of the chosen canonical diffusion isolates barrier crossing rather than local curvature, anisotropy or higher moments, yet provides no theorem establishing invariance of SSA or the DA spectrum under non-barrier geometric perturbations while retaining sensitivity to barrier height. The empirical checks on Gaussian mixtures and alanine dipeptide are consistent but do not rule out confounding (see the paragraph introducing the canonical process and the subsequent definition of SSA/DA).
- [§3.1, Eq. (12)–(15)] The closed-form spectrum under the isotropic-Gaussian null (generalizing Marchenko-Pastur) is stated without the intermediate expansion of the autocovariance operator or the verification that cross terms vanish; it is therefore unclear whether the analytic upper edge for lag selection remains valid once the diffusion is discretized or the score is estimated (see the derivation of the null spectrum).
- [§5.3, Figure 7] In the alanine-dipeptide experiment the claim that DA recovers the known slow backbone dihedrals relies on visual inspection of the leading directions; a quantitative comparison (e.g., overlap with the true slow subspace or comparison against a variance-ordered baseline) is missing, weakening the assertion that metastability rather than variance is being recovered.
minor comments (2)
- [§2.3 and §4] Notation for the lag parameter and the empirical autocovariance matrix is introduced inconsistently between the theoretical and experimental sections; a single consolidated definition would improve readability.
- [Figure 3] The caption of Figure 3 does not state the number of samples or the score-estimation procedure used for the SDXL experiment, making the reported SSA values difficult to reproduce.
Simulated Author's Rebuttal
We thank the referee for the constructive and detailed report. We address each major comment below, providing clarifications on the theoretical construction and indicating revisions that will strengthen the manuscript.
read point-by-point responses
-
Referee: [§2.1–2.2] The manuscript asserts that the autocovariance of the chosen canonical diffusion isolates barrier crossing rather than local curvature, anisotropy or higher moments, yet provides no theorem establishing invariance of SSA or the DA spectrum under non-barrier geometric perturbations while retaining sensitivity to barrier height. The empirical checks on Gaussian mixtures and alanine dipeptide are consistent but do not rule out confounding (see the paragraph introducing the canonical process and the subsequent definition of SSA/DA).
Authors: The canonical diffusion is the unique reversible process with stationary density f and constant scalar diffusion coefficient, so its infinitesimal generator is L = ∇log f · ∇ + Δ. The autocovariance operator is therefore completely determined by the score ∇log f, which encodes barrier heights via the underlying potential. We do not assert invariance under arbitrary geometric changes; rather, the constant-diffusion construction removes local anisotropy by design while preserving sensitivity to global mode separation induced by barriers. To make this explicit, we will add a short proposition in §2.2 relating SSA/DA to barrier height under controlled perturbations of local curvature, together with a brief discussion of the scope of the invariance. revision: yes
-
Referee: [§3.1, Eq. (12)–(15)] The closed-form spectrum under the isotropic-Gaussian null (generalizing Marchenko-Pastur) is stated without the intermediate expansion of the autocovariance operator or the verification that cross terms vanish; it is therefore unclear whether the analytic upper edge for lag selection remains valid once the diffusion is discretized or the score is estimated (see the derivation of the null spectrum).
Authors: We agree that the derivation was condensed. In the revised version we will move the full expansion of the autocovariance operator and the verification that cross terms vanish to a dedicated appendix. The analytic upper edge is derived for the continuous-time process with exact score; we will add a remark clarifying its status under Euler–Maruyama discretization and score estimation error, supported by additional Monte-Carlo checks that confirm the edge remains a reliable lag selector in the regimes used in the experiments. revision: yes
-
Referee: [§5.3, Figure 7] In the alanine-dipeptide experiment the claim that DA recovers the known slow backbone dihedrals relies on visual inspection of the leading directions; a quantitative comparison (e.g., overlap with the true slow subspace or comparison against a variance-ordered baseline) is missing, weakening the assertion that metastability rather than variance is being recovered.
Authors: We accept that a quantitative metric would strengthen the claim. In the revision we will compute the principal angles (or Grassmann distance) between the leading DA directions and the known slow dihedral subspace, and report the same overlap for a variance-ordered PCA baseline on identical data. These numbers, together with an updated Figure 7, will be added to §5.3. revision: yes
Circularity Check
No significant circularity; derivation relies on standard stochastic process definitions and independent random-matrix results
full rationale
The central construction selects the unique reversible diffusion with given stationary density f and constant scalar diffusion coefficient; this is a standard, externally defined object in the theory of reversible Markov processes and does not reduce to any fitted quantity or target mode-separation statistic. The null-spectrum derivation is obtained by applying classical Marchenko-Pastur-type analysis to the empirical autocovariance matrix under an isotropic-Gaussian null model, yielding a closed-form edge that is parameter-free with respect to the data and independent of the mode-separation claim. SSA and DA are then read off from the same autocovariance operator; they are not fitted to any external measure of barrier crossing. No load-bearing step invokes a self-citation whose content is itself unverified or whose uniqueness theorem is imported from the same authors without external grounding. The framework therefore remains self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Existence and uniqueness of a reversible diffusion with constant scalar diffusion coefficient having f as stationary distribution
Reference graph
Works this paper leans on
-
[1]
Diffusion maps.Applied and Computational Har- monic Analysis, 21(1):5–30, 2006
Ronald R Coifman and St ´ephane Lafon. Diffusion maps.Applied and Computational Har- monic Analysis, 21(1):5–30, 2006
work page 2006
-
[2]
Diffusion maps, spectral clustering and eigenfunctions of Fokker–Planck operators
Boaz Nadler, St ´ephane Lafon, Ronald R Coifman, and Ioannis G Kevrekidis. Diffusion maps, spectral clustering and eigenfunctions of Fokker–Planck operators. InAdvances in Neural Information Processing Systems (NeurIPS), volume 18, 2005
work page 2005
-
[3]
The dip test of unimodality.The Annals of Statistics, 13(1):70–84, 1985
John A Hartigan and Pamela M Hartigan. The dip test of unimodality.The Annals of Statistics, 13(1):70–84, 1985
work page 1985
-
[4]
Jakub Rydzewski. Spectral map: Embedding slow kinetics in collective variables.The Journal of Physical Chemistry Letters, 14(22):5216–5220, 2023
work page 2023
-
[5]
Jakub Rydzewski. Spectral map for slow collective variables, Markovian dynamics, and transi- tion state ensembles.Journal of Chemical Theory and Computation, 20(18):7775–7784, 2024
work page 2024
-
[6]
Lutz Molgedey and Heinz Georg Schuster. Separation of a mixture of independent signals using time delayed correlations.Physical Review Letters, 72(23):3634–3637, 1994
work page 1994
-
[7]
Guillermo P ´erez-Hern´andez, Fabian Paul, Toni Giorgino, Gianni De Fabritiis, and Frank No´e. Identification of slow molecular order parameters for Markov model construction.The Journal of Chemical Physics, 139(1):015102, 2013
work page 2013
-
[8]
Grigorios A Pavliotis.Stochastic Processes and Applications: Diffusion Processes, the Fokker–Planck and Langevin Equations, volume 60 ofTexts in Applied Mathematics. Springer, 2014
work page 2014
-
[9]
B. W. Silverman. Using kernel density estimates to investigate multimodality.Journal of the Royal Statistical Society: Series B (Methodological), 43(1):97–99, 1981
work page 1981
-
[10]
Denoising diffusion probabilistic models
Jonathan Ho, Ajay Jain, and Pieter Abbeel. Denoising diffusion probabilistic models. In Advances in Neural Information Processing Systems (NeurIPS), volume 33, pages 6840–6851, 2020
work page 2020
-
[11]
Generative modeling by estimating gradients of the data dis- tribution
Yang Song and Stefano Ermon. Generative modeling by estimating gradients of the data dis- tribution. InAdvances in Neural Information Processing Systems (NeurIPS), volume 32, 2019
work page 2019
-
[12]
Score-based generative modeling through stochastic differential equations
Yang Song, Jascha Sohl-Dickstein, Diederik P Kingma, Abhishek Kumar, Stefano Ermon, and Ben Poole. Score-based generative modeling through stochastic differential equations. In International Conference on Learning Representations (ICLR), 2021
work page 2021
-
[13]
Flow matching for generative modeling
Yaron Lipman, Ricky T Q Chen, Heli Ben-Hamu, Maximilian Nickel, and Matt Le. Flow matching for generative modeling. InInternational Conference on Learning Representations (ICLR), 2023
work page 2023
-
[14]
High-resolution image synthesis with latent diffusion models
Robin Rombach, Andreas Blattmann, Dominik Lorenz, Patrick Esser, and Bj ¨orn Ommer. High-resolution image synthesis with latent diffusion models. InProceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pages 10684–10695, 2022
work page 2022
-
[15]
SDXL: Improving latent diffusion models for high-resolution image synthesis
Dustin Podell, Zion English, Kyle Lacey, Andreas Blattmann, Tim Dockhorn, Jonas M ¨uller, Joe Penna, and Robin Rombach. SDXL: Improving latent diffusion models for high-resolution image synthesis. InThe Twelfth International Conference on Learning Representations (ICLR), 2024
work page 2024
-
[16]
Learning transferable visual models from natural language supervision
Alec Radford, Jong Wook Kim, Chris Hallacy, Aditya Ramesh, Gabriel Goh, Sandhini Agar- wal, Girish Sastry, Amanda Askell, Pamela Mishkin, Jack Clark, Gretchen Krueger, and Ilya Sutskever. Learning transferable visual models from natural language supervision. InInterna- tional Conference on Machine Learning (ICML), pages 8748–8763. PMLR, 2021
work page 2021
-
[17]
Computational Molecular Biology Group, Freie Universit ¨at Berlin. MDShare: Download- ing sample datasets for molecular dynamics analysis.https://markovmodel.github.io/ mdshare. Accessed: 2026-05-06. 10
work page 2026
-
[18]
Monte Carlo methods in statistical mechanics: Foundations and new algorithms
Alan Sokal. Monte Carlo methods in statistical mechanics: Foundations and new algorithms. In Cecile DeWitt-Morette, Pierre Cartier, and Antoine Folacci, editors,Functional Integration: Basics and Applications, pages 131–192. Springer, 1997
work page 1997
-
[19]
Practical Markov chain Monte Carlo.Statistical Science, 7(4):473–483, 1992
Charles J Geyer. Practical Markov chain Monte Carlo.Statistical Science, 7(4):473–483, 1992
work page 1992
-
[20]
A. W. van der Vaart.Asymptotic Statistics. Cambridge Series in Statistical and Probabilistic Mathematics. Cambridge University Press, 1998
work page 1998
-
[21]
Vladimir A Mar ˇcenko and Leonid A Pastur. Distribution of eigenvalues for some sets of random matrices.Mathematics of the USSR-Sbornik, 1(4):457–483, 1967
work page 1967
-
[22]
Monika Bhattacharjee, Arup Bose, and Apratim Dey. Joint convergence of sample cross- covariance matrices.ALEA: Latin American Journal of Probability and Mathematical Statis- tics, 20:395–423, 2023. doi: 10.30757/ALEA.v20-14
-
[23]
Santosh Kumar and S. Sai Charan. Spectral statistics for the difference of two wishart matrices. Journal of Physics A: Mathematical and Theoretical, 53(50):505202, 2020. doi: 10.1088/ 1751-8121/abc3fe
work page 2020
-
[24]
American Mathematical Society, 1992
Dan V V oiculescu, Kenneth J Dykema, and Alexandru Nica.Free Random Variables, volume 1 ofCRM Monograph Series. American Mathematical Society, 1992
work page 1992
-
[25]
Cambridge University Press, 2006
Alexandru Nica and Roland Speicher.Lectures on the Combinatorics of Free Probability, volume 335 ofLondon Mathematical Society Lecture Note Series. Cambridge University Press, 2006
work page 2006
-
[26]
Zhidong Bai and Jack W Silverstein.Spectral Analysis of Large Dimensional Random Matri- ces. Springer Series in Statistics. Springer, 2nd edition, 2010
work page 2010
-
[27]
Jinho Baik, G ´erard Ben Arous, and Sandrine P´ech´e. Phase transition of the largest eigenvalue for nonnull complex sample covariance matrices.The Annals of Probability, 33(5):1643–1697, 2005
work page 2005
-
[28]
Florent Benaych-Georges and Raj Rao Nadakuditi. The eigenvalues and eigenvectors of finite, low rank perturbations of large random matrices.Advances in Mathematics, 227(1):494–521, 2011
work page 2011
-
[29]
Zhigang Bao, L ´aszl´o Erd˝os, and Kevin Schnelli. Local stability of the free additive convolu- tion.Journal of Functional Analysis, 271(3):672–719, 2016
work page 2016
-
[30]
Ji Oon Lee and Kevin Schnelli. Tracy–Widom distribution for the largest eigenvalue of real sample covariance matrices with general population.The Annals of Applied Probability, 26 (6):3786–3839, 2016
work page 2016
-
[31]
Grigorios A. Pavliotis and Andrew M. Stuart.Multiscale Methods: Averaging and Homoge- nization, volume 53 ofTexts in Applied Mathematics. Springer, 2008
work page 2008
-
[32]
Anton Bovier and Frank den Hollander.Metastability: A Potential-Theoretic Approach, vol- ume 351 ofGrundlehren der mathematischen Wissenschaften. Springer, 2015
work page 2015
-
[33]
H. A. Kramers. Brownian motion in a field of force and the diffusion model of chemical reactions.Physica, 7(4):284–304, 1940
work page 1940
-
[34]
Princeton University Press, 1967
Edward Nelson.Dynamical Theories of Brownian Motion. Princeton University Press, 1967. 11 A Canonical Diffusion This appendix collects the uniqueness proof of the canonical diffusion (Theorem 1), the proof of similarity invariance (Proposition 1), and the design rationale for the choice of diffusion matrix. A.1 Proof of Theorem 1 We assumef:R d →(0,∞)is s...
work page 1967
-
[35]
at least one relaxation time out,
The2×2transformation ( ˜X, Z)7→(A, B)is orthogonal, so(A, B)has the same joint i.i.d.N(0, I d)distribution as( ˜X, Z), with columns ofAandBindependent of one another. Then ˜X= (A−B)/ √ 2andZ= (A+ B)/ √ 2, so direct expansion gives ˜X ˜X ⊤ = 1 2(AA⊤ +BB ⊤ −AB ⊤ −BA ⊤)andZ ˜X ⊤ + ˜XZ ⊤ = AA⊤ −BB ⊤, so 2N ˆCsym(τ) = (ασ 2 +βσ)| {z } P AA⊤ + (ασ 2 −βσ)| {z } ...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.